Abstract
Flaviviruses infect hundreds of millions of people annually, and no antiviral therapy is available1,2. We performed a genome-wide CRISPR/Cas9-based screen to identify host genes that, when edited, resulted in reduced flavivirus infection. Here, we validated nine human genes required for flavivirus infectivity, and these were associated with endoplasmic reticulum functions including translocation, protein degradation, and N-linked glycosylation. In particular, a subset of endoplasmic reticulum-associated signal peptidase complex (SPCS) proteins was necessary for proper cleavage of the flavivirus structural proteins (prM and E) and secretion of viral particles. Loss of SPCS1 expression resulted in markedly reduced yield of all Flaviviridae family members tested (West Nile, dengue, Zika, yellow fever, Japanese encephalitis, and hepatitis C viruses), but had little impact on alphavirus, bunyavirus, or rhabdovirus infection or the surface expression or secretion of diverse host proteins. We found that SPCS1 dependence could be bypassed by replacing the native prM protein leader sequences with a class I major histocompatibility complex (MHC) antigen leader sequence. Thus, SPCS1, either directly or indirectly via its interactions with host proteins, preferentially promotes the processing of specific protein cargo, and Flaviviridae have a unique dependence on this signal peptide processing pathway. SPCS1 and other signal processing pathway members could represent pharmacological targets for inhibiting infection by the expanding number of flaviviruses of medical concern.
Similar content being viewed by others
Main
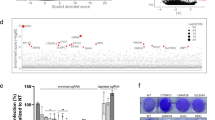
We performed a genome-wide inhibition of West Nile virus (WNV)-induced cell death screen using the CRISPR/Cas9 system3,4,5,6,7 and lentiviruses targeting 19,050 genes (Extended Data Fig. 1a). Whereas in the absence of lentivirus transduction cells did not survive WNV infection, colonies of lentivirus-transduced cells survived; single guide RNAs (sgRNAs) were amplified by PCR and sequenced. We identified 12 genes that were statistically enriched using MAGeCK8 (Supplementary Tables 1, 2). All 12 genes were endoplasmic reticulum-associated with annotated functions of carbohydrate modification, protein translocation and signal peptide processing, protein degradation, and heat shock response (Fig. 1a).
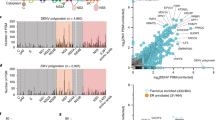
a, b, Genes were selected for validation based on statistical analysis (Supplementary Tables 1 and 2). Gene-edited 293T (a) and HeLa cells (b) were infected with WNV and analysed 12 h later for E protein. c–f, Effect of gene editing on ZIKV (c), JEV (d), DENV-2 (e), and YFV (f) infection in 293T cells. The results are the average of two or three independent experiments. g, h, 293T cells expressing indicated sgRNAs were infected with WNV (g) or JEV (h) and virus yield was determined. One of two independent experiments performed in triplicate is shown. Statistical significance was determined by ANOVA with a multiple comparisons correction (*P < 0.05, **P < 0.01, ***P < 0.0001; a–f). Error bars indicate s.e.m. ER, endoplasmic reticulum; ERAD, endoplasmic reticulum-associated degradation.
In validation studies, editing of nine genes resulted in reduced WNV antigen expression following infection of 293T or HeLa cells (Fig. 1a, b) without causing cytotoxicity (Extended Data Fig. 1b). We confirmed the efficiency of gene editing for the proteins for which we could obtain validated antibodies (Extended Data Fig. 1c). Validated genes were tested for effects on related flaviviruses: Zika (ZIKV), Japanese encephalitis (JEV), dengue serotype 2 (DENV-2), and yellow fever (YFV) viruses. Editing of six of these genes reduced infection by all four flaviviruses (Fig. 1c–f). Editing of STT3A, SEC63, SPCS1, or SPCS3 resulted in decreased yields of WNV and JEV (Fig. 1g, h). We observed less impact on unrelated positive- or negative-sense RNA viruses (Extended Data Fig. 1d).
As pathogenic flaviviruses are transmitted by arthropods, we evaluated the roles of orthologues of these genes in insect cells. Silencing of Drosophila orthologues reduced infection by WNV and DENV-2 (Fig. 2a, b) without appreciably affecting cell viability (Fig. 2c). Decreased WNV infection was also observed in mosquito cells after gene silencing (Fig. 2d). Depletion of Spase22-23 (orthologue of SPCS3) in adult Drosophila led to decreased WNV titres (Fig. 2e) and flies heterozygous for Spase12 (orthologue of SPCS1) showed reduced WNV infection (Fig. 2f). Overall, flavivirus infectivity in human and insect cells was dependent on analogous endoplasmic reticulum-associated genes.
a, b, Drosophila DL1 cells were treated with dsRNA for 3 days and infected with WNV (Kunjin) (a) or DENV-2 (b) for 30 h. Gene names of human orthologues are given in parentheses. The percentage of infected cells was normalized to the control β-galactosidase dsRNA. The data are expressed as the mean normalized value ± s.d. Statistically significant differences were determined by Student’s t-test (**P < 0.01; ***P < 0.001; ****P < 0.0001) and were compared to control dsRNA. The data are pooled from four experiments in duplicate. c, Cell viability. DL1 cells were treated with dsRNA and processed 4 days later. d, Aedes agypti AAG2 cells were treated with dsRNA for 3 days, infected with WNV (Kunjin) for 30 h, and processed for viral antigens. e. SPCS1 silenced Drosophila (Hs-Gal4>UAS–Spase22-23 IR (inverted repeat)) or sibling controls were infected with WNV (Kunjin) and titres measured 7 days later. The fold-change in titres of pools of ten flies from three experiments is shown (normalized mean ± s.d., *P < 0.05 by Student’s t-test). f, Wild-type or Spase12(EY10774)+/− sibling flies were infected with WNV (Kunjin) and titres measured 7 days later. Data from pools of five flies in three independent experiments is shown (*P < 0.05 by Student’s t-test).
Trans-complementation of gene-edited human cells with wild-type alleles rescued flavivirus infectivity (Extended Data Fig. 1e–g). Since we identified the genes encoding two (SPCS1 and SPCS3) of the five components of the Signal Peptidase Complex9,10, and found that insect SPCS genes also affected flavivirus infection, we focused our study on these genes. Gene silencing in human cells confirmed that SPCS genes were required for optimal flavivirus but not alphavirus infection (Extended Data Fig. 2 and data not shown).
We screened for clonal SPCS1 and SPCS3 knockout cells lines. Although we were unable to obtain SPCS3−/− clonal lines, SPCS1−/− 293T or Huh7.5 cell clones grew, with both alleles containing nonsense deletions (Fig. 3a and Extended Data Fig. 3). WNV, DENV, JEV, YFV, and ZIKV failed to accumulate in the supernatants of SPCS1−/− 293T cells (Fig. 3c–f), and WNV infectivity was restored in trans-complemented cells (Fig. 3h). However, SPCS1−/− cells supported infection by alphaviruses, bunyaviruses, and rhabdoviruses (Fig. 3i–k and Extended Data Fig. 3a). To corroborate these findings, we tested SPCS1−/− Huh7.5 cells and found reduced infection by WNV, ZIKV, JEV, and the related Flaviviridae member, hepatitis C virus (Extended Data Fig. 3e, f). In comparison, gene editing of the remaining SPCS genes, SEC11A and SEC11C, had minimal effects on infection (Extended Data Fig. 4).
a, Western blotting of SPCS1−/− 293T cells. b, Cells were transfected with YFV–luciferase replicon RNA (wild-type GDD or loss-of-function GVD). Firefly luciferase activity was measured and normalized to intracellular protein levels. The data reflect the average of two or three independent experiments performed in duplicate. c–h, Cells were infected with WNV (c, h), DENV-2 (d), JEV (e), YFV (f) or ZIKV (g), and viral yield measured. In h, cells were trans-complemented with an SPCS1 or control plasmid. Results are the average of two or three independent experiments performed in triplicate. i–k, Cells were infected with CHIKV (alphavirus), RVFV (bunyavirus), or VSV (rhabdovirus) and viral yield was measured. Results are the average of two or three independent experiments performed in triplicate. l, The polyprotein processing strategy of flaviviruses13. Red and blue arrows indicate sites of cleavage by host and viral (NS2B–NS3) proteases, respectively. m–o, Control or SPCS1−/− 293T (m, o) or Huh7.5 (n) cells were infected with WNV (m, o) or JEV (n). Lysates were blotted with (m) anti-WNV E, (n) anti-JEV E, or (o) anti-WNV NS1 monoclonal antibodies. Higher molecular mass bands (Ehi, Emed, and NS1hi) that react with anti-flavivirus monoclonal antibodies are indicated. One experiment of three is shown. p, 293T cells were infected with CHIKV or RVFV. Lysates were blotted with anti-CHIKV E2 or anti-RVFV Gn monoclonal antibodies. One experiment of two is shown. For gel source data, see Supplementary Fig. 1. FFU, focus-forming unit; PFU, plaque-forming unit.
To determine whether SPCS1 was required for viral translation, replication, or both, we used wild-type and loss-of-function11 flavivirus replicons encoding reporter genes12 (Fig. 3b and Extended Data Fig. 5). Transfection of control cells with replicon RNA resulted in low levels of reporter gene activity over the first several hours, which reflects translation of input viral RNA, whereas subsequent signal increases are due to RNA replication. In SPCS1−/− cells, high levels of reporter gene expression were observed, indicating that viral RNA translation and replication remained largely intact.
