Abstract
The human genome encodes the blueprint of life, but the function of the vast majority of its nearly three billion bases is unknown. The Encyclopedia of DNA Elements (ENCODE) project has systematically mapped regions of transcription, transcription factor association, chromatin structure and histone modification. These data enabled us to assign biochemical functions for 80% of the genome, in particular outside of the well-studied protein-coding regions. Many discovered candidate regulatory elements are physically associated with one another and with expressed genes, providing new insights into the mechanisms of gene regulation. The newly identified elements also show a statistical correspondence to sequence variants linked to human disease, and can thereby guide interpretation of this variation. Overall, the project provides new insights into the organization and regulation of our genes and genome, and is an expansive resource of functional annotations for biomedical research.
Similar content being viewed by others
Main

The human genome sequence provides the underlying code for human biology. Despite intensive study, especially in identifying protein-coding genes, our understanding of the genome is far from complete, particularly with regard to non-coding RNAs, alternatively spliced transcripts and regulatory sequences. Systematic analyses of transcripts and regulatory information are essential for the identification of genes and regulatory regions, and are an important resource for the study of human biology and disease. Such analyses can also provide comprehensive views of the organization and variability of genes and regulatory information across cellular contexts, species and individuals.
The Encyclopedia of DNA Elements (ENCODE) project aims to delineate all functional elements encoded in the human genome1,2,3. Operationally, we define a functional element as a discrete genome segment that encodes a defined product (for example, protein or non-coding RNA) or displays a reproducible biochemical signature (for example, protein binding, or a specific chromatin structure). Comparative genomic studies suggest that 3–8% of bases are under purifying (negative) selection4,5,6,7,8 and therefore may be functional, although other analyses have suggested much higher estimates9,10,11. In a pilot phase covering 1% of the genome, the ENCODE project annotated 60% of mammalian evolutionarily constrained bases, but also identified many additional putative functional elements without evidence of constraint2. The advent of more powerful DNA sequencing technologies now enables whole-genome and more precise analyses with a broad repertoire of functional assays.
Here we describe the production and initial analysis of 1,640 data sets designed to annotate functional elements in the entire human genome. We integrate results from diverse experiments within cell types, related experiments involving 147 different cell types, and all ENCODE data with other resources, such as candidate regions from genome-wide association studies (GWAS) and evolutionarily constrained regions. Together, these efforts reveal important features about the organization and function of the human genome, summarized below.
• The vast majority (80.4%) of the human genome participates in at least one biochemical RNA- and/or chromatin-associated event in at least one cell type. Much of the genome lies close to a regulatory event: 95% of the genome lies within 8 kilobases (kb) of a DNA–protein interaction (as assayed by bound ChIP-seq motifs or DNase I footprints), and 99% is within 1.7 kb of at least one of the biochemical events measured by ENCODE.
• Primate-specific elements as well as elements without detectable mammalian constraint show, in aggregate, evidence of negative selection; thus, some of them are expected to be functional.
• Classifying the genome into seven chromatin states indicates an initial set of 399,124 regions with enhancer-like features and 70,292 regions with promoter-like features, as well as hundreds of thousands of quiescent regions. High-resolution analyses further subdivide the genome into thousands of narrow states with distinct functional properties.
• It is possible to correlate quantitatively RNA sequence production and processing with both chromatin marks and transcription factor binding at promoters, indicating that promoter functionality can explain most of the variation in RNA expression.
• Many non-coding variants in individual genome sequences lie in ENCODE-annotated functional regions; this number is at least as large as those that lie in protein-coding genes.
• Single nucleotide polymorphisms (SNPs) associated with disease by GWAS are enriched within non-coding functional elements, with a majority residing in or near ENCODE-defined regions that are outside of protein-coding genes. In many cases, the disease phenotypes can be associated with a specific cell type or transcription factor.
ENCODE data production and initial analyses
Since 2007, ENCODE has developed methods and performed a large number of sequence-based studies to map functional elements across the human genome3. The elements mapped (and approaches used) include RNA transcribed regions (RNA-seq, CAGE, RNA-PET and manual annotation), protein-coding regions (mass spectrometry), transcription-factor-binding sites (ChIP-seq and DNase-seq), chromatin structure (DNase-seq, FAIRE-seq, histone ChIP-seq and MNase-seq), and DNA methylation sites (RRBS assay) (Box 1 lists methods and abbreviations; Supplementary Table 1, section P, details production statistics)3. To compare and integrate results across the different laboratories, data production efforts focused on two selected sets of cell lines, designated ‘tier 1’ and ‘tier 2’ (Box 1). To capture a broader spectrum of biological diversity, selected assays were also executed on a third tier comprising more than 100 cell types including primary cells. All data and protocol descriptions are available at http://www.encodeproject.org/, and a User’s Guide including details of cell-type choice and limitations was published recently3.
Integration methodology
For consistency, data were generated and processed using standardized guidelines, and for some assays, new quality-control measures were designed (see refs 3, 12 and http://encodeproject.org/ENCODE/dataStandards.html; A. Kundaje, personal communication). Uniform data-processing methods were developed for each assay (see Supplementary Information; A. Kundaje, personal communication), and most assay results can be represented both as signal information (a per-base estimate across the genome) and as discrete elements (regions computationally identified as enriched for signal). Extensive processing pipelines were developed to generate each representation (M. M. Hoffman et al., manuscript in preparation and A. Kundaje, personal communication). In addition, we developed the irreproducible discovery rate (IDR)13 measure to provide a robust and conservative estimate of the threshold where two ranked lists of results from biological replicates no longer agree (that is, are irreproducible), and we applied this to defining sets of discrete elements. We identified, and excluded from most analyses, regions yielding untrustworthy signals likely to be artefactual (for example, multicopy regions). Together, these regions comprise 0.39% of the genome (see Supplementary Information). The poster accompanying this issue represents different ENCODE-identified elements and their genome coverage.
Transcribed and protein-coding regions
We used manual and automated annotation to produce a comprehensive catalogue of human protein-coding and non-coding RNAs as well as pseudogenes, referred to as the GENCODE reference gene set14,15 (Supplementary Table 1, section U). This includes 20,687 protein-coding genes (GENCODE annotation, v7) with, on average, 6.3 alternatively spliced transcripts (3.9 different protein-coding transcripts) per locus. In total, GENCODE-annotated exons of protein-coding genes cover 2.94% of the genome or 1.22% for protein-coding exons. Protein-coding genes span 33.45% from the outermost start to stop codons, or 39.54% from promoter to poly(A) site. Analysis of mass spectrometry data from K562 and GM12878 cell lines yielded 57 confidently identified unique peptide sequences in intergenic regions relative to GENCODE annotation. Taken together with evidence of pervasive genome transcription16, these data indicate that additional protein-coding genes remain to be found.
In addition, we annotated 8,801 automatically derived small RNAs and 9,640 manually curated long non-coding RNA (lncRNA) loci17. Comparing lncRNAs to other ENCODE data indicates that lncRNAs are generated through a pathway similar to that for protein-coding genes17. The GENCODE project also annotated 11,224 pseudogenes, of which 863 were transcribed and associated with active chromatin18.
RNA
We sequenced RNA16 from different cell lines and multiple subcellular fractions to develop an extensive RNA expression catalogue. Using a conservative threshold to identify regions of RNA activity, 62% of genomic bases are reproducibly represented in sequenced long (>200 nucleotides) RNA molecules or GENCODE exons. Of these bases, only 5.5% are explained by GENCODE exons. Most transcribed bases are within or overlapping annotated gene boundaries (that is, intronic), and only 31% of bases in sequenced transcripts were intergenic16.
We used CAGE-seq (5′ cap-targeted RNA isolation and sequencing) to identify 62,403 transcription start sites (TSSs) at high confidence (IDR of 0.01) in tier 1 and 2 cell types. Of these, 27,362 (44%) are within 100 base pairs (bp) of the 5′ end of a GENCODE-annotated transcript or previously reported full-length messenger RNA. The remaining regions predominantly lie across exons and 3′ untranslated regions (UTRs), and some exhibit cell-type-restricted expression; these may represent the start sites of novel, cell-type-specific transcripts.
Finally, we saw a significant proportion of coding and non-coding transcripts processed into steady-state stable RNAs shorter than 200 nucleotides. These precursors include transfer RNA, microRNA, small nuclear RNA and small nucleolar RNA (tRNA, miRNA, snRNA and snoRNA, respectively) and the 5′ termini of these processed products align with the capped 5′ end tags16.
Protein bound regions
To identify regulatory regions directly, we mapped the binding locations of 119 different DNA-binding proteins and a number of RNA polymerase components in 72 cell types using ChIP-seq (Table 1, Supplementary Table 1, section N, and ref. 19); 87 (73%) were sequence-specific transcription factors. Overall, 636,336 binding regions covering 231 megabases (Mb; 8.1%) of the genome are enriched for regions bound by DNA-binding proteins across all cell types. We assessed each protein-binding site for enrichment of known DNA-binding motifs and the presence of novel motifs. Overall, 86% of the DNA segments occupied by sequence-specific transcription factors contained a strong DNA-binding motif, and in most (55%) cases the known motif was most enriched (P. Kheradpour and M. Kellis, manuscript in preparation).
Protein-binding regions lacking high or moderate affinity cognate recognition sites have 21% lower median scores by rank than regions with recognition sequences (Wilcoxon rank sum P value <10−16). Eighty-two per cent of the low-signal regions have high-affinity recognition sequences for other factors. In addition, when ChIP-seq peaks are ranked by their concordance with their known recognition sequence, the median DNase I accessibility is twofold higher in the bottom 20% of peaks than in the upper 80% (genome structure correction (GSC)20 P value <10−16), consistent with previous observations21,22,23,24. We speculate that low signal regions are either lower-affinity sites21 or indirect transcription-factor target regions associated through interactions with other factors (see also refs 25, 26).
