Abstract
Shoot growth depends on meristems, pools of stem cells that are maintained by a negative feedback loop between the CLAVATA pathway and the WUSCHEL homeobox gene1. CLAVATA signalling involves a secreted peptide, CLAVATA3 (CLV3)2, and its perception by cell surface leucine-rich repeat (LRR) receptors, including the CLV1 receptor kinase3 and a LRR receptor-like protein, CLV2 (ref. 4). However, the signalling mechanisms downstream of these receptors are poorly understood, especially for LRR receptor-like proteins, which lack a signalling domain5. Here we show that maize COMPACT PLANT2 (CT2) encodes the predicted α-subunit (Gα) of a heterotrimeric GTP binding protein. Maize ct2 phenotypes resemble Arabidopsis thaliana clavata mutants, and genetic, biochemical and functional assays indicate that CT2/Gα transmits a stem-cell-restrictive signal from a CLAVATA LRR receptor, suggesting a new function for Gα signalling in plants. Heterotrimeric GTP-binding proteins are membrane-associated molecular switches that are commonly activated by ligand binding to an associated seven-pass transmembrane (7TM) G-protein-coupled receptor (GPCR)6. Recent studies have questioned the idea that plant heterotrimeric G proteins interact with canonical GPCRs7, and our findings suggest that single pass transmembrane receptors act as GPCRs in plants, challenging the dogma that GPCRs are exclusively 7TM proteins.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Stahl, Y. & Simon, R. Plant primary meristems: shared functions and regulatory mechanisms. Curr. Opin. Plant Biol. 13, 53–58 (2010)
Fletcher, J. C., Brand, U., Running, M. P., Simon, R. & Meyerowitz, E. M. Signaling of cell fate decisions by CLAVATA3 in Arabidopsis shoot meristems. Science 283, 1911–1914 (1999)
Clark, S. E., Williams, R. W. & Meyerowitz, E. M. The CLAVATA1 gene encodes a putative receptor kinase that controls shoot and floral meristem size in Arabidopsis. Cell 89, 575–585 (1997)
Jeong, S., Trotochaud, A. E. & Clark, S. E. The Arabidopsis CLAVATA2 gene encodes a receptor-like protein required for the stability of the CLAVATA1 receptor-like kinase. Plant Cell 11, 1925–1934 (1999)
Nimchuk, Z. L., Tarr, P. T. & Meyerowitz, E. M. An evolutionarily conserved pseudokinase mediates stem cell production in plants. Plant Cell 23, 851–854 (2011)
Assmann, S. M. G proteins Go green: a plant G protein signaling FAQ sheet. Science 310, 71–73 (2005)
Urano, D. et al. G protein activation without a GEF in the plant kingdom. PLoS Genet. 8, e1002756 (2012)
Jackson, D., Veit, B. & Hake, S. Expression of maize KNOTTED1 related homeobox genes in the shoot apical meristem predicts patterns of morphogenesis in the vegetative shoot. Development 120, 405–413 (1994)
Bommert, P. et al. thick tassel dwarf1 encodes a putative maize ortholog of the Arabidopsis CLAVATA1 leucine-rich repeat receptor-like kinase. Development 132, 1235–1245 (2005)
Taguchi-Shiobara, F., Yuan, Z., Hake, S. & Jackson, D. The fasciated ear2 gene encodes a leucine-rich repeat receptor-like protein that regulates shoot meristem proliferation in maize. Genes Dev. 15, 2755–2766 (2001)
Ashikari, M., Wu, J., Yano, M., Sasaki, T. & Yoshimura, A. Rice gibberellin-insensitive dwarf mutant gene Dwarf 1 encodes the α-subunit of GTP-binding protein. Proc. Natl Acad. Sci. USA 96, 10284–10289 (1999)
van der Hoorn, R. A. et al. Structure-function analysis of Cf-9, a receptor-like protein with extracytoplasmic leucine-rich repeats. Plant Cell 17, 1000–1015 (2005)
Gallavotti, A., Yang, Y., Schmidt, R. J. & Jackson, D. The relationship between auxin transport and maize branching. Plant Physiol. 147, 1913–1923 (2008)
Fiers, M. et al. The 14-amino acid CLV3, CLE19, and CLE40 peptides trigger consumption of the root meristem in Arabidopsis through a CLAVATA2-dependent pathway. Plant Cell 17, 2542–2553 (2005)