We speculated that SPCS subunits, directly or indirectly, might regulate cleavage of the flavivirus polyprotein13. Flavivirus structural (prM and E) and non-structural (NS1 and NS4B) proteins are cleaved by unknown endoplasmic reticulum host signal peptidase(s) (Fig. 3l and refs 14, 15). Gene-edited 293T cells were infected with WNV or JEV, and lysates were analysed. Reduced levels of E and prM proteins were found in SPCS1−/− clones and SPCS1 or SPCS3 bulk gene-edited cells 12 h after infection, and by 24 h higher molecular mass bands reacted with anti-E or anti-prM/E antibodies16 (Fig. 3m, n and Extended Data Figs 3g, 6a, b). We next examined whether SPCS1 is required for cleavage of the viral non-structural proteins NS1–NS2A, 2K–NS4B, or NS2B–NS3. In SPCS1−/− cells, infection with WNV resulted in decreased expression of NS1 and the accumulation of higher molecular mass bands (Fig. 3o). We detected lower levels of NS4B protein in SPCS1−/− cells; in transfection studies with a tagged 2K–NS4B plasmid, a higher molecular mass band was observed. For NS1–NS2A and NS3, we did not detect aberrant cleavage (Extended Data Fig. 6). We also tested the effects on HCV E2 glycoprotein and found decreased levels in SPCS1−/− cells (Extended Data Fig. 7). In comparison, alphavirus or bunyavirus glycoproteins, which also require endoplasmic reticulum processing17,18, showed intact expression in SPCS1−/− cells (Fig. 3p and Extended Data Fig. 3b, c).
To isolate the effects of the SPCS complex from infection, we transfected a prM–E plasmid, which produces subviral particles (SVPs)19. Immunoblotting of cell lysates for E and prM proteins showed reduced levels and higher molecular mass bands in SPCS1- or SPCS3-deficient cells, and these changes correlated with a reduction in the number of secreted SVPs (Extended Data Fig. 8a–c). We tested whether cleavage of flavivirus protein signal sequences depended on SPCS1. We transfected WNV structural (capsid (C), prM, M, E) and secreted non-structural (NS1) genes with native or MHC class I (Kb) signal sequences into SPCS1−/− cells, and evaluated protein expression (Fig. 4).
a, 293T cells were transfected with prM or prM–E plasmids containing native (C–prM (green box) and prM–E (blue box)) or Kb (red box) leader sequences. Blots of lysates were probed with anti-prM/M monoclonal antibodies. One experiment of three is shown. b, 293T cells were transfected with E, prM–E, and E–NS1 plasmids containing native leader sequences (as in a and E–NS1 (orange box)) or a Kb leader. Blots of lysates were probed with anti-E monoclonal antibodies. A higher molecular mass band corresponds to uncleaved E–NS1. One experiment of two is shown. c, d, 293T cells were transfected with a prM–E plasmid. After 24 h, cells were labelled for 40 min (c) or 3 min (d) with 35S cysteine-methionine. Lysates were immunoprecipitated with an anti-E protein monoclonal antibody before SDS–PAGE. c, Top, cysteine–methionine was added for chase times (0–4 h). c (bottom) and d, Immunoprecipitates were untreated or treated with Endo H or PNGase F for 1 h at 37 °C. One experiment of two is shown. e, 293T cells were transfected with NS1, E–NS1, or prM–E–NS1 plasmids containing native viral or Kb leaders. Blots of lysates or supernatants were probed with anti-NS1 monoclonal antibodies. One experiment of two is shown. f, 293T cells were transfected with prM + E, prM–E, or E plasmids containing viral or Kb leaders. Left, blots of lysates were probed with anti-prM/M or anti-E monoclonal antibodies. One experiment of two is shown. Right, levels of SVPs in supernatant at 24 h. Data are pooled from independent experiments performed in triplicate (**P < 0.01, ***P < 0.001, ****P < 0.0001; unpaired t-test). g, 293T cells were transfected with prM–Flag. After 24 h, cells were labelled for 3 min with 35S cysteine–methionine. Lysates were immunoprecipitated with anti-Flag protein monoclonal antibodies. h, Model of processing of flavivirus structural and non-structural proteins based on infection and transfection studies and the literature14,15. Arrows indicate cleavage sites requiring SPCS1, sites affected by upstream SPCS1-dependent events, sites cleaved by the viral NS2B–NS3 protease, and sites cleaved via an SPCS1-independent pathway. For gel source data, see Supplementary Fig. 1.
Expression of C protein from a C–prM–E plasmid was equivalent in control and SPCS1−/− cells, although in the absence of the viral protease, C did not migrate at its normal size (Extended Data Fig. 8d). However, cleavage of the downstream proteins prM and E was reduced in SPCS1−/− cells. When NS2B–NS3 was supplied in trans, C was cleaved from prM–E and accumulated at the correct size in control and SPCS1−/− cells. Thus, expression or cleavage of C is not affected by SPCS1.
We next evaluated expression of prM and M. When the native prM leader sequence was used, expression of prM and its furin-cleavage product M was reduced in SPCS1−/− cells (Fig. 4a, groups 1 and 3). Substitution of the Kb leader rescued prM and M expression in SPCS1−/− cells only when prM was on a separate plasmid (Fig. 4a, group 2) but not as a prM–E plasmid (Fig. 4a, group 4). Thus, specific leader sequences determine the dependence of prM and M protein expression on SPCS1, and downstream proteins can modulate processing efficiency.
When E was transfected, its expression was largely independent of SPCS1 or the Kb leader sequence (Fig. 4b, groups 1 and 2). When E was cloned downstream of prM, accumulation of E was not detected in SPCS1−/− cells (Fig. 4b, groups 3 and 4). This finding suggested that the native leader sequence of E was not cleaved in SPCS1−/− cells when presented as an ‘internal’ leader sequence or that epistatic effects of the upstream prM protein reduced the stability of E protein. To test which of these possible explanations was correct, we performed 35S pulse-chase studies in prM–E-transfected cells. In control cells, only a single E protein band was visible, indicating rapid prM–E cleavage. However, prM–E and E bands were both present in SPCS1−/− cells (Fig. 4c, top) and remained in an endoplasmic reticulum-resident form (Fig. 4c, bottom). A short 3-min 35S pulse showed a delay in the cleavage of prM–E in SPCS1−/− cells (Fig. 4d).
We assessed the expression of NS1, which also requires endoplasmic reticulum-dependent signal sequence cleavage. When NS1 was transfected into cells, SPCS1 was not required for expression (Fig. 4e, group 1). When NS1 was cloned downstream of E (Fig. 4e, groups 2 and 3) or prM–E (Fig. 4e, groups 4 and 5), NS1 levels were reduced in SPCS1−/− cells. After blotting with an anti-NS1 monoclonal antibody, a 90-kDa band was visible in blots from SPCS1−/− cells (Fig. 4e, group 2), which probably represented uncleaved E–NS1; this result was corroborated by blotting for E protein (Fig. 4b, groups 5 and 6). Thus, placement of the NS1 leader sequence into an internal position rendered it more dependent on SPCS1 for cleavage.
Flavivirus SVPs can be produced after transfection of prM and E on single or separate plasmids20,21. Transfection of prM–E encoding native or Kb and native internal signal sequences resulted in loss of expression of prM and E or SVPs in SPCS1−/− cells (Fig. 4f, groups 5 and 6). When prM and E were co-transfected, the proteins were detected in SPCS1−/− cell lysates (Fig. 4f, groups 1 and 2) and supernatant, albeit at lower levels. In SPCS1−/− cells, prM negatively affected E but not NS1 production (Fig. 4f (compare groups 1, 2, and 7) and Extended Data Fig. 8e), possibly because of its chaperone-like function for E protein20. Defects in co-expression of prM and E in SPCS1−/− cells were corrected by inserting the Kb leader sequence in front of the prM gene (Fig. 4f, groups 3 and 4). A 3-min 35S pulse and immunoprecipitation experiment in SPCS1−/− cells showed an uncleaved form of prM (Fig. 4g).
To assess whether host surface proteins require SPCS1 for signal peptide processing, we profiled SPCS1−/− Jurkat T cells. Whereas ten antigens showed no difference in surface expression, levels of CD49d–CD29, ULBP1, and HLA-E were reduced by two-to-threefold (Extended Data Fig. 9a–c). A decrease in surface expression of ULBP1 has been reported in cells deficient in SPCS1 or SPCS2 expression22, although this phenotype was not explored. In an unbiased approach, we analysed the secretome of SPCS1−/− 293T cells by mass spectrometry. Of the approximately 245 secreted proteins identified, only 35 were downregulated in SPCS1−/− cells, and the fold-changes were small (Extended Data Fig. 10 and Supplementary Table 4). We validated 3 of 5 as being reduced in supernatants of SPCS1−/− cells (Supplementary Table 5). Despite profound effects on flavivirus protein processing, an absence of SPCS1 only modestly affected the expression of host proteins.