We organized all the information associated with each transcription factor—including the ChIP-seq peaks, discovered motifs and associated histone modification patterns—in FactorBook (http://www.factorbook.org; ref. 26), a public resource that will be updated as the project proceeds.
DNase I hypersensitive sites and footprints
Chromatin accessibility characterized by DNase I hypersensitivity is the hallmark of regulatory DNA regions27,28. We mapped 2.89 million unique, non-overlapping DNase I hypersensitive sites (DHSs) by DNase-seq in 125 cell types, the overwhelming majority of which lie distal to TSSs29. We also mapped 4.8 million sites across 25 cell types that displayed reduced nucleosomal crosslinking by FAIRE, many of which coincide with DHSs. In addition, we used micrococcal nuclease to map nucleosome occupancy in GM12878 and K562 cells30.
In tier 1 and tier 2 cell types, we identified a mean of 205,109 DHSs per cell type (at false discovery rate (FDR) 1%), encompassing an average of 1.0% of the genomic sequence in each cell type, and 3.9% in aggregate. On average, 98.5% of the occupancy sites of transcription factors mapped by ENCODE ChIP-seq (and, collectively, 94.4% of all 1.1 million transcription factor ChIP-seq peaks in K562 cells) lie within accessible chromatin defined by DNase I hotspots29. However, a small number of factors, most prominently heterochromatin-bound repressive complexes (for example, the TRIM28–SETDB1–ZNF274 complex31,32 encoded by the TRIM28, SETDB1 and ZNF274 genes), seem to occupy a significant fraction of nucleosomal sites.
Using genomic DNase I footprinting33,34 on 41 cell types we identified 8.4 million distinct DNase I footprints (FDR 1%)25. Our de novo motif discovery on DNase I footprints recovered ∼90% of known transcription factor motifs, together with hundreds of novel evolutionarily conserved motifs, many displaying highly cell-selective occupancy patterns similar to major developmental and tissue-specific regulators.
Regions of histone modification
We assayed chromosomal locations for up to 12 histone modifications and variants in 46 cell types, including a complete matrix of eight modifications across tier 1 and tier 2. Because modification states may span multiple nucleosomes, which themselves can vary in position across cell populations, we used a continuous signal measure of histone modifications in downstream analysis, rather than calling regions (M. M. Hoffman et al., manuscript in preparation; see http://code.google.com/p/align2rawsignal/). For the strongest, ‘peak-like’ histone modifications, we used MACS35 to characterize enriched sites. Table 2 describes the different histone modifications, their peak characteristics, and a summary of their known roles (reviewed in refs 36–39).
Our data show that global patterns of modification are highly variable across cell types, in accordance with changes in transcriptional activity. Consistent with previous studies40,41, we find that integration of the different histone modification information can be used systematically to assign functional attributes to genomic regions (see below).
DNA methylation
Methylation of cytosine, usually at CpG dinucleotides, is involved in epigenetic regulation of gene expression. Promoter methylation is typically associated with repression, whereas genic methylation correlates with transcriptional activity42. We used reduced representation bisulphite sequencing (RRBS) to profile DNA methylation quantitatively for an average of 1.2 million CpGs in each of 82 cell lines and tissues (8.6% of non-repetitive genomic CpGs), including CpGs in intergenic regions, proximal promoters and intragenic regions (gene bodies)43, although it should be noted that the RRBS method preferentially targets CpG-rich islands. We found that 96% of CpGs exhibited differential methylation in at least one cell type or tissue assayed (K. Varley et al., personal communication), and levels of DNA methylation correlated with chromatin accessibility. The most variably methylated CpGs are found more often in gene bodies and intergenic regions, rather than in promoters and upstream regulatory regions. In addition, we identified an unexpected correspondence between unmethylated genic CpG islands and binding by P300, a histone acetyltransferase linked to enhancer activity44.
Because RRBS is a sequence-based assay with single-base resolution, we were able to identify CpGs with allele-specific methylation consistent with genomic imprinting, and determined that these loci exhibit aberrant methylation in cancer cell lines (K. Varley et al., personal communication). Furthermore, we detected reproducible cytosine methylation outside CpG dinucleotides in adult tissues45, providing further support that this non-canonical methylation event may have important roles in human biology (K. Varley et al., personal communication).
Chromosome-interacting regions
Physical interaction between distinct chromosome regions that can be separated by hundreds of kilobases is thought to be important in the regulation of gene expression46. We used two complementary chromosome conformation capture (3C)-based technologies to probe these long-range physical interactions.
A 3C-carbon copy (5C) approach47,48 provided unbiased detection of long-range interactions with TSSs in a targeted 1% of the genome (the 44 ENCODE pilot regions) in four cell types (GM12878, K562, HeLa-S3 and H1 hESC)49. We discovered hundreds of statistically significant long-range interactions in each cell type after accounting for chromatin polymer behaviour and experimental variation. Pairs of interacting loci showed strong correlation between the gene expression level of the TSS and the presence of specific functional element classes such as enhancers. The average number of distal elements interacting with a TSS was 3.9, and the average number of TSSs interacting with a distal element was 2.5, indicating a complex network of interconnected chromatin. Such interwoven long-range architecture was also uncovered genome-wide using chromatin interaction analysis with paired-end tag sequencing (ChIA-PET)50 applied to identify interactions in chromatin enriched by RNA polymerase II (Pol II) ChIP from five cell types51. In K562 cells, we identified 127,417 promoter-centred chromatin interactions using ChIA-PET, 98% of which were intra-chromosomal. Whereas promoter regions of 2,324 genes were involved in ‘single-gene’ enhancer–promoter interactions, those of 19,813 genes were involved in ‘multi-gene’ interaction complexes spanning up to several megabases, including promoter–promoter and enhancer–promoter interactions51.
These analyses portray a complex landscape of long-range gene–element connectivity across ranges of hundreds of kilobases to several megabases, including interactions among unrelated genes (Supplementary Fig. 1, section Y). Furthermore, in the 5C results, 50–60% of long-range interactions occurred in only one of the four cell lines, indicative of a high degree of tissue specificity for gene–element connectivity49.
Summary of ENCODE-identified elements
Accounting for all these elements, a surprisingly large amount of the human genome, 80.4%, is covered by at least one ENCODE-identified element (detailed in Supplementary Table 1, section Q). The broadest element class represents the different RNA types, covering 62% of the genome (although the majority is inside of introns or near genes). Regions highly enriched for histone modifications form the next largest class (56.1%). Excluding RNA elements and broad histone elements, 44.2% of the genome is covered. Smaller proportions of the genome are occupied by regions of open chromatin (15.2%) or sites of transcription factor binding (8.1%), with 19.4% covered by at least one DHS or transcription factor ChIP-seq peak across all cell lines. Using our most conservative assessment, 8.5% of bases are covered by either a transcription-factor-binding-site motif (4.6%) or a DHS footprint (5.7%). This, however, is still about 4.5-fold higher than the amount of protein-coding exons, and about twofold higher than the estimated amount of pan-mammalian constraint.
Given that the ENCODE project did not assay all cell types, or all transcription factors, and in particular has sampled few specialized or developmentally restricted cell lineages, these proportions must be underestimates of the total amount of functional bases. However, many assays were performed on more than one cell type, allowing assessment of the rate of discovery of new elements. For both DHSs and CTCF-bound sites, the number of new elements initially increases rapidly with a steep gradient for the saturation curve and then slows with increasing number of cell types (Supplementary Figs 1 and 2, section R). With the current data, at the flattest part of the saturation curve each new cell type adds, on average, 9,500 DHS elements (across 106 cell types) and 500 CTCF-binding elements (across 49 cell types), representing 0.45% of the total element number. We modelled saturation for the DHSs and CTCF-binding sites using a Weibull distribution (r2 > 0.999) and predict saturation at approximately 4.1 million (standard error (s.e.) = 108,000) and 185,100 (s.e. = 18,020) sites, respectively, indicating that we have discovered around half of the estimated total DHSs. These estimates represent a lower bound, but reinforce the observation that there is more non-coding functional DNA than either coding sequence or mammalian evolutionarily constrained bases.
The impact of selection on functional elements
From comparative genomic studies, at least 3–8% of bases are under purifying (negative) selection4,5,6,7,8,9,10,11, indicating that these bases may potentially be functional. We previously found that 60% of mammalian evolutionarily constrained bases were annotated in the ENCODE pilot project, but also observed that many functional elements lacked evidence of constraint2, a conclusion substantiated by others52,53,54. The diversity and genome-wide occurrence of functional elements now identified provides an unprecedented opportunity to examine further the forces of negative selection on human functional sequences.
We examined negative selection using two measures that highlight different periods of selection in the human genome. The first measure, inter-species, pan-mammalian constraint (GERP-based scores; 24 mammals8), addresses selection during mammalian evolution. The second measure is intra-species constraint estimated from the numbers of variants discovered in human populations using data from the 1000 Genomes project55, and covers selection over human evolution. In Fig. 1, we plot both these measures of constraint for different classes of identified functional elements, excluding features overlapping exons and promoters that are known to be constrained. Each graph also shows genomic background levels and measures of coding-gene constraint for comparison. Because we plot human population diversity on an inverted scale, elements that are more constrained by negative selection will tend to lie in the upper and right-hand regions of the plot.
