Abstract
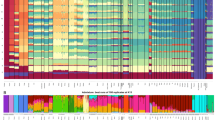
The peopling of the Americas has been the subject of extensive genetic, archaeological and linguistic research; however, central questions remain unresolved1,2,3,4,5. One contentious issue is whether the settlement occurred by means of a single6,7,8 migration or multiple streams of migration from Siberia9,10,11,12,13,14,15. The pattern of dispersals within the Americas is also poorly understood. To address these questions at a higher resolution than was previously possible, we assembled data from 52 Native American and 17 Siberian groups genotyped at 364,470 single nucleotide polymorphisms. Here we show that Native Americans descend from at least three streams of Asian gene flow. Most descend entirely from a single ancestral population that we call ‘First American’. However, speakers of Eskimo–Aleut languages from the Arctic inherit almost half their ancestry from a second stream of Asian gene flow, and the Na-Dene-speaking Chipewyan from Canada inherit roughly one-tenth of their ancestry from a third stream. We show that the initial peopling followed a southward expansion facilitated by the coast, with sequential population splits and little gene flow after divergence, especially in South America. A major exception is in Chibchan speakers on both sides of the Panama isthmus, who have ancestry from both North and South America.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Cavalli-Sforza, L. L., Menozzi, P. & Piazza, A. The History and Geography of Human Genes (Princeton Univ. Press, 1994)
Meltzer, D. J. First Peoples in a New World: Colonizing Ice Age America (Univ. of California Press, 2009)
Goebel, T., Waters, M. R. & O’Rourke, D. H. The late Pleistocene dispersal of modern humans in the Americas. Science 319, 1497–1502 (2008)
Dillehay, T. D. Probing deeper into first American studies. Proc. Natl Acad. Sci. USA 106, 971–978 (2009)
O’Rourke, D. H. & Raff, J. A. The human genetic history of the Americas: the final frontier. Curr. Biol. 20, R202–R207 (2010)
Tamm, E. et al. Beringian standstill and spread of Native American founders. PLoS ONE 2, e829 (2007)
Kitchen, A., Miyamoto, M. M. & Mulligan, C. J. A three-stage colonization model for the peopling of the Americas. PLoS ONE 3, e1596 (2008)
Fagundes, N. J. et al. Mitochondrial population genomics supports a single pre-Clovis origin with a coastal route for the peopling of the Americas. Am. J. Hum. Genet. 82, 583–592 (2008)
Greenberg, J. H., Turner, C. G. & Zegura, S. L. The settlement of the Americas: a comparison of the linguistic, dental, and genetic evidence. Curr. Anthropol. 27, 477–497 (1986)
Lell, J. T. et al. The dual origin and Siberian affinities of Native American Y chromosomes. Am. J. Hum. Genet. 70, 192–206 (2002)
Bortolini, M. C. et al. Y-chromosome evidence for differing ancient demographic histories in the Americas. Am. J. Hum. Genet. 73, 524–539 (2003)
Volodko, N. V. et al. Mitochondrial genome diversity in arctic Siberians, with particular reference to the evolutionary history of Beringia and Pleistocenic peopling of the Americas. Am. J. Hum. Genet. 82, 1084–1100 (2008)
Ray, N. et al. A statistical evaluation of models for the initial settlement of the American continent emphasizes the importance of gene flow with Asia. Mol. Biol. Evol. 27, 337–345 (2010)
de Azevedo, S. et al. Evaluating microevolutionary models for the early settlement of the New World: the importance of recurrent gene flow with Asia. Am. J. Phys. Anthropol. 146, 539–552 (2011)
Perego, U. A. et al. Distinctive Paleo-Indian migration routes from Beringia marked by two rare mtDNA haplogroups. Curr. Biol. 19, 1–8 (2009)
Alexander, D. H., Novembre, J. & Lange, K. Fast model-based estimation of ancestry in unrelated individuals. Genome Res. 19, 1655–1664 (2009)
Ruhlen, M. A Guide to the World’s Languages (Stanford Univ. Press, 1991)
Schroeder, K. B. et al. A private allele ubiquitous in the Americas. Biol. Lett. 3, 218–223 (2007)
Ray, N. PATHMATRIX: a geographical information system tool to compute effective distances among samples. Mol. Ecol. Notes 5, 177–180 (2005)
Wang, S. et al. Genetic variation and population structure in native Americans. PLoS Genet. 3, e185 (2007)
Yang, N. N. et al. Contrasting patterns of nuclear and mtDNA diversity in Native American populations. Ann. Hum. Genet. 74, 525–538 (2010)
Brown, M. D. et al. mtDNA haplogroup X: an ancient link between Europe/Western Asia and North America? Am. J. Hum. Genet. 63, 1852–1861 (1998)
Karafet, T. M. et al. Ancestral Asian source(s) of new world Y-chromosome founder haplotypes. Am. J. Hum. Genet. 64, 817–831 (1999)
Reich, D., Thangaraj, K., Patterson, N., Price, A. L. & Singh, L. Reconstructing Indian population history. Nature 461, 489–494 (2009)
Rasmussen, M. et al. Ancient human genome sequence of an extinct Palaeo-Eskimo. Nature 463, 757–762 (2010)
Balter, M. Archaeology. The peopling of the Aleutians. Science 335, 158–161 (2012)
Cooke, R. Prehistory of native Americans on the Central American land bridge: Colonization, dispersal, and divergence. J. Archaeol. Res. 13, 129–187 (2005)