The differential requirement of SPCS1 for viral and host protein processing suggests that components of the SPCS complex in mammalian and probably insect cells facilitate the cleavage of particular signal peptides in specific contexts. There may be additional requirements for some viruses, as interactions between SPCS1 and the HCV NS2 and E2 proteins have been reported23.
A recent study performed an analogous CRISPR-based screen with WNV24. Endoplasmic reticulum-associated genes were identified that prevented WNV-induced cell death. We identified three of these genes (EMC4, EMC6, and SEL1L), as did an siRNA screen25. Virtually all human gene ‘hits’ identified in our screen had insect orthologues required for optimal flavivirus infection. A subset of our genes were also identified in RNAi screens in Drosophila cells26,27. The endoplasmic reticulum is a focal site in the flavivirus lifecycle because it supports translation, polyprotein processing, replication, and virion morphogenesis. The identification of host gene targets that are selectively required for flavivirus infection but not cell survival provides intriguing candidates for pharmacological inhibition.
Methods
Cells and viruses
Vero, BHK21, HeLa, U2OS, Huh7.5, and 293T cells were cultured at 37 °C in Dulbecco’s Modified Eagle Medium (DMEM) supplemented with 10% fetal bovine serum (FBS). C6/36 Aedes albopictus cells were cultured at 28 °C in L15 supplemented with 10% FBS and 25 mM HEPES pH 7.3. Drosophila DL1 and Aedes Aag2 cells were cultured at 28 °C in Schneider’s medium supplemented with 10% FBS as described28. Jurkat cells were cultured at 37 °C in RPMI 1640 supplemented with 10% FBS and 10 mM HEPES pH 7.3. All cell lines were originally acquired from American Type Culture Collection or colleagues (Huh7.5) and were tested and judged free of mycoplasma contamination. The following viruses were used in screening and validation studies: WNV (New York 2000 and Kunjin), JEV (14-14-2 vaccine and Bennett strains), DENV-2 (16681 and New Guinea C strains), ZIKV (H/PF/2013), YFV (17D vaccine), CHIKV (2006 La Reunion OPY1), LACV (original strain), VSV (Indiana), HCV (J6/JFH), and SINV (Toto). With the exception of HCV (see below), all other viruses were propagated in BHK21, Vero, or C6/36 cells and titrated by standard plaque or focus-forming assays29.
Viral growth analysis
293T or Huh7.5 cells were infected with WNV (multiplicity of infection (MOI) 0.01), JEV (14-14-2 strain, MOI 0.05 or 0.5; Bennett strain, MOI 0.05), DENV-2 (MOI 3), YFV (MOI 1), ZIKV (MOI 0.05), CHIKV (MOI 0.01), SINV (MOI 0.01), RVFV (MOI 1), or VSV (MOI 0.01). After 2 h of incubation, cells were washed three times and samples were titrated on Vero cells. For HCV growth analysis, control and SPCS1 gene-edited Huh7.5 cells were inoculated at an MOI of 1 with virus derived from a growth-adapted JFH-1 infectious clone30. Cells were rinsed 6 h after infection to remove unbound virus and samples were collected every 24 h for 7 days. Viral titres in the supernatant were quantified by focus-forming assay, as described previously31.
Pooled sgRNA screen and data analysis
A pooled library encompassing 122,411 different sgRNAs against 19,050 human genes was derived by the Zhang laboratory32 and obtained from a commercial source (Addgene). The library was packaged using a lentivirus expression system and 293T cells were transfected using FugeneHD (Promega). Forty-eight hours after transfection, supernatants were collected, clarified by centrifugation (3,500 rpm × 20 min), filtered, and aliquotted for storage at −80 °C.
For the screen, we generated clonal 293T-Cas9 cells by transfecting the lentiCas9-Blast plasmid (Addgene 52962) using FugeneHD transfection reagent (Promega), blasticidin selection, and limiting dilution. These 293T-Cas9 cells were transduced with lentiviruses encoding individual sgRNAs at an MOI of 0.3. After selection with puromycin for 7 days, ~2 × 108 cells were infected with WNV (MOI of 1) and then incubated for 2–3 weeks. In parallel, untransduced 293T-Cas9 cells were infected to ensure virus-induced infection and cell death. The experiments were performed in parallel as either duplicate or triplicate technical replicates in two independent biological experiments.
Genomic DNA was extracted from the uninfected cells (5 × 107) or the cells (3 × 107) that survived WNV infection, and sgRNA sequences were amplified5, and subjected to next generation sequencing using an Illumina HiSeq 2500 platform. The sgRNA sequences against specific genes were recovered after removal of the tag sequences using the FASTX-Toolkit (http://hannonlab.cshl.edu/fastx_toolkit/) and cutadapt 1.8.1.
Gene validation
The cut-off for candidate gene ‘hits’ was made using a published computational tool (MAGeCK)8 and reflected sequencing data showing multiple different sgRNAs per gene, the number of sequencing reads per gene, and the enrichment of a given sgRNA compared to the uninfected cell library (Supplementary Tables 1, 2). From this, we identified 12 genes that showed statistically significant enrichment compared to uninfected cells. These candidate genes were tested for validation by using 3–5 independent sgRNAs per gene from the parent library and cloning them into the plasmid pSpCas9(BB)-2A-Puro (PX459) V2.0 (Addgene plasmid 62988). The control sgRNAs were used from the parent library. Plasmids were transfected into 293T or HeLa cells using FugeneHD transfection reagent and puromycin was added one day later. Three days later, puromycin was removed, and cells were allowed to recover for three additional days before infection with different viruses.
For flow cytometric analyses, gene-edited 293T cells were infected with WNV (MOI 5, 12 h), JEV (MOI 50, 22 h), ZIKV (MOI 10, 30 h), DENV-2 (MOI 3, 32 h), YFV (MOI 3, 38 h), CHIKV (MOI 1, 6 h), SINV (MOI 10, 6 h), LACV (MOI 5, 6 h), or VSV-GFP (MOI 1, 5.5 h). Gene-edited HeLa cells were infected with WNV (MOI 3, 24 h). At the indicated times, cells were fixed with 1% paraformaldehyde (PFA) diluted in PBS for 20 min at room temperature and permeabilized with Perm buffer (HBSS (Invitrogen), 10 mM HEPES, 0.1% (w/v) saponin (Sigma), and 0.025% NaN3 (Sigma)) for 10 min at room temperature. Cells then were rinsed one additional time with Perm buffer, transferred to a U-bottom plate, and incubated for 1 h at 4 °C with 1 μg ml−1 of the following virus-specific antibodies: WNV (human E16 (ref. 33)); DENV2 (mouse E18 (ref. 34)); JEV (mouse E18 (ref. 34)); YFV (mouse E60 (ref. 34)); CHIKV (CHK-11 (ref. 35)); SINV (ascites fluid, ATCC VR-1248AF), LACV (807-31 and 807-33, gift from A. Pekosz). After washing, cells were incubated with an Alexa Fluor 647-conjugated goat anti-mouse or anti-human IgG (Invitrogen) for 1 h at 4 °C. Cells were fixed in 1% PFA in PBS, processed on a FACS Array (BD Biosciences), and analysed using FlowJo software (Tree Star).
Validation also was performed by an infectious virus yield assay. Bulk gene-edited 293T cells were infected with WNV (MOI 0.01) or JEV (MOI 0.5). Supernatants were collected at specific times after infection and focus-forming assays were performed in 96-well plates as described previously36. Following infection, cell monolayers were overlaid with 100 μl per well of medium (1 × DMEM, 4% FBS) containing 1% carboxymethylcellulose, and incubated for 22 h (WNV) or 36 h (JEV) at 37 °C with 5% CO2. Cells were then fixed by adding 100 μl per well of 1% PFA directly onto the overlay at room temperature for 40 min. Cells were washed twice with PBS, permeabilized (in 1 × PBS, 0.1% saponin, and 0.1% BSA) for 20 min, and incubated with antibodies specific for WNV (humanized E16 (ref. 33)) or JEV (mouse E18 (ref. 34)) E glycoprotein for 1 h at room temperature. After being rinsed twice, cells were incubated with species-specific HRP-conjugated secondary antibodies (Sigma). After further washing, foci were developed by incubating in 50 μl per well of TrueBlue peroxidase substrate (KPL) for 10 min at room temperature, after which time cells were washed twice in water. Well images were captured using Immuno Capture software (Cell Technology Ltd), and foci counted using BioSpot software (Cell Technology Ltd).
Insect cell and fly infections
dsRNAs were generated as described37. To silence genes using RNAi, insect cells were passaged into serum-free medium containing dsRNAs targeting the indicated genes. Cells were serum-starved for 1 h, after which complete medium was added and cells were incubated for 3 days. Cells were infected with WNV (Kunjin strain) at an MOI of 4 or DENV-2 (NGC strain) at an MOI of 1 for 30 h and then processed for microscopy with automated image analysis as described38. Control (hs>+) or Spcs3-depleted (hs>Spase22-23 IR (Bloomington)) 4–7-day-old female flies were heat shocked (37 °C) for 1 h for three consecutive days to deplete the gene of interest and challenged with WNV (Kunjin) (5 PFU). At day 7 after infection, pools of 10 flies were crushed and titred by plaque assay. Three independent experiments were performed. Heterozygous flies (Spase12(EY10774)) were outcrossed to wild-type flies and either wild-type or Spase12 heterozygous sibling controls were challenged with WNV (Kunjin) for 7 days and groups of 5 flies were titred.
siRNA treatments in human cells
Human U2OS cells were transfected with siRNAs against control, SPCS1, SPCS2, or SPCS3 genes for three days, infected with WNV (Kunjin) or DENV-2 (MOI 1) for 18 h, and then processed for microscopy with automated image analysis as described38.