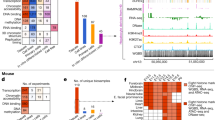
a, Levels of pan-mammalian constraint (mean GERP score; 24 mammals8, x axis) compared to diversity, a measure of negative selection in the human population (mean expected heterozygosity, inverted scale, y axis) for ENCODE data sets. Each point is an average for a single data set. The top-right corners have the strongest evolutionary constraint and lowest diversity. Coding (C), UTR (U), genomic (G), intergenic (IG) and intronic (IN) averages are shown as filled squares. In each case the vertical and horizontal cross hairs show representative levels for the neutral expectation for mammalian conservation and human population diversity, respectively. The spread over all non-exonic ENCODE elements greater than 2.5 kb from TSSs is shown. The inner dashed box indicates that parts of the plot have been magnified for the surrounding outer panels, although the scales in the outer plots provide the exact regions and dimensions magnified. The spread for DHS sites (b) and RNA elements (d) is shown in the plots on the left. RNA elements are either long novel intronic (dark green) or long intergenic (light green) RNAs. The horizontal cross hairs are colour-coded to the relevant data set in d. c, Spread of transcription factor motif instances either in regions bound by the transcription factor (orange points) or in the corresponding unbound motif matches in grey, with bound and unbound points connected with an arrow in each case showing that bound sites are generally more constrained and less diverse. e, Derived allele frequency spectrum for primate-specific elements, with variations outside ENCODE elements in black and variations covered by ENCODE elements in red. The increase in low-frequency alleles compared to background is indicative of negative selection occurring in the set of variants annotated by the ENCODE data. f, Aggregation of mammalian constraint scores over the glucocorticoid receptor (GR) transcription factor motif in bound sites, showing the expected correlation with the information content of bases in the motif. An interactive version of this figure is available in the online version of the paper.
For DNase I elements (Fig. 1b) and bound motifs (Fig. 1c), most sets of elements show enrichment in pan-mammalian constraint and decreased human population diversity, although for some cell types the DNase I sites do not seem overall to be subject to pan-mammalian constraint. Bound transcription factor motifs have a natural control from the set of transcription factor motifs with equal sequence potential for binding but without binding evidence from ChIP-seq experiments—in all cases, the bound motifs show both more mammalian constraint and higher suppression of human diversity.
Consistent with previous findings, we do not observe genome-wide evidence for pan-mammalian selection of novel RNA sequences (Fig. 1d). There are also a large number of elements without mammalian constraint, between 17% and 90% for transcription-factor-binding regions as well as DHSs and FAIRE regions. Previous studies could not determine whether these sequences are either biochemically active, but with little overall impact on the organism, or under lineage-specific selection. By isolating sequences preferentially inserted into the primate lineage, which is only feasible given the genome-wide scale of this data, we are able to examine this issue specifically. Most primate-specific sequence is due to retrotransposon activity, but an appreciable proportion is non-repetitive primate-specific sequence. Of 104,343,413 primate-specific bases (excluding repetitive elements), 67,769,372 (65%) are found within ENCODE-identified elements. Examination of 227,688 variants segregating in these primate-specific regions revealed that all classes of elements (RNA and regulatory) show depressed derived allele frequencies, consistent with recent negative selection occurring in at least some of these regions (Fig. 1e). An alternative approach examining sequences that are not clearly under pan-mammalian constraint showed a similar result (L. Ward and M. Kellis, manuscript submitted). This indicates that an appreciable proportion of the unconstrained elements are lineage-specific elements required for organismal function, consistent with long-standing views of recent evolution56, and the remainder are probably ‘neutral’ elements2 that are not currently under selection but may still affect cellular or larger scale phenotypes without an effect on fitness.
The binding patterns of transcription factors are not uniform, and we can correlate both inter- and intra-species measures of negative selection with the overall information content of motif positions. The selection on some motif positions is as high as protein-coding exons (Fig. 1f; L. Ward and M. Kellis, manuscript submitted). These aggregate measures across motifs show that the binding preferences found in the population of sites are also relevant to the per-site behaviour. By developing a per-site metric of population effect on bound motifs, we found that highly constrained bound instances across mammals are able to buffer the impact of individual variation57.
ENCODE data integration with known genomic features
Promoter-anchored integration
Many of the ENCODE assays directly or indirectly provide information about the action of promoters. Focusing on the TSSs of protein-coding transcripts, we investigated the relationships between different ENCODE assays, in particular testing the hypothesis that RNA expression (output) can be effectively predicted from patterns of chromatin modification or transcription factor binding (input). Consistent with previous reports58, we observe two relatively distinct types of promoter: (1) broad, mainly (C+G)-rich, TATA-less promoters; and (2) narrow, TATA-box-containing promoters. These promoters have distinct patterns of histone modifications, and transcription-factor-binding sites are selectively enriched in each class (Supplementary Fig. 1, section Z).
We developed predictive models to explore the interaction between histone modifications and measures of transcription at promoters, distinguishing between modifications known to be added as a consequence of transcription (such as H3K36me3 and H3K79me2) and other categories of histone marks59. In our analyses, the best models had two components: an initial classification component (on/off) and a second quantitative model component. Our models showed that activating acetylation marks (H3K27ac and H3K9ac) are roughly as informative as activating methylation marks (H3K4me3 and H3K4me2) (Fig. 2a). Although repressive marks, such as H3K27me3 or H3K9me3, show negative correlation both individually and in the model, removing these marks produces only a small reduction in model performance. However, for a subset of promoters in each cell line, repressive histone marks (H3K27me3 or H3K9me3) must be used to predict their expression accurately. We also examined the interplay between the H3K79me2 and H3K36me3 marks, both of which mark gene bodies, probably reflecting recruitment of modification enzymes by polymerase isoforms. As described previously, H3K79me2 occurs preferentially at the 5′ ends of gene bodies and H3K36me3 occurs more 3′, and our analyses support the previous model in which the H3K79me2 to H3K36me3 transition occurs at the first 3′ splice site60.
a, b, Correlative models between either histone modifications or transcription factors, respectively, and RNA production as measured by CAGE tag density at TSSs in K562 cells. In each case the scatter plot shows the output of the correlation models (x axis) compared to observed values (y axis). The bar graphs show the most important histone modifications (a) or transcription factors (b) in both the initial classification phase (top bar graph) or the quantitative regression phase (bottom bar graph), with larger values indicating increasing importance of the variable in the model. Further analysis of other cell lines and RNA measurement types is reported elsewhere59,79. AUC, area under curve; Gini, Gini coefficient; RMSE, root mean square error.
Few previous studies have attempted to build qualitative or quantitative models of transcription genome-wide from transcription factor levels because of the paucity of documented transcription-factor-binding regions and the lack of coordination around a single cell line. We thus examined the predictive capacity of transcription-factor-binding signals for the expression levels of promoters (Fig. 2b). In contrast to the profiles of histone modifications, most transcription factors show enriched binding signals in a narrow DNA region near the TSS, with relatively higher binding signals in promoters with higher CpG content. Most of this correlation could be recapitulated by looking at the aggregate binding of transcription factors without specific transcription factor terms. Together, these correlation models indicate both that a limited set of chromatin marks are sufficient to ‘explain’ transcription and that a variety of transcription factors might have broad roles in general transcription levels across many genes. It is important to note that this is an inherently observational study of correlation patterns, and is consistent with a variety of mechanistic models with different causal links between the chromatin, transcription factor and RNA assays. However, it does indicate that there is enough information present at the promoter regions of genes to explain most of the variation in RNA expression.
We developed predictive models similar to those used to model transcriptional activity to explore the relationship between levels of histone modification and inclusion of exons in alternately spliced transcripts. Even accounting for expression level, H3K36me3 has a positive contribution to exon inclusion, whereas H3K79me2 has a negative contribution (H. Tilgner et al., manuscript in preparation). By monitoring the RNA populations in the subcellular fractions of K562 cells, we found that essentially all splicing is co-transcriptional61, further supporting a link between chromatin structure and splicing.
Transcription-factor-binding site-anchored integration
Transcription-factor-binding sites provide a natural focus around which to explore chromatin properties. Transcription factors are often multifunctional and can bind a variety of genomic loci with different combinations and patterns of chromatin marks and nucleosome organization. Hence, rather than averaging chromatin mark profiles across all binding sites of a transcription factor, we developed a clustering procedure, termed the Clustered Aggregation Tool (CAGT), to identify subsets of binding sites sharing similar but distinct patterns of chromatin mark signal magnitude, shape and hidden directionality30. For example, the average profile of the repressive histone mark H3K27me3 over all 55,782 CTCF-binding sites in H1 hESCs shows poor signal enrichment (Fig. 3a). However, after grouping profiles by signal magnitude we found a subset of 9,840 (17.6%) CTCF-binding sites that exhibit significant flanking H3K27me3 signal. Shape and orientation analysis further revealed that the predominant signal profile for H3K27me3 around CTCF peak summits is asymmetric, consistent with a boundary role for some CTCF sites between active and polycomb-silenced domains. Further examples are provided in Supplementary Figs 5 and 6 of section E. For TAF1, predominantly found near TSSs, the asymmetric sites are orientated with the direction of transcription. However, for distal sites, such as those bound by GATA1 and CTCF, we also observed a high proportion of asymmetric histone patterns, although independent of motif directionality. In fact, all transcription-factor-binding data sets in all cell lines show predominantly asymmetric patterns (asymmetry ratio >0.6) for all chromatin marks but not for DNase I signal (Fig. 3b). This indicates that most transcription-factor-bound chromatin events correlate with structured, directional patterns of histone modifications, and that promoter directionality is not the only source of orientation at these sites.
a, Results of clustered aggregation of H3K27me3 modification signal around CTCF-binding sites (a multifunctional protein involved with chromatin structure). The first three plots (left column) show the signal behaviour of the histone modification over all sites (top) and then split into the high and low signal components. The solid lines show the mean signal distribution by relative position with the blue shaded area delimiting the tenth and ninetieth percentile range. The high signal component is then decomposed further into six different shape classes on the right (see ref. 30 for details). The shape decomposition process is strand aware. b, Summary of shape asymmetry for DNase I, nucleosome and histone modification signals by plotting an asymmetry ratio for each signal over all transcription-factor-binding sites. All histone modifications measured in this study show predominantly asymmetric patterns at transcription-factor-binding sites. An interactive version of this figure is available in the online version of the paper.