Müller, R., Bleckmann, A. & Simon, R. The receptor kinase CORYNE of Arabidopsis transmits the stem cell-limiting signal CLAVATA3 independently of CLAVATA1. Plant Cell 20, 934–946 (2008)
Ullah, H. et al. Modulation of cell proliferation by heterotrimeric G protein in Arabidopsis. Science 292, 2066–2069 (2001)
Bommert, P., Nagasawa, N. S. & Jackson, D. Quantitative variation in maize kernel row number is controlled by the FASCIATED EAR2 locus. Nature Genet. 45, 334–337 (2013)
Brown, P. J. et al. Distinct genetic architectures for male and female inflorescence traits of maize. PLoS Genet. 7, e1002383 (2011)
Katritch, V., Cherezov, V. & Stevens, R. C. Structure-function of the G protein-coupled receptor superfamily. Annu. Rev. Pharmacol. Toxicol. 53, 531–556 (2013)
Pandey, S., Nelson, D. C. & Assmann, S. M. Two novel GPCR-type G proteins are abscisic acid receptors in Arabidopsis. Cell 136, 136–148 (2009)
Perfus-Barbeoch, L., Jones, A. M. & Assmann, S. M. Plant heterotrimeric G protein function: insights from Arabidopsis and rice mutants. Curr. Opin. Plant Biol. 7, 719–731 (2004)
Lease, K. A. et al. A mutant Arabidopsis heterotrimeric G-protein β subunit affects leaf, flower, and fruit development. Plant Cell 13, 2631–2641 (2001)
Shpak, E. D., Berthiaume, C. T., Hill, E. J. & Torii, K. U. Synergistic interaction of three ERECTA-family receptor-like kinases controls Arabidopsis organ growth and flower development by promoting cell proliferation. Development 131, 1491–1501 (2004)
Shiu, S. H. & Bleecker, A. B. Plant receptor-like kinase gene family: diversity, function, and signaling. Sci. STKE 2001, re22 (2001)
Iyer-Pascuzzi, A. S. et al. Imaging and analysis platform for automatic phenotyping and trait ranking of plant root systems. Plant Physiol. 152, 1148–1157 (2010)
Freeling, M. & Walbot, V. The Maize Handbook. (Springer-Verlag, 1994)
Mohanty, A. et al. Advancing cell biology and functional genomics in maize using fluorescent protein-tagged lines. Plant Physiol. 149, 601–605 (2009)
Chen, J. G., Gao, Y. & Jones, A. M. Differential roles of Arabidopsis heterotrimeric G-protein subunits in modulating cell division in roots. Plant Physiol. 141, 887–897 (2006)
Dennehey, B. K., Retersen, W. L., Ford-Santino, C., Pajeau, M. & Armstrong, C. L. Comparison of selective agents for use with the selectable marker gene bar in maize transformation. Plant Cell Tissue Organ Cult. 36, 1–7 (1994)
Harlow, E. & Lane, D. Antibodies: A Laboratory Manual. (Cold Spring Harbor Laboratory Press, 1988)
Larsson, C. & Widell, S. in Aqueous Two-Phase Systems: Methods and Protocols Vol. 11 Methods in Biotechnology, (ed Hatti-Kaul, R. ) 159–166 (Humana Press, 2000)
McDonald, W. H., Ohi, R., Miyamoto, D. T., Mitchison, T. J. & Yates. J. R. III Comparison of three directly coupled HPLC MS/MS strategies for identification of proteins from complex mixtures: single-dimension LC-MS/MS, 2-phase MudPIT, and 3-phase MudPIT. Int. J. Mass Spectrom. 219, 245–251 (2002)
Taylor, P. et al. Automated 2D peptide separation on a 1D nano-LC-MS system. J. Proteome Res. 8, 1610–1616 (2009)
Washburn, M. P., Wolters, D. & Yates, J. R. III. Large-scale analysis of the yeast proteome by multidimensional protein identification technology. Nature Biotechnol. 19, 242–247 (2001)
Acknowledgements
The authors thank T. Mulligan for plant care; T. Rocheford for growing the ct2 targeted EMS screens; F. Albeanu for help with two-photon microscopy; P. Yin for assistance with genotyping; K. Rivera and D. Pappin for performing proteomic experiments; and T. Zadrozny, Q. Wu and W. Boss for assistance and advice with two-phase partitioning. P.B. was supported by DFG grant Bo 3012/1-1, A.G. was supported by the BARD Postdoctoral award no FI-431-2008, B.I.J was supported by Dupont-Pioneer (CSHL-Collaborative agreement), P.B. and D.J. were supported by the National Science Foundation Plant Genome Program, DBI-0604923.