Replicon transfection and analysis
Two types of replicons were used.
SP6-generated YFV replicons. The wild-type and NS5 polymerase mutant (GDD→GVD) YFV replicons (YF-FFLuc2A, wild-type and GVD) have been published previously12 and were a gift from R. Kuhn. Capped replicon RNA was generated using SP6 polymerase with an mMESSAGE mMACHINE kit according to the manufacturer’s instructions (Thermo Fisher Scientific). RNA was purified using an RNEasy kit (Qiagen) and 2 μg was transfected into control or SPCS1−/− Huh7.5 cells using Lipofectamine 3000 according to the manufacturer’s instructions (Thermo Fisher Scientific). At specified times, cells were collected, lysed, and processed for firefly luciferase activity using a commercial kit (Promega). Cleared lysates were tested for Fluc activity using the Dual-Luciferase Reporter Assay System (Promega) and the protein concentration was quantified using a BCA assay kit (ThermoFisher). Fluc activity (relative light units, RLU) was normalized by subtracting background luminescence of transfected cells collected at the time of transfection, then the adjusted RLU was divided by the total protein content (in μg) to yield RLU per μg protein.
cDNA launched WNV replicons. The construction of wild-type and NS5 polymerase mutant (GDD→GVD) WNV replicons (lineage I, strain New York 1999) was based on a previously described cDNA launched molecular clone system39. The backbone of this strategy, a plasmid containing a truncated WNV genome under the control of a CMV promoter (pWNV-backbone), was designed to be complemented via ligation of a structural gene DNA fragment; transfection of pWNV-backbone alone does not result in production of a self-replicating RNA molecule. Using overlap extension PCR and unique restriction endonuclease sites, pWNV-backbone was modified by the introduction of a fragment downstream of the CMV promoter encoding [5′UTR-cyclization sequence of capsid-FMDV2a protease-signal sequence of E-NS1] to complement the NS2→NS5-3′UTR already present in the pWNV-backbone plasmid, generating the replicon plasmid pWNVI-rep. The reporter gene GFP was then cloned upstream of the FMDV2a protease sequence via a unique MluI site to generate pWNVI-rep–GFP. The construction and organization of this WNV lineage I replicon is analogous to a previously described lineage II WNV replicon (pWNVIIrep–GFP)40. Finally, QuikChange mutagenesis (Agilent Technologies) was used to delete the enhancer portion of the CMV immediate early enhancer/promoter, generating pWNVI-minCMV-rep–GFP, and to generate the GDD→GVD NS5 polymerase variant. Although the CMV enhancer–promoter combination commonly found in cloning vectors results in robust and constitutive expression, inclusion of only the minimal CMV promoter (no enhancer) results in low-level expression41. As such, direct transfection of pWNVI-minCMV-rep–GFP results in a dim GFP signal, which reflects translation of the RNA generated by DNA-dependent RNA translation. RNA polymerase-dependent replication of the wild-type (but not GVD mutant) replicon results in higher production of GFP over time. The eGFP is bracketed by the FMDV2a autocleavage site, and does not rely on host or viral proteases for processing. Wild-type and NS5 GVD variants of pWNVI-minCMV-rep–GFP (200 ng) were transfected into 104 control or gene-edited 293T cells (96-well plates) using Lipofectamine 2000. At various times after transfection, cells were collected, cooled to 4 °C, stained sequentially with a biotinylated anti-9NS1 (ref. 42) (or biotin anti-CHIKV negative control monoclonal antibodies) and Alexa 647-conjugated streptavidin. In some samples, cells were fixed with 4% PFA in PBS (10 min, room temperature) and permeabilized with 0.1% (w/v) saponin. Cells were processed for two-colour flow cytometry using a MACs Quant Analyzer 10 (Miltenyi Biotec).
Plasmid transfections
293T gene-edited cells were transfected with the following genes that were derived from a WNV infectious cDNA clone43 and then cloned into a pHLsec backbone (gift from D. Fremont): V5–C–prM–E, prM, prM–Flag (3 × Flag), E, prM–E, prM–E–NS1, E–NS1, NS1, NS1–NS2A–Flag (includes full-length NS1 and 231 amino acids of NS2A fused to a C-terminal 3 × Flag), and 2K–NS4B–haemagglutinin tag (HA). These plasmids were obtained from colleagues (e.g., 2K-NS4B-HA44, gift from A. Garcia-Sastre) or in some cases engineered to contain either native WNV signal sequences (C–prM, 18 amino acids beyond the C terminus of C; prM-E, 17 C-terminal amino acids of prM; E-NS1, 24 C-terminal amino acids of E) or the signal sequence of mouse Kb class I MHC (N-terminal 21 amino acids). Plasmids were transfected into gene-edited 293T cells using FugeneHD reagent (Promega) according to the manufacturer’s instructions. Supernatants containing prM-E subviral particles (SVPs) were collected 24 h after transfection, filtered through a 0.2-μm filter, and stored aliquotted at −80 °C. For the capture ELISA, Nunc MaxiSorp polystyrene 96-well plates were coated overnight at 4 °C with mouse E60 monoclonal antibodies34 (5 μg ml−1) in a pH 9.3 carbonate buffer. Plates were washed three times in enzyme-linked immunosorbent assay (ELISA) wash buffer (PBS with 0.02% Tween 20) and blocked for 1 h at 37 °C with ELISA block buffer (PBS, 2% bovine serum albumin, and 0.02% Tween 20). Supernatants from prM-E plasmid transfected cells were captured on plates coated with E60 for 90 min at room temperature. Subsequently, plates were rinsed five times in wash buffer and then incubated with humanized anti-WNV E16 (1 μg ml−1 in block buffer) for 1 h at room temperature. Plates were washed five times and then incubated with pre-absorbed biotinylated goat anti-human IgG antibody (1 μg ml−1; Jackson Laboratories) for 1 h at room temperature in blocking buffer. Plates were washed again five times and then sequentially incubated with 2 μg ml−1 of horseradish peroxidase-conjugated streptavidin (Vector Laboratories) and tetramethylbenzidine substrate (Dako). The reaction was stopped with the addition of 2 N H2SO4 to the medium, and emission (450 nm) was read using an iMark microplate reader (Bio-Rad).
Western blotting
For virus infected samples, cells were infected with WNV (MOI 200–1,000, 24 h), JEV (MOI 150, 45 h), CHIKV (MOI 5, 12 h), SINV (MOI 5, 16 h), RVFV (MOI 2.5, 16 h), or HCV (MOI 5, 48 or 72 h). Cells (106) were lysed directly in 30 μl RIPA buffer (Cell Signaling) with 0.1% SDS and a cocktail of protease inhibitors (Sigma-Aldrich). Samples were prepared in LDS buffer (Life Technologies) under non-reducing or reducing (dithiothreitol) conditions. After heating (70 °C, 10 min), samples were electrophoresed using 7% Tris-Acetate or 4–12%, 10% or 12% Bis-Tris gels (Life Technologies) and proteins were transferred to PVDF membranes using an iBlot2 Dry Blotting System (Life Technologies). Membranes were blocked with 5% non-fat dry powdered milk and probed with antibodies against SPCS1 (11847-1-AP, Proteintech), SPCS2 (14872-1-AP, Proteintech), SPCS3 (ab91222, Abcam), SEC11A (14753-1-AP, Proteintech), SEC11C (HPA026816, Sigma) and SEC61B (ab15576, Abcam). For studies with prM–E, prM, E, NS1, NS1–2A–Flag, or 2K–NS4B–Flag-transfected or virus-infected cells, membranes were probed with anti-E (human E16; mouse CHK-4835; mouse anti-JEV, oligoclonal pool), anti-NS1 (mouse 8-NS1), anti-NS3 (W1018-54, USBio), anti-NS4B (rabbit polyclonal antibody45, gift from W. I. Lipkin) anti-prM (human CR429316 or rabbit WNV-M (IMG-5099A, IMGENEX)), anti-Flag (F1804, Sigma), and the relevant secondary antibodies. For validation of the secretome experiments, supernatants were electrophoresed and PVDF membranes were probed with anti-CXCL16 (ab101404, Abcam), anti-SFRP1 (ab126613, Abcam), anti-RNASET2 (ab169655, Abcam), anti-LGALS3BP(ab81489, Abcam),anti-SLITL2 (ab173758, Abcam), anti-PEDF (ab157207, Abcam), anti-NPC2 (19888-1-AP, Proteintech), anti-CREG1 (12220-1-AP, Proteintech), and the relevant secondary antibodies. Western blots were developed using SuperSignal West Pico Chemiluminescent Substrate or SuperSignal West Femto Maximum Sensitivity Substrate (Life Technologies).