We also examined nucleosome occupancy relative to the symmetry properties of chromatin marks around transcription-factor-binding sites. Around TSSs, there is usually strong asymmetric nucleosome occupancy, often accounting for most of the histone modification signal (for instance, see Supplementary Fig. 4, section E). However, away from TSSs, there is far less concordance. For example, CTCF-binding sites typically show arrays of well-positioned nucleosomes on either side of the peak summit (Supplementary Fig. 1, section E)62. Where the flanking chromatin mark signal is high, the signals are often asymmetric, indicating differential marking with histone modifications (Supplementary Figs 2 and 3, section E). Thus, we confirm on a genome-wide scale that transcription factors can form barriers around which nucleosomes and histone modifications are arranged in a variety of configurations62,63,64,65. This is explored in further detail in refs 25, 26 and 30.
Transcription factor co-associations
Transcription-factor-binding regions are nonrandomly distributed across the genome, with respect to both other features (for example, promoters) and other transcription-factor-binding regions. Within the tier 1 and 2 cell lines, we found 3,307 pairs of statistically co-associated factors (P <1 × 10−16, GSC) involving 114 out of a possible 117 factors (97%) (Fig. 4a). These include expected associations, such as Jun and Fos, and some less expected novel associations, such as TCF7L2 with HNF4-α and FOXA2 (ref. 66; a full listing is given in Supplementary Table 1, section F). When one considers promoter and intergenic regions separately, this changes to 3,201 pairs (116 factors, 99%) for promoters and 1,564 pairs (108 factors, 92%) for intergenic regions, with some associations more specific to these genomic contexts (for example, the cluster of HDAC2, GABPA, CHD2, GTF2F1, MXI1 and MYC in promoter regions and SP1, EP300, HDAC2 and NANOG in intergenic regions (Fig. 4b)). These general and context-dependent associations lead to a network representation of the co-binding with many interesting properties, explored in refs 19, 25 and 26. In addition, we also identified a set of regions bound by multiple factors representing high occupancy of transcription factor (HOT) regions67.
a, Significant co-associations of transcription factor pairs using the GSC statistic across the entire genome in K562 cells. The colour strength represents the extent of association (from red (strongest), orange, to yellow (weakest)), whereas the depth of colour represents the fit to the GSC20 model (where white indicates that the statistical model is not appropriate) as indicated by the key. Most transcription factors have a nonrandom association to other transcription factors, and these associations are dependent on the genomic context, meaning that once the genome is separated into promoter proximal and distal regions, the overall levels of co-association decrease, but more specific relationships are uncovered. b, Three classes of behaviour are shown. The first column shows a set of associations for which strength is independent of location in promoter and distal regions, whereas the second column shows a set of transcription factors that have stronger associations in promoter-proximal regions. Both of these examples are from data in K562 cells and are highlighted on the genome-wide co-association matrix (a) by the labelled boxes A and B, respectively. The third column shows a set of transcription factors that show stronger association in distal regions (in the H1 hESC line). An interactive version of this figure is available in the online version of the paper.
Genome-wide integration
To identify functional regions genome-wide, we next integrated elements independent of genomic landmarks using either discriminative training methods, where a subset of known elements of a particular class were used to train a model that was then used to discover more instances of this class, or using methods in which only data from ENCODE assays were used without explicit knowledge of any annotation.
For discriminative training, we used a three-step process to predict potential enhancers, described in Supplementary Information and ref. 67. Two alternative discriminative models converged on a set of ∼13,000 putative enhancers in K562 cells67. In the second approach, two methodologically distinct unbiased approaches (see refs 40, 68 and M. M. Hoffman et al., manuscript in preparation) converged on a concordant set of histone modification and chromatin-accessibility patterns that can be used to segment the genome in each of the tier 1 and tier 2 cell lines, although the individual loci in each state in each cell line are different. With the exception of RNA polymerase II and CTCF, the addition of transcription factor data did not substantially alter these patterns. At this stage, we deliberately excluded RNA and methylation assays, reserving these data as a means to validate the segmentations.
Our integration of the two segmentation methods (M. M. Hoffman et al., manuscript in preparation) established a consensus set of seven major classes of genome states, described in Table 3. The standard view of active promoters, with a distinct core promoter region (TSS and PF states), leading to active gene bodies (T, transcribed state), is rediscovered in this model (Fig. 5a, b). There are three ‘active’ distal states. We tentatively labelled two as enhancers (predicted enhancers, E, and predicted weak enhancers, WE) due to their occurrence in regions of open chromatin with high H3K4me1, although they differ in the levels of marks such as H3K27ac, currently thought to distinguish active from inactive enhancers. The other active state (CTCF) has high CTCF binding and includes sequences that function as insulators in a transfection assay. The remaining repressed state (R) summarizes sequences split between different classes of actively repressed or inactive, quiescent chromatin. We found that the CTCF-binding-associated state is relatively invariant across cell types, with individual regions frequently occupying the CTCF state across all six cell types (Fig. 5c). Conversely, the E and T states have substantial cell-specific behaviour, whereas the TSS state has a bimodal behaviour with similar numbers of cell-invariant and cell-specific occurrences. It is important to note that the consensus summary classes do not capture all the detail discovered in the individual segmentations containing more states.
a, Illustrative region with the two segmentation methods (ChromHMM and Segway) in a dense view and the combined segmentation expanded to show each state in GM12878 cells, beneath a compressed view of the GENCODE gene annotations. Note that at this level of zoom and genome browser resolution, some segments appear to overlap although they do not. Segmentation classes are named and coloured according to the scheme in Table 3. Beneath the segmentations are shown each of the normalized signals that were used as the input data for the segmentations. Open chromatin signals from DNase-seq from the University of Washington group (UW DNase) or the ENCODE open chromatin group (Openchrom DNase) and FAIRE assays are shown in blue; signal from histone modification ChIP-seq in red; and transcription factor ChIP-seq signal for Pol II and CTCF in green. The mauve ChIP-seq control signal (input control) at the bottom was also included as an input to the segmentation. b, Association of selected transcription factor (left) and RNA (right) elements in the combined segmentation states (x axis) expressed as an observed/expected ratio (obs./exp.) for each combination of transcription factor or RNA element and segmentation class using the heat-map scale shown in the key besides each heat map. c, Variability of states between cell lines, showing the distribution of occurrences of the state in the six cell lines at specific genome locations: from unique to one cell line to ubiquitous in all six cell lines for five states (CTCF, E, T, TSS and R). d, Distribution of methylation level at individual sites from RRBS analysis in GM12878 cells across the different states, showing the expected hypomethylation at TSSs and hypermethylation of genes bodies (T state) and repressed (R) regions.
The distribution of RNA species across segments is quite distinct, indicating that underlying biological activities are captured in the segmentation. Polyadenylated RNA is heavily enriched in gene bodies. Around promoters, there are short RNA species previously identified as promoter-associated short RNAs (Fig. 5b)16,69. Similarly, DNA methylation shows marked distinctions between segments, recapitulating the known biology of predominantly unmethylated active promoters (TSS states) followed by methylated gene bodies42 (T state, Fig. 5d). The two enhancer-enriched states show distinct patterns of DNA methylation, with the less active enhancer state (by H3K27ac/H3K4me1 levels) showing higher methylation. These states also have an excess of RNA elements without poly(A) tails and methyl-cap RNA, as assayed by CAGE sequences, compared to matched intergenic controls, indicating a specific transcriptional mode associated with active enhancers70. Transcription factors also showed distinct distributions across the segments (Fig. 5b). A striking pattern is the concentration of transcription factors in the TSS-associated state. The enhancers contain a different set of transcription factors. For example, in K562 cells, the E state is enriched for binding by the proteins encoded by the EP300, FOS, FOSL1, GATA2, HDAC8, JUNB, JUND, NFE2, SMARCA4, SMARCB1, SIRT6 and TAL1 genes. We tested a subset of these predicted enhancers in both mouse and fish transgenic models (examples in Fig. 6), with over half of the elements showing activity, often in the corresponding tissue type.
Randomly sampled E state segments (see Table 3) from the K562 segmentation were cloned for mouse- and fish-based transgenic enhancer assays. a, Representative LacZ-stained transgenic embryonic day (E)11.5 mouse embryo obtained with construct hs2065 (EN167, chr10: 46052882–46055670, GRCh37). Highly reproducible staining in the blood vessels was observed in 9 out of 9 embryos resulting from independent transgenic integration events. b, Representative green fluorescent protein reporter transgenic medaka fish obtained from a construct with a basal hsp70 promoter on meganuclease-based transfection. Reproducible transgenic expression in the circulating nucleated blood cells and the endothelial cell walls was seen in 81 out of 100 transgenic tests of this construct.
The segmentation provides a linear determination of functional state across the genome, but not an association of particular distal regions with genes. By using the variation of DNase I signal across cell lines, 39% of E (enhancer associated) states could be linked to a proposed regulated gene29 concordant with physical proximity patterns determined by 5C49 or ChIA-PET.