Author information
Authors and Affiliations
Contributions
P.B., B.I.J., A.G. and D.J. designed experiments and analysed data; P.B. performed all experiments, except peptide assays and contributions to imaging, phenotyping and biochemical assays (B.I.J.) and fluorescence imaging (A.G.); and P.B. and D.J. wrote the paper.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Extended data figures and tables
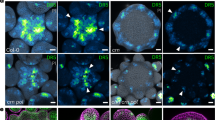
Extended Data Figure 1 Phenotypic analysis of ct2 mutants.
a, Wild-type seedling. b, ct2 seedling showing reduced height. c, Cleared wild-type vegetative SAM. d, Cleared ct2 vegetative SAM showing increased diameter (± denotes s.d.; n = 10 for each genotype). e, Wild-type SAM longitudinal section probed in situ for KNOTTED1 (KN1) mRNA. f, ct2 SAM longitudinal section probed in situ for KN1 mRNA showing a wider expression domain but normal patterning. g, h, Transverse sections of wild-type and ct2 SAM probed in situ for KN1. i, SEM of wild-type ear primordium showing a regular shaped inflorescence meristem (IM) and regular arrangement of spikelet pair meristems (SPM) on the flanks. j, ct2 ear primordium showing fasciation of the inflorescence meristem and irregular arrangement of SPM. k, Wild-type tassel primordium. l, ct2 tassel primordium showing reduced length of main rachis and tassel branches and increased spikelet density. Scale bars represent 100 µm (e–h), 1 mm (i, j) and 2 mm (k, l).
Extended Data Figure 2 ct2 alleles and complementation by the CT2–YFP transgene.
a, Schematic of the ct2 locus with mutations marked and agarose gel electrophoresis of partial ct2-Ref (left) and ct2-258 (right) complementary DNA (cDNA) fragments compared to the corresponding wild type. cDNA fragments show increased size for the ct2-Ref allele owing to the insertion within exon 14, and reduced size for the ct2-258 allele owing to skipping of exons 12 and 13. Primer locations are indicated in a. All 3 EMS induced alleles had mutations that induced aberrant splicing, which led to the introduction of premature stop codons. ct2-258, had a G to A transition at the last nucleotide of exon 12, disrupting the 5′ splice site of exon 12. This mutation caused skipping of exons 12 and 13. The G to A transition at the last nucleotide of exon 4 of the ct2-2 allele disrupted the natural 5′ splice site, resulting in activation of a cryptic 5′ splice site five nucleotides downstream. Another G to A transition at the last nucleotide of intron 2 in the ct2-27 allele disrupted the natural 3′ splice site generating a frame-shifted mRNA missing the first seven nucleotides of exon 3. b, ct2 and CT2–YFP transgene genotyping. The genotyping scheme relies on an AccI site single nucleotide polymorphism that is lost in ct2-Ref and amplification of the YFP insertion in the transgene, as described in the Methods. c, Leaf length was indistinguishable between transgenic, homozygous ct2-Ref and wild-type plants, but significantly longer than non-transgenic ct2-Ref homozygous mutants (P value = 0.001, Student’s t test; n = 10 for each genotype; error bars represent s.d.), indicating complementation. d, Seedling phenotypes of transgenic, wild-type plants. e, Seedling phenotype of transgenic, homozygous ct2-Ref plants. f, Seedling phenotype of non-transgenic, homozygous ct2-Ref plants; plants are dwarfed and develop shorter leaves. Pictures d–f are shown at identical magnification.
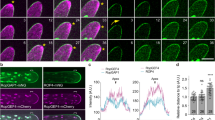
Extended Data Figure 3 CT2–YFP expression.
a, CT2–YFP expression in median optical section of the inflorescence meristem (IM) of an ear primordium, note expression is enhanced in outer layers. b, CT2–YFP expression in a spikelet pair meristem (SPM), showing enhanced expression in the L1 (arrows). c, More centrally localized CT2–YFP expression within a developing spikelet meristem (SM). d, CT2–YFP expression during early stages of floral development, in stamen (st) and carpel (ca) primordia. In both organs, YFP signal is enhanced in the L1 layer (arrow). e, Expression during ovule development, in the ovule (ov) and in the outer cell layers of the stamen primordia (st). f, CT2–YFP expression in roots that were counterstained with propidium iodide is localized along the cell periphery. Image e is a three-dimensional reconstruction of a Z-stack series; scale bars, 100 μm.
Extended Data Figure 4 ct2 double mutant analysis.