Metabolic labelling, pulse-chase, and immunoprecipitation experiments
Pulse-labelling and pulse-chase experiments were performed as described previously46. After starvation in methionine/cysteine-free DMEM for 30 min, 293T cells were labelled metabolically with 300 or 500 μCi ml−1 [35S]-methionine/cysteine (PerkinElmer Life Sciences) at 37 °C for 3 or 40 min. Cells then were washed three times in PBS and immediately lysed or incubated in DMEM supplemented with non-radiolabelled cysteine (500 μg ml−1) and methionine (100 μg ml−1). Cells lysis was performed in 400 μl of 50 mM Tris-HCl, pH 7.4, 150 mM NaCl, 1 mM PMSF, 1 mM EDTA, 5 μg ml−1 aprotinin, 5 μg ml−1 leupeptin, 1% Triton X-100, 1% sodium deoxycholate, 0.1% SDS. After preclearing with an irrelevant human monoclonal antibody protein A-agarose (Thermo Fisher Scientific) complex, lysates were incubated for 1 h at 4 °C with humanized monoclonal E16 and E60 monoclonal antibodies or anti-Flag and then with protein A-agarose for 2 h. The immunoprecipitates were washed seven times in 50 mM Tris-HCl, pH 7.4, 150 mm NaCl, 1 mM PMSF, 1 mM EDTA, 5 μg ml−1 aprotinin, 5 μg ml−1 leupeptin, 1% Triton X-100, 1% sodium deoxycholate, and 0.1% SDS, and then analysed by SDS–PAGE under reducing conditions, followed by fluorography. Some immunoprecipitates were incubated with 20 mU endoglycosidase H or PNGase F (New England BioLabs) for 1 h at 37 °C before SDS–PAGE and fluorography.
293T cell viability assay
A Vybrant MTT cell viability assay (Life Technologies) was used according to the manufacturer’s instructions. Briefly, 10 μl of 12 mM MTT (4,5-dimethylthiazol-2-yl-2-5-diphenyltetrazolium bromide) was added to 105 293T cells (different gene-edited lines, with or without WNV infection) in 100 μl phenol-red free medium. Cells were incubated for 4 h at 37 °C, at which time medium was removed and formazan crystals solubilized in 100 μl of DMSO were added for 10 min at 37 °C. Liquid was analysed for absorbance at 540 nm using a Synergy H1 Hybrid Plate Reader (Biotek).
Flow and mass cytometry analysis of Jurkat T cells
The antibodies and conjugates used are listed in Supplementary Table 6. For flow cytometry studies, wild-type and SPCS1 gene-edited Jurkat T cells were incubated with fluorophore-conjugated monoclonal antibodies for 30 min at 4 °C and then washed three times in PBS containing 5% FBS. Cells were immediately processed on an LSRII flow cytometer and data were analysed using FlowJo 10.0.7. For mass cytometry studies, wild-type and SPCS1 gene-edited Jurkat T cells were incubated with monoclonal antibodies conjugated with transition element isotopes and analysed on a CyTOF 2 mass cytometer (Fluidigm DVS Sciences). Data were analysed using Cytobank (http://wustl.cytobank.org) and FlowJo 10.0.7.
Secretome analysis of SPCS1−/− 293T cells
Wild-type and SPCS1−/− 293T cells were cultured in poly-d-lysine treated flasks in FreeStyle 293 Expression Medium (ThermoFisher) supplemented with 10% FBS. At 90% confluence, cells were washed four times with pre-warmed PBS, then twice with pre-warmed FreeStyle 293 Expression Medium, and maintained in FreeStyle 293 Expression Medium without FBS for 48 h. Supernatants were collected and clarified by centrifugation at 1,000g for 5 min, and then 10,000g for 20 min at 4 °C. Samples were concentrated with Amicon Ultra-15 Centrifugal Filter Units (Millipore) at 5,000g for 1 h in the presence of 1 × protease inhibitors (S8830, Sigma). The concentrates were collected and stored at −80 °C. After thawing on ice, the samples were exchanged twice in digestion buffer (Tris, 0.1 M, pH 8.5 containing 8 M urea) by centrifugation (~4,000g, 2 h) in Amicon Ultracel 3K units to a volume of ~100 μl. The solubilized samples were reduced with 2 mM DTT (ThermoScientific) for 30 min at 37 °C followed by alkylation at room temperature for 30 min with 7 mM iodoacetamide (Sigma) in the dark. The alkylated samples were treated with 7 mM DTT for 15 min at room temperature. After dilution, the samples were digested with LysC (1 μg) (Sigma) overnight at 37 °C with agitation (ThermoMixer). After dilution of the samples to 1.5 M urea with Tris buffer, trypsin was added (5 μg) (Sigma) was added and the incubation was continued overnight at 37 °C with mixing. The digested samples were acidified with to a concentration of 1% tri-flouro acetic acid (TFA). The peptides were desalted with a SepPak (50 mg) with 0.1%TFA/70% acetonitrile in an elution volume (2 ml). The lyophilized peptides were quantified with a fluorescent assay (Thermo Fisher) and 2 μg was labelled with TMT-6 reagents according to the vendor. The labelled peptides were desalted and the samples were transferred to PCR tubes (0.5 ml) and positioned in 96-well holders for robotic solid phase extraction (SPE). Each digest was extracted sequentially with one C4 tip (Glygen BIOMEK NT3C04) and one porous graphite carbon micro-tip (Glygen BIOMEK NT3CAR) with the following auto-pipetting steps: (i) wet tips with AcN/FA (60%/1%) (10 × 25 μl); (ii) equilibrate tips with AcN/FA (1%/1%) (10 × 25 μl); (iii) extract peptides with repetitive aspirations of the digest (50 × 25 μl); (iv) wash loaded tips with AcN/FA (1%/1%) (10 × 25 μl); and (v) elute peptides with AcN/FA (60%/1%) (5 × 65 μl). The SPE eluents were pooled and dried in a SpeedVac centrifuge and transferred to an autosampler vial for LC–MS analysis.
The remainder of the peptides were dissolved in the binding buffer (100 mM Tris, pH 7.8 containing NaCl (0.5 M), MnCl2 (1 mM) and CaCl2 (1 mM). The dried lectins (Con-A and WGA) Sigma were dissolved in binding buffer (4 mg ml−1). The rCA120 (10 mg ml−1), Con-A and WGA were added to the peptide solution (36 μl and 10 μl, respectively). After incubation at room temperature, the mixture was transferred to a YM-10 Microcon filter unit. After centrifugation (14,000g) for 10 min and washing with binding buffer (100 μl), the filter unit was transferred to another tube. The peptides were released with the addition of PNGase (10 units) in 100 μl of ammonium bicarbonate buffer (50 mM) after incubation at 37 °C for 1.5 h. The enzyme addition and incubation was repeated and the peptides recovered with one wash of PNGase buffer. The peptides were acidified to 5% formic acid and desalted, labelled with TMT-6, and prepared for LC–MS as described above for the total pool of peptides.
LC–MS analysis
LC–ESI/MS/MS analysis was conducted with a Q-Exactive Plus mass spectrometer coupled to an EASY-nanoLC 1000 system (Thermo-Fisher). For each Hp-RP fraction, 2 μl of sample was loaded onto a 75 μm i.d. × 25 cm Acclaim PepMap 100 RP column (Thermo-Fisher Scientific). Peptide separations were started with 95% mobile phase A (0.1% FA) for 5 min and increased to 30% B (100% ACN, 0.1% FA) over 180 min, followed by a 25-min gradient to 45% B, a 5-min gradient to 95% B and wash at 90% B for 7 min, with a flow rate of 300 nl min−1. Full-scan mass spectra were acquired by the Orbitrap mass analyser in the mass-to-charge ratio (m/z) of 375–1,400 and with a mass resolving power set to 70,000. Fifteen data-dependent high-energy collisional dissociations were performed with a mass resolving power set to 35,000, a fixed first m/z of 100, an isolation width of 0.7 m/z, and the normalized collision energy (NCE) setting of 32. The maximum injection time was 50 ms for parent-ion analysis and 105 ms for product-ion analysis. Target ions already selected for MS/MS were excluded dynamically for 30 s. An automatic gain control target value of 3 × 106 ions was used for full MS scans and 105 ions for MS/MS scans. Peptide ions with charge states of one or greater than six were excluded from MS/MS interrogation.
Protein identification and quantification with TMT
All raw data were processed using Proteome Discoverer (version 2.1.0.81, Thermo-Fischer Scientific). MS/MS spectra were searched with SequestHT engine against the human UniRef database (69,021 entries; version 2014_05), assuming the digestion enzyme was trypsin with a maximum of 2 missed cleavage allowed. The searches were performed with a fragment ion mass tolerance of 0.02 Da and a parent ion tolerance of 20 ppm. Deamidation of asparagine and glutamine, acetylation and TMT 6-plex derivatization of N termini and oxidation of methionine were specified in Proteome Discoverer as variable modifications. Iodoacetamide derivatization of cysteine and TMT 6-plex derivatization of lysine were specified as fixed modifications. Peptide spectral matches (PSM) were validated using percolator based on q-values at a 1% FDR47. Peptides were filtered to 1% FDR and grouped into proteins at 1% FDR as specified in Proteome Discoverer. The intensities of TMT reporter ions were determined with Proteome Discoverer at a mass tolerance of 0.01 Da and used for peptide quantifications. The median values of peptide intensities that can be assigned to a same protein was used to represent protein intensities. Peptide identifications that can be assigned to more than one protein were removed from protein quantification
Proteomic Data Analysis
Normalization in protein ratios was applied in that the median ratios are log2 0. Data analysis was performed with the free software environment for statistical computing and graphics, R (http://www.R-project.org). Gene ontology analysis was carried out using the Database for Annotation, Visualization and Integrated Discovery (DAVID)48,49. Data from duplicated LC/MS/MS analysis were first averaged and protein abundance ratios were log2-transformed before statistical analysis. A one-way ANOVA with Benjamini-Hochberg correction was performed to assess the statistical significance in protein abundance changes between wild type and SPCS1−/− cells.