To provide a fine-grained regional classification, we turned to a self organizing map (SOM) to cluster genome segmentation regions based on their assay signal characteristics (Fig. 7). The segmentation regions were initially randomly assigned to a 1,350-state map in a two-dimensional toroidal space (Fig. 7a). This map can be visualized as a two-dimensional rectangular plane onto which the various signal distributions can be plotted. For instance, the rectangle at the bottom left of Fig. 7a shows the distribution of the genome in the initial randomized map. The SOM was then trained using the twelve different ChIP-seq and DNase-seq assays in the six cell types previously analysed in the large-scale segmentations (that is, over 72-dimensional space). After training, the SOM clustering was again visualized in two dimensions, now showing the organized distribution of genome segments (lower right of panel, Fig. 7a). Individual data sets associated with the genome segments in each SOM map unit (hexagonal cells) can then be visualized in the same framework to learn how each additional kind of data is distributed on the chromatin state map. Figure 7b shows CAGE/TSS expression data overlaid on the randomly initialized (left) and trained map (right) panels. In this way the trained SOM highlighted cell-type-specific TSS clusters (bottom panels of Fig. 7b), indicating that there are sets of tissue-specific TSSs that are distinguished from each other by subtle combinations of ENCODE chromatin data. Many of the ultra-fine-grained state classifications revealed in the SOM are associated with specific gene ontology (GO) terms (right panel of Fig. 7c). For instance, the left panel of Fig. 7c identifies ten SOM map units enriched with genomic regions associated with genes associated with the GO term ‘immune response’. The central panel identifies a different set of map units enriched for the GO term ‘sequence-specific transcription factor activity’. The two map units most enriched for this GO term, indicated by the darkest green colouring, contain genes with segments that are high in H3K27me3 in H1 hESCs, but that differ in H3K27me3 levels in HUVECs. Gene function analysis with the GO ontology tool (GREAT71) reveals that the map unit with high H3K27me3 levels in both cell types is enriched in transcription factor genes with known neuronal functions, whereas the neighbouring map unit is enriched in genes involved in body patterning. The genome browser shots at the bottom of Fig. 7c pick out an example region for each of the two SOM map units illustrating the difference in H3K27me3 signal. Overall, we have 228 distinct GO terms associated with specific segments across one or more states (A. Mortazavi, personal communication), and can assign over one-third of genes to a GO annotation solely on the basis of its multicellular histone patterns. Thus, the SOM analysis provides a fine-grained map of chromatin data across multiple cell types, which can then be used to relate chromatin structure to other data types at differing levels of resolution (for instance, the large cluster of units containing any active TSS, its subclusters composed of units enriched in TSSs active in only one cell type, or individual map units significantly enriched for specific GO terms).
a–c, The training of the SOM (a) and analysis of the results (b, c) are shown. Initially we arbitrarily placed genomic segments from the ChromHMM segmentation on to the toroidal map surface, although the SOM does not use the ChromHMM state assignments (a). We then trained the map using the signal of the 12 different ChIP-seq and DNase-seq assays in the six cell types analysed. Each unit of the SOM is represented here by a hexagonal cell in a planar two-dimensional view of the toroidal map. Curved arrows indicate that traversing the edges of two dimensional view leads back to the opposite edge. The resulting map can be overlaid with any class of ENCODE or other data to view the distribution of that data within this high-resolution segmentation. In panel a the distributions of genome bases across the untrained and trained map (left and right, respectively) are shown using heat-map colours for log10 values. b, The distribution of TSSs from CAGE experiments of GENCODE annotation on the planar representations of either the initial random organization (left) or the final trained SOM (right) using heat maps coloured according to the accompanying scales. The bottom half of b expands the different distributions in the SOM for all expressed TSSs (left) or TSSs specifically expressed in two example cell lines, H1 hESC (centre) and HepG2 (right). c, The association of Gene Ontology (GO) terms on the same representation of the same trained SOM. We assigned genes that are within 20 kb of a genomic segment in a SOM unit to that unit, and then associated this set of genes with GO terms using a hypergeometric distribution after correcting for multiple testing. Map units that are significantly associated to GO terms are coloured green, with increasing strength of colour reflecting increasing numbers of genes significantly associated with the GO terms for either immune response (left) or sequence-specific transcription factor activity (centre). In each case, specific SOM units show association with these terms. The right-hand panel shows the distribution on the same SOM of all significantly associated GO terms, now colouring by GO term count per SOM unit. For sequence-specific transcription factor activity, two example genomic regions are extracted at the bottom of panel c from neighbouring SOM units. These are regions around the DBX1 (from SOM unit 26,31, left panel) and IRX6 (SOM unit 27,30, right panel) genes, respectively, along with their H3K27me3 ChIP-seq signal for each of the tier 1 and 2 cell types. For DBX1, representative of a set of primarily neuronal transcription factors associated with unit 26,31, there is a repressive H3K27me3 signal in both H1 hESCs and HUVECs; for IRX6, representative of a set of body patterning transcription factors associated with SOM unit 27,30, the repressive mark is restricted largely to the embryonic stem (ES) cell. An interactive version of this figure is available in the online version of the paper.
The classifications presented here are necessarily limited by the assays and cell lines studied, and probably contain a number of heterogeneous classes of elements. Nonetheless, robust classifications can be made, allowing a systematic view of the human genome.
Insights into human genomic variation
We next explored the potential impact of sequence variation on ENCODE functional elements. We examined allele-specific variation using results from the GM12878 cells that are derived from an individual (NA12878) sequenced in the 1000 Genomes project, along with her parents. Because ENCODE assays are predominantly sequence-based, the trio design allows each GM12878 data set to be divided by the specific parental contributions at heterozygous sites, producing aggregate haplotypic signals from multiple genomic sites. We examined 193 ENCODE assays for allele-specific biases using 1,409,992 phased, heterozygous SNPs and 167,096 insertions/deletions (indels) (Fig. 8). Alignment biases towards alleles present in the reference genome sequence were avoided using a sequence specifically tailored to the variants and haplotypes present in NA12878 (a ‘personalized genome’)72. We found instances of preferential binding towards each parental allele. For example, comparison of the results from the POLR2A, H3K79me2 and H3K27me3 assays in the region of NACC2 (Fig. 8a) shows a strong paternal bias for H3K79me2 and POL2RA and a strong maternal bias for H3K27me3, indicating differential activity for the maternal and paternal alleles.
a, Representative allele-specific information from GM12878 cells for selected assays around the first exon of the NACC2 gene (genomic region Chr9: 138950000–138995000, GRCh37). Transcription signal is shown in green, and the three sections show allele-specific data for three data sets (POLR2A, H3K79me2 and H3K27me3 ChIP-seq). In each case the purple signal is the processed signal for all sequence reads for the assay, whereas the blue and red signals show sequence reads specifically assigned to either the paternal or maternal copies of the genome, respectively. The set of common SNPs from dbSNP, including the phased, heterozygous SNPs used to provide the assignment, are shown at the bottom of the panel. NACC2 has a statistically significant paternal bias for POLR2A and the transcription-associated mark H3K79me2, and has a significant maternal bias for the repressive mark H3K27me3. b, Pair-wise correlations of allele-specific signal within single genes (below the diagonal) or within individual ChromHMM segments across the whole genome for selected DNase-seq and histone modification and transcription factor ChIP-seq assays. The extent of correlation is coloured according to the heat-map scale indicated from positive correlation (red) through to anti-correlation (blue). An interactive version of this figure is available in the online version of the paper.
Figure 8b shows the correlation of selected allele-specific signals across the whole genome. For instance, we found a strong allelic correlation between POL2RA and BCLAF1 binding, as well as negative correlation between H3K79me2 and H3K27me3, both at genes (Fig. 8b, below the diagonal, bottom left) and chromosomal segments (top right). Overall, we found that positive allelic correlations among the 193 ENCODE assays are stronger and more frequent than negative correlations. This may be due to preferential capture of accessible alleles and/or the specific histone modification and transcription factor, assays used in the project.
Rare variants, individual genomes and somatic variants
We further investigated the potential functional effects of individual variation in the context of ENCODE annotations. We divided NA12878 variants into common and rare classes, and partitioned these into those overlapping ENCODE annotation (Fig. 9a and Supplementary Tables 1 and 2, section K). We also predicted potential functional effects: for protein-coding genes, these are either non-synonymous SNPs or variants likely to induce loss of function by frame-shift, premature stop, or splice-site disruption; for other regions, these are variants that overlap a transcription-factor-binding site. We found similar numbers of potentially functional variants affecting protein-coding genes or affecting other ENCODE annotations, indicating that many functional variants within individual genomes lie outside exons of protein-coding genes. A more detailed analysis of regulatory variant annotation is described in ref. 73.
a, Breakdown of variants in a single genome (NA12878) by both frequency (common or rare (that is, variants not present in the low-coverage sequencing of 179 individuals in the pilot 1 European panel of the 1000 Genomes project55)) and by ENCODE annotation, including protein-coding gene and non-coding elements (GENCODE annotations for protein-coding genes, pseudogenes and other ncRNAs, as well as transcription-factor-binding sites from ChIP-seq data sets, excluding broad annotations such as histone modifications, segmentations and RNA-seq). Annotation status is further subdivided by predicted functional effect, being non-synonymous and missense mutations for protein-coding regions and variants overlapping bound transcription factor motifs for non-coding element annotations. A substantial proportion of variants are annotated as having predicted functional effects in the non-coding category. b, One of several relatively rare occurrences, where alignment to an individual genome sequence (paternal and maternal panels) shows a different readout from the reference genome. In this case, a paternal-haplotype-specific CTCF peak is identified. c, Relative level of somatic variants from a whole-genome melanoma sample that occur in DHSs unique to different cell lines. The coloured bars show cases that are significantly enriched or suppressed in somatic mutations. Details of ENCODE cell types can be found at http://encodeproject.org/ENCODE/cellTypes.html. An interactive version of this figure is available in the online version of the paper.