Spikelet density of ct2-Ref; td1-Ref double mutants was significantly higher than either single mutant (***P value = 0.0001; Student’s t test). In contrast, spikelet density in ct2-Ref; fea2-0 double mutants was not significantly different (NS) from that of ct2 single mutants (P value = 0.42; Student’s t test), even though each single mutant was significantly higher than normal (P value = 0.001; Student’s t test). However, fea2 mutants were significantly different from ct2;fea2 double mutants (**P value 0.001; Student’s t test), suggesting that CT2 also signals independently of FEA2 in control of spikelet density; n ≥ 10 for each genotype; error bars represent s.d. This experiment was conducted with 2 independent biological replicates.
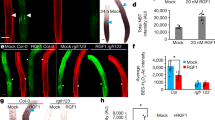
Extended Data Figure 5 Full fractionation western blots and loading controls and FEA2 glycosylation and localization.
a, Western blot probed with an anti-FEA2 peptide antibody and a secondary HRP-coupled antibody (GE); loading quantities shown on right side by 0.1% Ponceau S stain. The anti-FEA2 peptide antibody detects a band of ∼70 kDa in B73 membrane enriched fractions (lane 3). Specificity of the antiserum is evident by the lack of a cross-reacting band in the membrane-enriched fraction from fea2 ears (lane 4). b, FEA2 is glycosylated. It runs at ∼70 kDa in B73 membrane-enriched fractions (lane 1) and is reduced in size upon treatment with PNGaseF glycosidase (lane 2). c, Subcellular fractionation of FEA2 by aqueous two-phase partitioning. The lower (L) and upper (U) phase fractions from B73 inflorescences were resolved on a 10% SDS–PAGE gel, and probed with antisera against BiP lumenal binding protein, or plasma membrane H+ ATPase (Agrisera), or FEA2. Similar loading is shown by Ponceau staining. The endogenous FEA2 distribution is similar to the plasma membrane H+ ATPase, suggesting that it localizes predominantly to the plasma membrane. d, Western blot probed with an anti-GFP peptide antibody and a secondary HRP coupled antibody (GE), loading quantities shown on right side by 0.1% Ponceau S stain. e, Complete scan of the IP western blot shown in Fig. 3d, showing full fractionation and all lanes on same gel. The blot was probed with an anti-FEA2 peptide antibody and a secondary HRP coupled antibody (GE). MF, membrane fraction. This experiment was conducted with 3 biological replicates.
Extended Data Figure 6 Overlapping sizes of FEA2 and CT2 protein complexes.
In the lower panel, membrane enriched extracts from wild-type ears were fractionated by gel filtration, and fractions separated by SDS–PAGE and probed with the FEA2 antiserum. FEA2 was detected in complexes ranging from ∼350 to 550 kDa, with a peak around 400 kDa. The positions of the marker proteins are indicated. In the upper panel, the CT2–YFP fusion protein was detected by spectroscopy as it eluted from the column. In cytoplasmic extracts, CT2–YFP in the soluble fraction (CT2–YFP SOL) eluted at ∼100 kDa, suggesting an interaction with unknown cytoplasmic protein(s). In contrast, a membrane-enriched fraction (CT2–YFP MF) showed that CT2–YFP membrane complexes were a broad range of sizes that overlapped with the FEA2 complex size at ∼400 kDa (shaded area).
Supplementary information
Supplementary Table 1 - Mass spectroscopy analysis of CT2-YFP IPs
We performed 2 independent IP/ Mudpit mass spec runs using CT2-YFP, and we only considered hits with 2 or more significantly matching peptides. The list of potential CT2 interactors includes matches that were present in both runs, after protein matches that were found in control (non-transgenic) IPs were eliminated from the list. The IP mass spec analysis detected CT2 itself, the heterotrimeric G protein subunit (annotated as ZmAGB1), and three predicted LRR receptors, see file for additional information. (XLSX 53 kb)
Rights and permissions
About this article
Cite this article
Bommert, P., Je, B., Goldshmidt, A. et al. The maize Gα gene COMPACT PLANT2 functions in CLAVATA signalling to control shoot meristem size. Nature 502, 555–558 (2013). https://doi.org/10.1038/nature12583
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature12583
This article is cited by
-
Characterization of sub-tropical maize (Zea Mays L.) inbred lines for the variation in kernel row numbers (KRNs)
Cereal Research Communications (2024)
-
A role for heritable transcriptomic variation in maize adaptation to temperate environments
Genome Biology (2023)
-
Transcriptomic analysis reveals the regulation of early ear-length development in maize
Plant Growth Regulation (2023)
-
Role of Heterotrimeric G-Proteins in Improving Abiotic Stress Tolerance of Crop Plants
Journal of Plant Growth Regulation (2023)
-
Novel insights into maize (Zea mays) development and organogenesis for agricultural optimization
Planta (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.