Statistical analysis
No statistical methods were used to predetermine sample size. The experiments were not randomized and the investigators were not blinded to allocation during experiments and outcome assessment. Statistical significance was assigned when P values were <0.05 using GraphPad Prism Version 5.04. Viral antigen staining after expression of sgRNA was analysed using a one-way ANOVA adjusting for repeated measures with a Dunnett’s multiple comparison test or with a Mann–Whitney test depending on the number of comparison groups. Analysis of levels of E protein in the supernatant from CRISPR-Cas9 gene edited cells was analysed by a one-way ANOVA. Analysis of siRNA in insect and human cells was performed using a Student’s t-test or ANOVA.
Change history
15 July 2016
The HTML was replaced with the final print version of the manuscript (instead of the near-final version).
References
Diamond, M. S. & Pierson, T. C. Molecular insight into Dengue virus pathogenesis and its implications for disease control. Cell 162, 488–492 (2015)
Weaver, S. C. et al. Zika virus: History, emergence, biology, and prospects for control. Antiviral Res. 130, 69–80 (2016)
Cong, L. et al. Multiplex genome engineering using CRISPR/Cas systems. Science 339, 819–823 (2013)
Jinek, M. et al. RNA-programmed genome editing in human cells. eLife 2, e00471 (2013)
Shalem, O. et al. Genome-scale CRISPR-Cas9 knockout screening in human cells. Science 343, 84–87 (2014)
Koike-Yusa, H., Li, Y., Tan, E. P., Velasco-Herrera, Mdel. C. & Yusa, K. Genome-wide recessive genetic screening in mammalian cells with a lentiviral CRISPR-guide RNA library. Nat. Biotechnol. 32, 267–273 (2014)
Wang, T., Wei, J. J., Sabatini, D. M. & Lander, E. S. Genetic screens in human cells using the CRISPR-Cas9 system. Science 343, 80–84 (2014)
Li, W. et al. MAGeCK enables robust identification of essential genes from genome-scale CRISPR/Cas9 knockout screens. Genome Biol. 15, 554 (2014)
Evans, E. A., Gilmore, R. & Blobel, G. Purification of microsomal signal peptidase as a complex. Proc. Natl Acad. Sci. USA 83, 581–585 (1986)
Meyer, H. A. & Hartmann, E. The yeast SPC22/23 homolog Spc3p is essential for signal peptidase activity. J. Biol. Chem. 272, 13159–13164 (1997)
Khromykh, A. A., Kenney, M. T. & Westaway, E. G. trans-Complementation of flavivirus RNA polymerase gene NS5 by using Kunjin virus replicon-expressing BHK cells. J. Virol. 72, 7270–7279 (1998)
Jones, C. T., Patkar, C. G. & Kuhn, R. J. Construction and applications of yellow fever virus replicons. Virology 331, 247–259 (2005)
Lindenbach, B. D., Murray, C. L., Thiel, H. J. & Rice, C. M. in Fields Virology Vol. 1 (eds Knipe, D. M. & Howley, P. M. ) 712–746 (Lippincott Williams & Wilkins, 2013)
Chambers, T. J., Grakoui, A. & Rice, C. M. Processing of the yellow fever virus nonstructural polyprotein: a catalytically active NS3 proteinase domain and NS2B are required for cleavages at dibasic sites. J. Virol. 65, 6042–6050 (1991)
Falgout, B., Pethel, M., Zhang, Y. M. & Lai, C. J. Both nonstructural proteins NS2B and NS3 are required for the proteolytic processing of dengue virus nonstructural proteins. J. Virol. 65, 2467–2475 (1991)
Throsby, M. et al. Isolation and characterization of human monoclonal antibodies from individuals infected with West Nile Virus. J. Virol. 80, 6982–6992 (2006)
Barth, B. U., Wahlberg, J. M. & Garoff, H. The oligomerization reaction of the Semliki Forest virus membrane protein subunits. J. Cell Biol. 128, 283–291 (1995)
Löber, C., Anheier, B., Lindow, S., Klenk, H. D. & Feldmann, H. The Hantaan virus glycoprotein precursor is cleaved at the conserved pentapeptide WAASA. Virology 289, 224–229 (2001)
Schalich, J. et al. Recombinant subviral particles from tick-borne encephalitis virus are fusogenic and provide a model system for studying flavivirus envelope glycoprotein functions. J. Virol. 70, 4549–4557 (1996)
Lorenz, I. C., Allison, S. L., Heinz, F. X. & Helenius, A. Folding and dimerization of tick-borne encephalitis virus envelope proteins prM and E in the endoplasmic reticulum. J. Virol. 76, 5480–5491 (2002)
Hanna, S. L. et al. N-linked glycosylation of west nile virus envelope proteins influences particle assembly and infectivity. J. Virol. 79, 13262–13274 (2005)
Gowen, B. G. et al. A forward genetic screen reveals novel independent regulators of ULBP1, an activating ligand for natural killer cells. eLife 4, e08474 (2015)
Suzuki, R. et al. Signal peptidase complex subunit 1 participates in the assembly of hepatitis C virus through an interaction with E2 and NS2. PLOS Pathog. 9, e1003589 (2013)
Ma, H. et al. A CRISPR-based screen identifies genes essential for West-Nile-Virus-induced cell death. Cell Rep. 12, 673–683 (2015)
Krishnan, M. N. et al. RNA interference screen for human genes associated with West Nile virus infection. Nature 455, 242–245 (2008)
Yasunaga, A. et al. Genome-wide RNAi screen identifies broadly-acting host factors that inhibit arbovirus infection. PLOS Pathog. 10, e1003914 (2014)
Sessions, O. M. et al. Discovery of insect and human dengue virus host factors. Nature 458, 1047–1050 (2009)
Rose, P. P. et al. Natural resistance-associated macrophage protein is a cellular receptor for sindbis virus in both insect and mammalian hosts. Cell Host Microbe 10, 97–104 (2011)
Brien, J. D., Lazear, H. M. & Diamond, M. S. Propagation, quantification, detection, and storage of West Nile virus. Curr. Protoc. Microbiol. 31, 15D 13 11–15D 13 18 (2013)
Jiang, J. & Luo, G. Cell culture-adaptive mutations promote viral protein-protein interactions and morphogenesis of infectious hepatitis C virus. J. Virol. 86, 8987–8997 (2012)
Sabo, M. C. et al. Neutralizing monoclonal antibodies against Hepatitis C Virus E2 protein bind discontinuous epitopes and inhibit infection at a postattachment step. J. Virol. 85, 7005–7019 (2011)
Sanjana, N. E., Shalem, O. & Zhang, F. Improved vectors and genome-wide libraries for CRISPR screening. Nat. Meth. 11, 783–784 (2014)
Oliphant, T. et al. Development of a humanized monoclonal antibody with therapeutic potential against West Nile virus. Nat. Med. 11, 522–530 (2005)
Oliphant, T. et al. Antibody recognition and neutralization determinants on domains I and II of West Nile Virus envelope protein. J. Virol. 80, 12149–12159 (2006)
Pal, P. et al. Development of a highly protective combination monoclonal antibody therapy against Chikungunya virus. PLoS Pathog. 9, e1003312 (2013)
Fuchs, A., Pinto, A. K., Schwaeble, W. J. & Diamond, M. S. The lectin pathway of complement activation contributes to protection from West Nile virus infection. Virology 412, 101–109 (2011)
Boutros, M. et al. Genome-wide RNAi analysis of growth and viability in Drosophila cells. Science 303, 832–835 (2004)
Hackett, B. A. et al. RNASEK is required for internalization of diverse acid-dependent viruses. Proc. Natl Acad. Sci. USA 112, 7797–7802 (2015)
Lin, T. Y. et al. A novel approach for the rapid mutagenesis and directed evolution of the structural genes of west nile virus. J. Virol. 86, 3501–3512 (2012)
Pierson, T. C. et al. A rapid and quantitative assay for measuring antibody-mediated neutralization of West Nile virus infection. Virology 346, 53–65 (2006)
Mishin, V. P., Cominelli, F. & Yamshchikov, V. F. A ‘minimal’ approach in design of flavivirus infectious DNA. Virus Res. 81, 113–123 (2001)
Chung, K. M. et al. Antibodies against West Nile Virus nonstructural protein NS1 prevent lethal infection through Fc gamma receptor-dependent and -independent mechanisms. J. Virol. 80, 1340–1351 (2006)
Beasley, D. W. et al. Envelope protein glycosylation status influences mouse neuroinvasion phenotype of genetic lineage 1 West Nile virus strains. J. Virol. 79, 8339–8347 (2005)
Muñoz-Jordán, J. L. et al. Inhibition of alpha/beta interferon signaling by the NS4B protein of flaviviruses. J. Virol. 79, 8004–8013 (2005)
Medigeshi, G. R. et al. West Nile virus infection activates the unfolded protein response, leading to CHOP induction and apoptosis. J. Virol. 81, 10849–10860 (2007)
Miner, J. J. et al. Cytoplasmic domain of P-selectin glycoprotein ligand-1 facilitates dimerization and export from the endoplasmic reticulum. J. Biol. Chem. 286, 9577–9586 (2011)
Käll, L., Canterbury, J. D., Weston, J., Noble, W. S. & MacCoss, M. J. Semi-supervised learning for peptide identification from shotgun proteomics datasets. Nat. Methods 4, 923–925 (2007)
Huang, W., Sherman, B. T. & Lempicki, R. A. Systematic and integrative analysis of large gene lists using DAVID bioinformatics resources. Nat. Protocols 4, 44–57 (2009)
Huang, W., Sherman, B. T. & Lempicki, R. A. Bioinformatics enrichment tools: paths toward the comprehensive functional analysis of large gene lists. Nucleic Acids Res. 37, 1–13 (2009)
Acknowledgements
This work was supported by NIH grants U19 AI083019 (M.S.D.), U19 AI106772 (M.S.D.), R01 AI104972 (M.S.D.), and T32 AI007163 (E.F.) and by the Washington University Institute of Clinical and Translational Sciences (UL1 TR000448 from the National Center for Advancing Translational Sciences and P41 GM103422-35 from the National Institute of General Medical Sciences). T.C.P and K.A.D are supported by the intramural program of NIAID. We thank R. Kuhn, A. Garcia-Sastre, H. Zhao, D. Fremont, X. Wang, and R. Townsend for reagents, experimental advice, and data analysis; P. Erdmann-Gilmore, R. Connors, Y. Mi, and H. Lin for expert technical assistance; and X. de Lamballerie and the European Virus Archive goes Global (EVAg) for consenting to the use of H/PF/2013 ZIKV strain for this study under a material transfer agreement with the EVAg parter, Aix-Marseille Université.