Wang, S. et al. Geographic patterns of genome admixture in Latin American Mestizos. PLoS Genet. 4, e1000037 (2008)
Bryc, K. et al. Colloquium paper: genome-wide patterns of population structure and admixture among Hispanic/Latino populations. Proc. Natl Acad. Sci. USA 107 (Suppl 2). 8954–8961 (2010)
Wall, J. D. et al. Genetic variation in Native Americans, inferred from Latino SNP and resequencing data. Mol. Biol. Evol. 28, 2231–2237 (2011)
Price, A. L. et al. Sensitive detection of chromosomal segments of distinct ancestry in admixed populations. PLoS Genet. 5, e1000519 (2009)
Browning, S. R. & Browning, B. L. Rapid and accurate haplotype phasing and missing-data inference for whole-genome association studies by use of localized haplotype clustering. Am. J. Hum. Genet. 81, 1084–1097 (2007)
Patterson, N., Price, A. L. & Reich, D. Population structure and eigenanalysis. PLoS Genet. 2, e190 (2006)
Saitou, N. & Nei, M. The neighbor-joining method: a new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 4, 406–425 (1987)
Liu, K. & Muse, S. V. PowerMarker: an integrated analysis environment for genetic marker analysis. Bioinformatics 21, 2128–2129 (2005)
Greenberg, J. H. Language in the Americas (Stanford Univ. Press, 1987)
Campbell, L. American Indian languages: the historical linguistics of Native America (Oxford Univ. Press, 1997)
Acknowledgements
We thank the volunteers who provided the samples that made this study possible. We thank E. D. Ruiz for assistance in the collection involving the Mixtec, Zapotec and Mixe; and P. Herrera for assistance in the collection involving the Quechua; A. Carnevale, M. Crawford, M. Metspalu, F. C. Nielsen, X. Soberon, R. Villems and E. Willerslev for facilitating sharing of data from Mexican, Siberian and Arctic populations; C. Stevens and A. Crenshaw for assistance with genotyping; and P. Bellwood, D. Bolnick, K. Bryc, J. Diamond, T. Dillehay, R. Gonzalez-José, M. Hammer, J. Hill, B. Kemp, S. LeBlanc, D. Meltzer, P. Moorjani, A. Moreno-Estrada, B. Pakendorf, J. Pickrell, M. Ruhlen, D. G. Smith, M. Stoneking, N. Tuross and A. Williams for critiques and discussions. Support was provided by National Institutes of Health grants NS043538 (A.R.-L.), NS037484 and MH075007 (N.B.F.), GM079558 (A.D.), GM079558-S1 (A.D.), GM057672 (K.K.K. and J.R.K.), and HG006399 (D.R., N.P. & A.L.P); by a Biotechnology and Biological Sciences Research Council grant BB/1021213/1; by a National Science Foundation HOMINID grant BCS-1032255 (D.R. and N.P.); by a Canadian Institutes of Health Research grant (D.L.); by a Universidad de Antioquia CODI grant (G.B.); by a Fondo de Investigación Sanitaria grant PS 09/2368 (A.C.); by a Ministerio de Ciencia e Innovacion grant SAF2011-26983 (A.S.); by a Wenner-Gren Foundation grant ICRG-65 (A.D. and R.S.); by Russian Foundation for Basic Research grants 06-04-048182 (R.S.) and 02-06-80524a (L.O.); by a Siberian Branch Russian Academy of Sciences field grant (L.O.); by a Centre National de la Recherche Scientifique Programme Interdisciplinaire de Recherche Amazonie grant (J.-M.D.); and by startup funds from Harvard Medical School (D.R.) and the Harvard School of Public Health (A.L.P.).
Author information
Authors and Affiliations
Contributions
D.R., N.B.F., A.L.P. and A.R-L. conceived the project. D.R., N.P., D.C., A.T., S.M., N.R. and A.R-L. performed analyses. D.R. and A. R.-L. wrote the paper with input from all the co-authors. A.R.-L. assembled the sample collection, directed experimental work and coordinated the study. All other authors contributed to the collection of samples and data.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Additional information
The data analysed here are available for non-profit research on population history under an inter-institutional data access agreement with the Universidad de Antioquia, Colombia; queries regarding data access should be sent to A.R.-L. (a.ruizlin@ucl.ac.uk).
Supplementary information
Supplementary Information
This file contains Supplementary Text 1-7, Supplementary Figures 1-5 and Supplementary Tables 1-3. (PDF 2279 kb)
Rights and permissions
About this article
Cite this article
Reich, D., Patterson, N., Campbell, D. et al. Reconstructing Native American population history. Nature 488, 370–374 (2012). https://doi.org/10.1038/nature11258
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11258
This article is cited by
-
Network of large pedigrees reveals social practices of Avar communities
Nature (2024)
-
Native American ancestry and breast cancer risk in Colombian and Mexican women: ruling out potential confounding through ancestry-informative markers
Breast Cancer Research (2023)
-
Bidirectional dispersals during the peopling of the North American Arctic
Scientific Reports (2023)
-
Comparing Pruning and Thresholding with Continuous Shrinkage Polygenic Score Methods in a Large Sample of Ancestrally Diverse Adolescents from the ABCD Study®
Behavior Genetics (2023)
-
Genomic history of coastal societies from eastern South America
Nature Ecology & Evolution (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.