To study further the potential effects of NA12878 genome variants on transcription-factor-binding regions, we performed peak calling using a constructed personal diploid genome sequence for NA12878 (ref. 72). We aligned ChIP-seq sequences from GM12878 separately against the maternal and paternal haplotypes. As expected, a greater fraction of reads were aligned than to the reference genome (see Supplementary Information, Supplementary Fig. 1, section K). On average, approximately 1% of transcription-factor-binding sites in GM12878 cells are detected in a haplotype-specific fashion. For instance, Fig. 9b shows a CTCF-binding site not detected using the reference sequence that is only present on the paternal haplotype due to a 1-bp deletion (see also Supplementary Fig. 2, section K). As costs of DNA sequencing decrease further, optimized analysis of ENCODE-type data should use the genome sequence of the individual or cell being analysed when possible.
Most analyses of cancer genomes so far have focused on characterizing somatic variants in protein-coding regions. We intersected four available whole-genome cancer data sets with ENCODE annotations (Fig. 9c and Supplementary Fig. 2, section L). Overall, somatic variation is relatively depleted from ENCODE annotated regions, particularly for elements specific to a cell type matching the putative tumour source (for example, skin melanocytes for melanoma). Examining the mutational spectrum of elements in introns for cases where a strand-specific mutation assignment could be made reveals that there are mutational spectrum differences between DHSs and unannotated regions (0.06 Fisher’s exact test, Supplementary Fig. 3, section L). The suppression of somatic mutation is consistent with important functional roles of these elements within tumour cells, highlighting a potential alternative set of targets for examination in cancer.
Common variants associated with disease
In recent years, GWAS have greatly extended our knowledge of genetic loci associated with human disease risk and other phenotypes. The output of these studies is a series of SNPs (GWAS SNPs) correlated with a phenotype, although not necessarily the functional variants. Notably, 88% of associated SNPs are either intronic or intergenic74. We examined 4,860 SNP–phenotype associations for 4,492 SNPs curated in the National Human Genome Research Institute (NHGRI) GWAS catalogue74. We found that 12% of these SNPs overlap transcription-factor-occupied regions whereas 34% overlap DHSs (Fig. 10a). Both figures reflect significant enrichments relative to the overall proportions of 1000 Genomes project SNPs (about 6% and 23%, respectively). Even after accounting for biases introduced by selection of SNPs for the standard genotyping arrays, GWAS SNPs show consistently higher overlap with ENCODE annotations (Fig. 10a, see Supplementary Information). Furthermore, after partitioning the genome by density of different classes of functional elements, GWAS SNPs were consistently enriched beyond all the genotyping SNPs in function-rich partitions, and depleted in function-poor partitions (see Supplementary Fig. 1, section M). GWAS SNPs are particularly enriched in the segmentation classes associated with enhancers and TSSs across several cell types (see Supplementary Fig. 2, section M).
a, Overlap of lead SNPs in the NHGRI GWAS SNP catalogue (June 2011) with DHSs (left) or transcription-factor-binding sites (right) as red bars compared with various control SNP sets in blue. The control SNP sets are (from left to right): SNPs on the Illumina 2.5M chip as an example of a widely used GWAS SNP typing panel; SNPs from the 1000 Genomes project; SNPs extracted from 24 personal genomes (see personal genome variants track at http://main.genome-browser.bx.psu.edu (ref. 80)), all shown as blue bars. In addition, a further control used 1,000 randomizations from the genotyping SNP panel, matching the SNPs with each NHGRI catalogue SNP for allele frequency and distance to the nearest TSS (light blue bars with bounds at 1.5 times the interquartile range). For both DHSs and transcription-factor-binding regions, a larger proportion of overlaps with GWAS-implicated SNPs is found compared to any of the controls sets. b, Aggregate overlap of phenotypes to selected transcription-factor-binding sites (left matrix) or DHSs in selected cell lines (right matrix), with a count of overlaps between the phenotype and the cell line/factor. Values in blue squares pass an empirical P-value threshold ≤0.01 (based on the same analysis of overlaps between randomly chosen, GWAS-matched SNPs and these epigenetic features) and have at least a count of three overlaps. The P value for the total number of phenotype–transcription factor associations is <0.001. c, Several SNPs associated with Crohn’s disease and other inflammatory diseases that reside in a large gene desert on chromosome 5, along with some epigenetic features indicative of function. The SNP (rs11742570) strongly associated to Crohn’s disease overlaps a GATA2 transcription-factor-binding signal determined in HUVECs. This region is also DNase I hypersensitive in HUVECs and T-helper TH1 and TH2 cells. An interactive version of this figure is available in the online version of the paper.
Examining the SOM of integrated ENCODE annotations (see above), we found 19 SOM map units showing significant enrichment for GWAS SNPs, including many SOM units previously associated with specific gene functions, such as the immune response regions. Thus, an appreciable proportion of SNPs identified in initial GWAS scans are either functional or lie within the length of an ENCODE annotation (∼500 bp on average) and represent plausible candidates for the functional variant. Expanding the set of feasible functional SNPs to those in reasonable linkage disequilibrium, up to 71% of GWAS SNPs have a potential causative SNP overlapping a DNase I site, and 31% of loci have a candidate SNP that overlaps a binding site occupied by a transcription factor (see also refs 73, 75).
The GWAS catalogue provides a rich functional categorization from the precise phenotypes being studied. These phenotypic categorizations are nonrandomly associated with ENCODE annotations and there is marked correspondence between the phenotype and the identity of the cell type or transcription factor used in the ENCODE assay (Fig. 10b). For example, five SNPs associated with Crohn’s disease overlap GATA2-binding sites (P value 0.003 by random permutation or 0.001 by an empirical approach comparing to the GWAS-matched SNPs; see Supplementary Information), and fourteen are located in DHSs found in immunologically relevant cell types. A notable example is a gene desert on chromosome 5p13.1 containing eight SNPs associated with inflammatory diseases. Several are close to or within DHSs in T-helper type 1 (TH1) and TH2 cells as well as peaks of binding by transcription factors in HUVECs (Fig. 10c). The latter cell line is not immunological, but factor occupancy detected there could be a proxy for binding of a more relevant factor, such as GATA3, in T cells. Genetic variants in this region also affect expression levels of PTGER4 (ref. 76), encoding the prostaglandin receptor EP4. Thus, the ENCODE data reinforce the hypothesis that genetic variants in 5p13.1 modulate the expression of flanking genes, and furthermore provide the specific hypothesis that the variants affect occupancy of a GATA factor in an allele-specific manner, thereby influencing susceptibility to Crohn’s disease.
Nonrandom association of phenotypes with ENCODE cell types strengthens the argument that at least some of the GWAS lead SNPs are functional or extremely close to functional variants. Each of the associations between a lead SNP and an ENCODE annotation remains a credible hypothesis of a particular functional element class or cell type to explore with future experiments. Supplementary Tables 1–3, section M, list all 14,885 pairwise associations across the ENCODE annotations. The accompanying papers have a more detailed examination of common variants with other regulatory information19,25,29,73,75,77.
Concluding remarks
The unprecedented number of functional elements identified in this study provides a valuable resource to the scientific community as well as significantly enhances our understanding of the human genome. Our analyses have revealed many novel aspects of gene expression and regulation as well as the organization of such information, as illustrated by the accompanying papers (see http://www.encodeproject.org/ENCODE/pubs.html for collected ENCODE publications). However, there are still many specific details, particularly about the mechanistic processes that generate these elements and how and where they function, that require additional experiments to elucidate.
The large spread of coverage—from our highest resolution, most conservative set of bases implicated in GENCODE protein-coding gene exons (2.9%) or specific protein DNA binding (8.5%) to the broadest, most general set of marks covering the genome (approximately 80%), with many gradations in between—presents a spectrum of elements with different functional properties discovered by ENCODE. A total of 99% of the known bases in the genome are within 1.7 kb of any ENCODE element, whereas 95% of bases are within 8 kb of a bound transcription factor motif or DNase I footprint. Interestingly, even using the most conservative estimates, the fraction of bases likely to be involved in direct gene regulation, even though incomplete, is significantly higher than that ascribed to protein-coding exons (1.2%), raising the possibility that more information in the human genome may be important for gene regulation than for biochemical function. Many of the regulatory elements are not constrained across mammalian evolution, which so far has been one of the most reliable indications of an important biochemical event for the organism. Thus, our data provide orthologous indicators for suggesting possible functional elements.
Importantly, for the first time we have sufficient statistical power to assess the impact of negative selection on primate-specific elements, and all ENCODE classes display evidence of negative selection in these unique-to-primate elements. Furthermore, even with our most conservative estimate of functional elements (8.5% of putative DNA/protein binding regions) and assuming that we have already sampled half of the elements from our transcription factor and cell-type diversity, one would estimate that at a minimum 20% (17% from protein binding and 2.9% protein coding gene exons) of the genome participates in these specific functions, with the likely figure significantly higher.