Author information
Authors and Affiliations
Contributions
R.Z. performed the primary CRISPR/Cas9 screen and viral protein transfection experiments. Validation studies in cells with different viruses were performed by R.Z., P.Z., M.J.G, A.Z., E.F., and M.S.D. J.J.M. performed the pulse-chase experiments and mass cytometry. J.P.W. performed studies with replicons. K.R. and H.R performed the insect cell, insect, and siRNA experiments. Q.Z. performed the secretome analysis and mass spectrometry. R.Z., S.C., T.C.P., and M.S.D. designed the experiments. K.A.D. and T.C.P. provided key reagents. Q.Z. performed data analysis. S.C. and M.S.D. wrote the initial draft of the manuscript, with the other authors contributing to editing into the final form.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
Reviewer Information Nature thanks W. Wei and the other anonymous reviewer(s) for their contribution to the peer review of this work.
Extended data figures and tables
Extended Data Figure 1 Results of CRISPR-Cas9 screen.
a, Scheme of gene-editing screen. b, Analysis of cell viability of gene-edited cells. WNV-infected CRISPR-Ca9 edited bulk cells were evaluated for cell viability using a metabolic MTT assay 24 h after infection. The results were pooled from several independent experiments performed in duplicate and data were compared to cells edited with a control sgRNA. None of the differences were statistically significant compared to the control. c, Western blotting confirms the efficiency of gene editing of SEC61B, SPCS1, and SPCS3. β-actin is included as a loading control. d, Effect of gene editing on infection by other RNA viruses. sgRNA-edited bulk selected cell populations were infected with alphaviruses (SINV or CHIKV), a bunyavirus (LACV) or a rhabdovirus (VSV). Cells were analysed for intracellular viral antigen staining by flow cytometry using virus-specific monoclonal antibodies. The data are representative of two independent experiments and are expressed as relative infection (viral antigen expression) compared to the sgRNA control. d–f, Trans-complementation of sgRNA gene-edited cells with Flag-tagged genes. d, Individual sgRNA bulk gene-edited cell lines were trans-complemented with cDNA expressing C-terminal Flag-tagged versions of their respective genes and GFP or an empty vector control and GFP. Transfected cells were analysed by flow cytometry for expression of the Flag-tag in the GFP+ cells. The data are representative of two independent experiments. e, Western blotting of SPCS1 and SPCS3 trans-complemented genes after incubation with an anti-Flag antibody. f, Individual sgRNA cell lines were trans-complemented with cDNA expressing C-terminal Flag-tagged versions of their respective genes and GFP or an empty vector control and GFP. Transfected cells were infected with WNV (MOI 5) and 12 h later cells were stained for intracellular E antigen and processed by flow cytometry. The data are representative of three independent experiments performed in triplicate and reflect the percentage of WNV-infected cells in the fraction that expressed GFP. The indicated comparisons were statistically significant (****P < 0.0001), as determined by the Mann–Whitney test. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 2 Gene silencing of SPCS genes in human U2OS cells.
Human U2OS cells were transfected with either control or SPCS1, SPCS2, or SPCS3 siRNAs and infected with WNV (Kunjin) or DENV-2 (MOI 1) for 18 h. Left, the percentage of infected cells was determined by automated fluorescence microscopy. The data are expressed as the mean normalized value ± s.d. **P < 0.01; ***P < 0.0001 compared to control siRNA by ANOVA with a multiple comparison correction. The data are pooled from three independent experiments assayed in quadruplicate. No reduction in infection of CHIKV or SINV was observed after SPCS gene silencing (data not shown). Right, western blotting of SPCS proteins in gene-silenced U2OS cells. Representative results are shown and tubulin is included as a loading control. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 3 Viral infection in SPCS1−/− cells.
a–c, Alphaviruses replicate and are processed efficiently in 293T cells in the absence of expression of SPCS1. a, SINV infection in control and SPCS1−/− clonal cells. Cells were infected (MOI 0.01) and supernatants were collected and analysed by FFA. The results are the average of two independent experiments performed in triplicate. b, Control and SPCS1−/− gene-edited 293T cells were infected with SINV. At the indicated time, lysates were prepared, electrophoresed and western blotted with anti-SINV E2 ascites fluid (ATCC VR-1248AF). c, Control or SPCS1−/− 293T cells were infected with CHIKV (MOI 5). After 8 h, cells were labelled for 30 min with [35S]cysteine/methionine. Excess cold cysteine/methionine was added for indicated chase times (0, 1 or 4 h). An uninfected control established the specificity of the immunoprecipitation. After 35S labelling, lysates were prepared and immunoprecipitated with anti-E2 monoclonal antibodies (CHK-48). Immunoprecipitates were left untreated (blank) or treated with Endo H (E) or PNGase F (P) for 1 h at 37 °C before SDS–PAGE and fluorography. d, Sequencing of SPCS1 alleles in gene-edited 293T and Huh7 cell clones after puromycin selection and limiting dilution cloning. The sgRNA targeting site and the ‘PAM’ sequences are highlighted at the top of the wild-type gene, and the sequence of edited alleles are indicated. e, Western blotting of bulk-selected or clonal (clone 7) Huh7.5 cells (control and SPCS1 sgRNA selected) for expression of SPCS1 (~12 kDa). f, WNV, HCV, ZIKV, and JEV (Bennett strain) infection in control and SPCS1-deficient Huh7.5 cells. Cells were infected at an MOI of 0.01 (WNV, ZIKV, JEV) or 1 (HCV) and supernatants were collected and analysed by FFA. The results are the average of two independent experiments performed in triplicate. g, Control or SPCS1−/− Huh7.5 cells were infected at an MOI of 150 for 45 h with a pathogenic JEV isolate (Bennett strain). Lysates were blotted with an anti-JEV E monoclonal antibody. Higher molecular mass bands (Ehi and Emed) that reacted specifically with the anti-E monoclonal antibody are indicated. One representative experiment of two is shown and loading controls (β-actin) are included. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 4 Gene editing of SEC11A and SEC11C do not substantively affect infection of several enveloped viruses.
Top, 293T cells were treated with the indicated sgRNA and isolated in bulk after puromycin drug selection. Western blotting confirmed gene editing of SEC11A (left, 20 kDa) or SEC11C (middle, 22 kDa). No difference in levels or migration pattern of WNV E was observed in SEC11A or SEC11C gene-edited cells (right) after WNV infection at an MOI of 200 for 24 h. Spaces between the western blots indicate cropping to remove lanes that were not relevant to this experiment. Bottom, control or gene-edited 293T cells were infected with viruses and supernatants were harvested after infection for titration. Left, WNV (MOI 0.01, 72 h) or YFV (MOI 1, 72 h); right, SINV (MOI 0.01, 72 h), CHIKV (MOI 0.01, 36 h), VSV (MOI 0.01, 36 h), or RVFV (MOI 1, 72 h). Results are representative of two independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 5 Effect of sgRNA on translation and replication of wild-type and NS5 GVD polymerase mutant WNV replicons.
A cDNA launched WNV replicon (a, wild-type; b, GVD polymerase ‘dead’ mutant) with a minimal CMV promoter (GFP–NS1–NS2A–NS2B–NS3–NS4A–NS4B–NS5) was transfected into the indicated gene-edited 293T cells. At 48 and 72 h after transfection, cells were collected and analysed for GFP expression by flow cytometry. After transfection with the wild-type replicon, WNV replication was lower in STT3A gene-edited cells, as determined by ANOVA with a multiple comparisons correction (P < 0.05 at 48 and 72 h). Data are the average of three independent experiments. Note, the GVD replicon data with sgRNA control (translation only) are provided for comparison in a.
Extended Data Figure 6 Processing of WNV proteins in SPCS1 and SPCS3 gene-edited 293T cells.
a, Normal (left) and over-exposed (right) western blot in SPCS1 and SPCS3 bulk gene-edited 293T cells. The over-exposure is shown to highlight the accumulation of high molecular mass bands that react with anti-E protein antibody. Control, SPCS1 and SPCS3 gene-edited 293T cells were infected with WNV or mock-infected for the indicated times. Lysates were western blotted with an anti-E (human E16) monoclonal antibodies. Under these electrophoresis conditions, natively processed E protein migrates at ~50 to 55 kDa. Higher molecular mass bands (Emed (probably prM–E) and Ehi (probably prM–E–NS1)) that react specifically with the E monoclonal antibody are present only in SPCS1 and SPCS3 gene-edited 293T cells. The data are representative of two independent experiments and a loading control (β-actin) is shown. b, Western blot of SPCS1 and SPCS3 bulk gene-edited 293T cells. Control, SPCS1, and SPCS3 gene-edited 293T cells were infected with WNV or mock-infected for the indicated times. Lysates were western blotted with an anti-prM human monoclonal antibody (CR4293) that recognizes a shared epitope on prM and E. Higher molecular mass bands (prM–Ehi) probably represent uncleaved polyprotein forms and are present only in SPCS1 and SPCS3 gene-edited 293T cells. The data are representative of two independent experiments and a loading control (β-actin) is shown. c, Control or SPCS1−/− cells were infected with WNV or left unmanipulated (−) and 24 h later cell lysates were generated and probed with a polyclonal antibody against NS4B. The results are representative of two independent experiments and loading controls (β-actin) are shown. d, Control or SPCS1−/− cells were transfected with a 2K-NS4B–HA plasmid. One day later, lysates were probed with an anti-HA antibody. Results are representative of two independent experiments and loading controls (β-actin) are shown. Cleaved (NS4B) and uncleaved (2K–NS4B) bands are indicated on the right of the gel. e, Control or SPCS1−/− cells were transfected with NS1, NS1–NS2A–Flag, or control plasmids. One day later, lysates were probed with anti-NS1 (left) or anti-FLAG (right) antibodies. Cleavage of NS1–NS2A–Flag results in expression of the C-terminal Flag tag exclusively with the residual NS2A protein. The results are representative of three independent experiments and loading controls (β-actin) are shown. Note, expression of NS1–NS2A results in two forms of NS1 (NS1 and NS1′) owing to a ribosomal frameshift event that occurs at a heptanucleotide motif near the beginning of the NS2A gene. f, Control or SPCS1−/− cells were infected with WNV or left uninfected and 24 h later cell lysates were generated and probed with a monoclonal antibody against NS3. The results are representative of two independent experiments and loading controls (β-actin) are shown. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 7 Western blotting for HCV E2 in control and SPCS1 gene-edited Huh7.5 cells.
Control or SPCS1 gene-edited cells were infected with HCV (MOI 5; +) or left untreated (−) and 48 or 72 h later cell lysates were generated and probed with a mouse monoclonal antibody against HCV E2 protein. The results are representative of two independent experiments and a normal and over-exposed blot are shown. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 8 Effect of sgRNA on WNV structural protein processing and production.
a–c, The indicated gene-edited 293T cells were transfected with a plasmid encoding WNV prM–E and subjected to western blotting with hE16 (anti-E) (a) or CR4293 (anti-prM-E) (b). Note the shift of the prM–E bands to high molecular mass in bulk gene-edited cells with reduced expression of SPCS1 or SPCS3. The results are representative of three independent experiments and a loading control (β-actin) is shown. c, 293T cells expressing the indicated sgRNA were transfected with a plasmid encoding prM–E. After 24 h, supernatants were collected and SVPs were quantified by capture ELISA. The results are the average of several independent experiments performed in triplicate. The asterisks indicate SVP levels in the supernatant that are statistically different compared to control cells (****P < 0.001, ANOVA with a multiple comparison correction). d, Control or SPCS1−/− clonal 293T cells were transfected with a single C–prM–E plasmid containing an N-terminal V5 tag fused to C (purple box) and native C–prM (green box) and prM–E (blue box) leader sequences. In some experiments, a cDNA launched WNV replicon was co-transfected to facilitate the cleavage of C from prM by the viral NS2B–NS3 protease. Lysates were prepared 24 h later and probed with an anti-V5 (top) or anti-E (bottom) antibody. Note, two separate gels were run for blotting with anti-V5 (C) and anti-E. One representative experiment of two is shown and a loading control (β-actin) for the top (anti-V5) gel is included. e, Control and SPCS1−/− 293T cells were transfected with E or NS1 with or without prM co-transfection. One day after transfection, cells were collected and lysates were western blotted with antibodies against E (left) or NS1 (right). Molecular mass markers and specific proteins are indicated to the left and right of each gel, respectively. The results are representative of three independent experiments. For gel source data, see Supplementary Fig. 1.
Extended Data Figure 9 Expression of immune system antigens on the surface of SPCS1 gene-edited cells.
a, b, Control and SPCS1 gene-edited Jurkat cells were incubated with monoclonal antibodies against the indicated cell surface antigens. After washing, cells were fixed with paraformaldehyde and then processed by flow cytometry (a) or mass cytometry (b). The histograms are as follows: black, isotype control in wild-type cells; blue, control cells; red, SPCS1 gene-edited cells. Results are representative of three independent experiments for flow cytometry and one run on a mass cytometer in triplicate. c, Western blotting of bulk-selected Jurkat cells (control and SPCS1 sgRNA selected) for expression of SPCS1 (~12 kDa). For gel source data, see Supplementary Fig. 1.
Extended Data Figure 10 Secretome analysis in control and SPCS1−/− cell supernatants.
a, Volcano plot from one-way ANOVA for secreted protein abundances between control and SPCS1−/− 293T cells. The areas of dots are proportional to the log2 standard deviation of protein ratios. The vertical dashed lines delimit fold changes ± 1.1 and the horizontal dashed line delimits P value ≤ 0.05. The red dots show secreted proteins using the SP_PIR classification. Values < log2 0 indicate secreted proteins that show reduced expression in SPCS1−/− 293T cells. b, c, Western blotting of supernatants from control and SPCS1−/− 293T cells. b, Proteins (LGALS3BP, RNASET2, and NPC2) identified as downregulated in SPCS1−/− 293T cells by mass spectrometry (see Supplementary Tables 4 and 5). c, Proteins identified as having similar or possibly higher levels in supernatants of SPCS1−/− 293T cells. For gel source data, see Supplementary Fig. 1.
Supplementary information
Supplementary Table 1
This table shows frequency of sgRNA deep sequencing reads. Data was obtained by deep-sequencing sgRNA from uninfected or surviving WNV-infected cells. Three independent libraries were sequenced from each of two biologically independent experiments. Results are sorted by the top fold-enrichment of an individual sgRNA from WNV-infected cells compared to uninfected cells. (XLSX 5651 kb)
Supplementary Table 2
This file contains a list of gene hits and scores after MaGeck analysis. (XLSX 803 kb)
Supplementary Table 3
This file contains sgRNA sequences used for gene validation. (XLSX 9 kb)
Supplementary Table 4
This file contains results of secretome analysis from control and SPCS1-/- 293T cells after tandem mass spectrometry (MALDI-TOF/TOF) and peptide fragmentation spectra analysis. (XLSX 71 kb)
Supplementary Table 5
This file contains a summary of Western blot validation of secretome analysis in cells and supernatants of control and SPCS1-/- 293T cells. (XLSX 33 kb)
Supplementary Table 6
This file contains a list of antibodies used for flow and mass cytometry for staining of control and SPCS1-/- Jurkat cells. (XLSX 35 kb)
Supplementary Figure
This file contains Supplementary Figure 1, gel source data. (PDF 11515 kb)
Rights and permissions
About this article
Cite this article
Zhang, R., Miner, J., Gorman, M. et al. A CRISPR screen defines a signal peptide processing pathway required by flaviviruses. Nature 535, 164–168 (2016). https://doi.org/10.1038/nature18625
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature18625
This article is cited by
-
Cellular glycan modification by B3GAT1 broadly restricts influenza virus infection
Nature Communications (2022)
-
Stabilization and structure determination of integral membrane proteins by termini restraining
Nature Protocols (2022)
-
Surfaceome CRISPR screen identifies OLFML3 as a rhinovirus-inducible IFN antagonist
Genome Biology (2021)
-
A genome-wide CRISPR screen identifies host factors that regulate SARS-CoV-2 entry
Nature Communications (2021)
-
Distributed synthesis of sarcolemmal and sarcoplasmic reticulum membrane proteins in cardiac myocytes
Basic Research in Cardiology (2021)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.