The broad coverage of ENCODE annotations enhances our understanding of common diseases with a genetic component, rare genetic diseases, and cancer, as shown by our ability to link otherwise anonymous associations to a functional element. ENCODE and similar studies provide a first step towards interpreting the rest of the genome—beyond protein-coding genes—thereby augmenting common disease genetic studies with testable hypotheses. Such information justifies performing whole-genome sequencing (rather than exome only, 1.2% of the genome) on rare diseases and investigating somatic variants in non-coding functional elements, for instance, in cancer. Furthermore, as GWAS analyses typically associate disease to SNPs in large regions, comparison to ENCODE non-coding functional elements can help pinpoint putative causal variants in addition to refinement of location by fine-mapping techniques78. Combining ENCODE data with allele-specific information derived from individual genome sequences provides specific insight on the impact of a genetic variant. Indeed, we believe that a significant goal would be to use functional data such as that derived from this project to assign every genomic variant to its possible impact on human phenotypes.
So far, ENCODE has sampled 119 of 1,800 known transcription factors and general components of the transcriptional machinery on a limited number of cell types, and 13 of more than 60 currently known histone or DNA modifications across 147 cell types. DNase I, FAIRE and extensive RNA assays across subcellular fractionations have been undertaken on many cell types, but overall these data reflect a minor fraction of the potential functional information encoded in the human genome. An important future goal will be to enlarge this data set to additional factors, modifications and cell types, complementing the other related projects in this area (for example, Roadmap Epigenomics Project, http://www.roadmapepigenomics.org/, and International Human Epigenome Consortium, http://www.ihec-epigenomes.org/). These projects will constitute foundational resources for human genomics, allowing a deeper interpretation of the organization of gene and regulatory information and the mechanisms of regulation, and thereby provide important insights into human health and disease. Co-published ENCODE-related papers can be explored online via the Nature ENCODE explorer (http://www.nature.com/ENCODE), a specially designed visualization tool that allows users to access the linked papers and investigate topics that are discussed in multiple papers via thematically organized threads.
Methods Summary
For full details of Methods, see Supplementary Information.
References
ENCODE Project Consortium. The ENCODE (ENCyclopedia Of DNA Elements) Project. Science 306, 636–640 (2004)
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007)
The ENCODE Project Consortium. A user’s guide to the encyclopedia of DNA elements (ENCODE). PLoS Biol. 9, e1001046 (2011)
Mouse Genome Sequencing Consortium. Initial sequencing and comparative analysis of the mouse genome. Nature 420, 520–562 (2002)
Chiaromonte, F. et al. The share of human genomic DNA under selection estimated from human-mouse genomic alignments. Cold Spring Harb. Symp. Quant. Biol. 68, 245–254 (2003)
Cooper, G. M. et al. Distribution and intensity of constraint in mammalian genomic sequence. Genome Res. 15, 901–913 (2005)
Parker, S. C., Hansen, L., Abaan, H. O., Tullius, T. D. & Margulies, E. H. Local DNA topography correlates with functional noncoding regions of the human genome. Science 324, 389–392 (2009)
Lindblad-Toh, K. et al. A high-resolution map of human evolutionary constraint using 29 mammals. Nature 478, 476–482 (2011)
Pheasant, M. & Mattick, J. S. Raising the estimate of functional human sequences. Genome Res. 17, 1245–1253 (2007)
Ponting, C. P. & Hardison, R. C. What fraction of the human genome is functional? Genome Res. 21, 1769–1776 (2011)
Asthana, S. et al. Widely distributed noncoding purifying selection in the human genome. Proc. Natl Acad. Sci. USA 104, 12410–12415 (2007)
Landt, S. G. et al. ChIP-seq guidelines and practices used by the ENCODE and modENCODE consortia. Genome Res. http://dx.doi.org/10.1101/gr.136184.111 (2012)
Li, Q., Brown, J. B., Huang, H. & Bickel, P. J. Measuring reproducibility of high-throughput experiments. Ann. Appl. Stat. 5, 1752–1779 (2011)
Harrow, J. et al. GENCODE: The reference human genome annotation for the ENCODE project. Genome Res. http://dx.doi.org/10.1101/gr.135350.111 (2012)
Howald, C. et al. Combining RT-PCR-seq and RNA-seq to catalog all genic elements encoded in the human genome. Genome Res. http://dx.doi.org/10.1101/gr.134478.111 (2012)
Djebali, S. et al. Landscape of transcription in human cells. Naturehttp://dx.doi.org/10.1038/nature11233 (this issue)
Derrien, T. et al. The GENCODE v7 catalog of human long noncoding RNAs: Analysis of their gene structure, evolution, and expression. Genome Res. http://dx.doi.org/10.1101/gr.132159.111 (2012)
Pei, B. et al. The GENCODE pseudogene resource. Genome Biol. 13, R51 (2012)
Gerstein, M. B. et al. Architecture of the human regulatory network derived from ENCODE data. Nature http://dx.doi.org/10.1038/nature11245 (this issue)
Bickel, P. J., Boley, N., Brown, J. B., Huang, H. Y. & Zhang, N. R. Subsampling methods for genomic inference. Ann. Appl. Stat. 4, 1660–1697 (2010)
Kaplan, T. et al. Quantitative models of the mechanisms that control genome-wide patterns of transcription factor binding during early Drosophila development. PLoS Genet. 7, e1001290 (2011)
Li, X. Y. et al. The role of chromatin accessibility in directing the widespread, overlapping patterns of Drosophila transcription factor binding. Genome Biol. 12, R34 (2011)
Pique-Regi, R. et al. Accurate inference of transcription factor binding from DNA sequence and chromatin accessibility data. Genome Res. 21, 447–455 (2011)
Zhang, Y. et al. Primary sequence and epigenetic determinants of in vivo occupancy of genomic DNA by GATA1. Nucleic Acids Res. 37, 7024–7038 (2009)
Neph, S. et al. An expansive human regulatory lexicon encoded in transcription factor footprints. Nature http://dx.doi.org/10.1038/nature11212 (this issue)
Whitfield, T. W. et al. Functional analysis of transcription factor binding sites in human promoters. Genome Biol. 13, R50 (2012)
Gross, D. S. & Garrard, W. T. Nuclease hypersensitive sites in chromatin. Annu. Rev. Biochem. 57, 159–197 (1988)
Urnov, F. D. Chromatin remodeling as a guide to transcriptional regulatory networks in mammals. J. Cell. Biochem. 88, 684–694 (2003)
Thurman, R. E. et al. The accessible chromatin landscape of the human genome. Naturehttp://dx.doi.org/10.1038/nature11232 (this issue)
Kundaje, A. et al. Ubiquitous heterogeneity and asymmetry of the chromatin environment at regulatory elements. Genome Res. http://dx.doi.org/10.1101/gr.136366.111 (2012)
Schultz, D. C., Ayyanathan, K., Negorev, D., Maul, G. G. & Rauscher, F. J., III SETDB1: a novel KAP-1-associated histone H3, lysine 9-specific methyltransferase that contributes to HP1-mediated silencing of euchromatic genes by KRAB zinc-finger proteins. Genes Dev. 16, 919–932 (2002)
Frietze, S., O’Geen, H., Blahnik, K. R., Jin, V. X. & Farnham, P. J. ZNF274 recruits the histone methyltransferase SETDB1 to the 3′ ends of ZNF genes. PLoS ONE 5, e15082 (2010)
Boyle, A. P. et al. High-resolution genome-wide in vivo footprinting of diverse transcription factors in human cells. Genome Res. 21, 456–464 (2011)
Hesselberth, J. R. et al. Global mapping of protein-DNA interactions in vivo by digital genomic footprinting. Nature Methods 6, 283–289 (2009)
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008)
Kouzarides, T. Chromatin modifications and their function. Cell 128, 693–705 (2007)
Li, B., Carey, M. & Workman, J. L. The role of chromatin during transcription. Cell 128, 707–719 (2007)
Hon, G. C., Hawkins, R. D. & Ren, B. Predictive chromatin signatures in the mammalian genome. Hum. Mol. Genet. 18, R195–R201 (2009)
Zhou, V. W., Goren, A. & Bernstein, B. E. Charting histone modifications and the functional organization of mammalian genomes. Nature Rev. Genet. 12, 7–18 (2011)
Ernst, J. et al. Mapping and analysis of chromatin state dynamics in nine human cell types. Nature 473, 43–49 (2011)
Hon, G., Wang, W. & Ren, B. Discovery and annotation of functional chromatin signatures in the human genome. PLoS Comput. Biol. 5, e1000566 (2009)
Ball, M. P. et al. Targeted and genome-scale strategies reveal gene-body methylation signatures in human cells. Nature Biotechnol. 27, 361–368 (2009)
Meissner, A. et al. Genome-scale DNA methylation maps of pluripotent and differentiated cells. Nature 454, 766–770 (2008)
Ogryzko, V. V., Schiltz, R. L., Russanova, V., Howard, B. H. & Nakatani, Y. The transcriptional coactivators p300 and CBP are histone acetyltransferases. Cell 87, 953–959 (1996)
Lister, R. et al. Human DNA methylomes at base resolution show widespread epigenomic differences. Nature 462, 315–322 (2009)
Dekker, J. Gene regulation in the third dimension. Science 319, 1793–1794 (2008)
Dostie, J. et al. Chromosome Conformation Capture Carbon Copy (5C): a massively parallel solution for mapping interactions between genomic elements. Genome Res. 16, 1299–1309 (2006)
Lajoie, B. R., van Berkum, N. L., Sanyal, A. & Dekker, J. My5C: web tools for chromosome conformation capture studies. Nature Methods 6, 690–691 (2009)
Sanyal, A., Lajoie, B., Jain, G. & Dekker, J. The long-range interaction landscape of gene promoters. Nature http://dx.doi.org/10.1038/nature11279 (this issue)
Fullwood, M. J. et al. An oestrogen-receptor-alpha-bound human chromatin interactome. Nature 462, 58–64 (2009)
Li, G. et al. Extensive promoter-centered chromatin interactions provide a topological basis for transcription regulation. Cell 148, 84–98 (2012)
Borneman, A. R. et al. Divergence of transcription factor binding sites across related yeast species. Science 317, 815–819 (2007)
Odom, D. T. et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nature Genet. 39, 730–732 (2007)
Schmidt, D. et al. Five-vertebrate ChIP-seq reveals the evolutionary dynamics of transcription factor binding. Science 328, 1036–1040 (2010)
A map of human genome variation from population-scale sequencing. Nature 467, 1061–1073 (2010)
King, M. C. & Wilson, A. C. Evolution at two levels in humans and chimpanzees. Science 188, 107–116 (1975)
Spivakov, M. et al. Analysis of variation at transcription factor binding sites in Drosophila and humans. Genome Biol. 13, R49 (2012)
Sandelin, A. et al. Mammalian RNA polymerase II core promoters: insights from genome-wide studies. Nature Rev. Genet. 8, 424–436 (2007)
Dong, X. et al. Modeling gene expression using chromatin features in various cellular contexts. Genome Biol. 13, R53 (2012)
Huff, J. T., Plocik, A. M., Guthrie, C. & Yamamoto, K. R. Reciprocal intronic and exonic histone modification regions in humans. Nature Struct. Mol. Biol. 17, 1495–1499 (2010)
Tilgner, H. et al. Deep sequencing of subcellular RNA fractions shows splicing to be predominantly co-transcriptional in the human genome but inefficient for lncRNAs. Genome Res. http://dx.doi.org/10.1101/gr.134445.111 (2012)
Fu, Y., Sinha, M., Peterson, C. L. & Weng, Z. The insulator binding protein CTCF positions 20 nucleosomes around its binding sites across the human genome. PLoS Genet. 4, e1000138 (2008)
Kornberg, R. D. & Stryer, L. Statistical distributions of nucleosomes: nonrandom locations by a stochastic mechanism. Nucleic Acids Res. 16, 6677–6690 (1988)
Schones, D. E. et al. Dynamic regulation of nucleosome positioning in the human genome. Cell 132, 887–898 (2008)
Valouev, A. et al. Determinants of nucleosome organization in primary human cells. Nature 474, 516–520 (2011)
Frietze, S. et al. Cell type-specific binding patterns reveal that TCF7L2 can be tethered to the genome by association with GATA3. Genome Biol. 13, R52 (2012)
Yip, K. Y. et al. Classification of human genomic regions based on experimentally-determined binding sites of more than 100 transcription-related factors. Genome Biol. 13, R48 (2012)
Hoffman, M. M. et al. Unsupervised pattern discovery in human chromatin structure through genomic segmentation. Nature Methods 9, 473–476 (2012)
Kapranov, P. et al. RNA maps reveal new RNA classes and a possible function for pervasive transcription. Science 316, 1484–1488 (2007)
Koch, F. et al. Transcription initiation platforms and GTF recruitment at tissue-specific enhancers and promoters. Nature Struct. Mol. Biol. 18, 956–963 (2011)
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nature Biotechnol. 28, 495–501 (2010)
Rozowsky, J. et al. AlleleSeq: analysis of allele-specific expression and binding in a network framework. Mol. Syst. Biol. 7, 522 (2011)
Boyle, A. P. et al. Annotation of functional variation in personal genomes using RegulomeDB. Genome Res. http://dx.doi.org/10.1101/gr.137323.112 (2012)
Hindorff, L. A. et al. Potential etiologic and functional implications of genome-wide association loci for human diseases and traits. Proc. Natl Acad. Sci. USA 106, 9362–9367 (2009)
Schaub, M. A., Boyle, A. P., Kundaje, A., Batzoglou, S. & Snyder, M. Linking disease associations with regulatory information in the human genome. Genome Res. http://dx.doi.org/10.1101/gr.136127.111 (2012)
Libioulle, C. et al. Novel Crohn disease locus identified by genome-wide association maps to a gene desert on 5p13.1 and modulates expression of PTGER4. PLoS Genet. 3, e58 (2007)
Vernot, B. et al. Personal and population genomics of human regulatory variation. Genome Res. http://dx.doi.org/10.1101/gr.134890.111 (2012)
Harismendy, O. et al. 9p21 DNA variants associated with coronary artery disease impair interferon-γ signalling response. Nature 470, 264–268 (2011)
Cheng, C. et al. Understanding transcriptional regulation by integrative analysis of transcription factor binding data. Genome Res. http://dx.doi.org/10.1101/gr.136838.111 (2012)
Schuster, S. C. et al. Complete Khoisan and Bantu genomes from southern Africa. Nature 463, 943–947 (2010)
Acknowledgements
We thank additional members of our laboratories and institutions who have contributed to the experimental and analytical components of this project. We thank D. Leja for assistance with production of the figures. The Consortium is funded by grants from the NHGRI as follows: production grants: U54HG004570 (B. E. Bernstein); U01HG004695 (E. Birney); U54HG004563 (G. E. Crawford); U54HG004557 (T. R. Gingeras); U54HG004555 (T. J. Hubbard); U41HG004568 (W. J. Kent); U54HG004576 (R. M. Myers); U54HG004558 (M. Snyder); U54HG004592 (J. A. Stamatoyannopoulos). Pilot grants: R01HG003143 (J. Dekker); RC2HG005591 and R01HG003700 (M. C. Giddings); R01HG004456-03 (Y. Ruan); U01HG004571 (S. A. Tenenbaum); U01HG004561 (Z. Weng); RC2HG005679 (K. P. White). This project was supported in part by American Recovery and Reinvestment Act (ARRA) funds from the NHGRI through grants U54HG004570, U54HG004563, U41HG004568, U54HG004592, R01HG003143, RC2HG005591, R01HG003541, U01HG004561, RC2HG005679 and R01HG003988 (L. Pennacchio). In addition, work from NHGRI Groups was supported by the Intramural Research Program of the NHGRI (L. Elnitski, ZIAHG200323; E. H. Margulies, ZIAHG200341). Research in the Pennachio laboratory was performed at Lawrence Berkeley National Laboratory and at the United States Department of Energy Joint Genome Institute, Department of Energy Contract DE-AC02-05CH11231, University of California.
Author information
Authors and Affiliations
Consortia
Contributions
See the consortium author list for details of author contributions.
Corresponding author
Ethics declarations
Competing interests
The author declare no competing financial interests.
Supplementary information
Supplementary Information 1
This file contains Supplementary Text and Data, Methods and References – see Contents list for full details. (PDF 1875 kb)
Supplementary Table 1-section U
This data fie shows the GENCODE Gene Annotation Statistics. (XLS 20 kb)
Supplementary Movie 1
This video shows the transient expression of a piece of human DNA predicted to an enhancer in a Medaka fish embryo. The prediction is from k562 human cell line, which is a red blood cell precursor derived line. The expression in Medaka is in the mature red blood cells (these are nucleated cells in fish). The expression is green fluorescent protein. The video was made in the Wittbrodt laboratory at the University of Heidelberg. (MOV 930 kb)
Supplementary Movie 2
This video shows the transient expression of a piece of human DNA predicted to an enhancer in a Medaka fish embryo. The prediction is from k562 human cell line, which is a red blood cell precursor derived line. The expression in Medaka is in the mature red blood cells (these are nucleated cells in fish). The expression is green fluorescent protein. The video was made in the Wittbrodt laboratory at the University of Heidelberg. (MOV 637 kb)
Supplementary Information 2
This file contains Supplementary Figures E1-E6, K1-K2, L1-L3, M1-M2, R1-R2, Y1 and Z1, Supplementary Tables E1-E2, K1-K2 and L1 and additional references. (PDF 2511 kb)
Supplementary Table 1-section F
This data file shows the TF Co‐associations. (XLS 2170 kb)
Supplementary Table 1-section M
This data file shows the GWAS SNP phenotype associations across TF and DHS ENCODE annotations. (XLS 609 kb)
Supplementary Table 2-section M
This file contains the GWAS SNP pair‐wise associations across DHS ENCODE annotations. (TXT 907 kb)
Supplementary Table 3-section M
This file contains the GWAS SNP pair‐wise associations across TF ENCODE annotations. (TXT 2395 kb)
Supplementary Table 1-section N
This data file shows the ENCODE TF Classification in detail. (XLS 34 kb)
Supplementary Table 1-section P
This data file shows the ENCODE Data Production Summary. (XLS 38 kb)
Supplementary Table 1-section Q
This data file shows the ENCODE Element Counts and Lengths by Data Type. (XLS 323 kb)
Rights and permissions
This article is distributed under the terms of the Creative Commons Attribution-Non-Commercial-Share Alike licence (http://creativecommons.org/licenses/by-nc-sa/3.0/).
About this article
Cite this article
The ENCODE Project Consortium. An integrated encyclopedia of DNA elements in the human genome. Nature 489, 57–74 (2012). https://doi.org/10.1038/nature11247
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nature11247
This article is cited by
-
Analysis of REST binding sites with canonical and non-canonical motifs in human cell lines
BMC Medical Genomics (2024)
-
Conserved role of hnRNPL in alternative splicing of epigenetic modifiers enables B cell activation
EMBO Reports (2024)
-
Transposable elements-mediated recruitment of KDM1A epigenetically silences HNF4A expression to promote hepatocellular carcinoma
Nature Communications (2024)
-
Decoding chromatin states by proteomic profiling of nucleosome readers
Nature (2024)
-
Advances in the multimodal analysis of the 3D chromatin structure and gene regulation
Experimental & Molecular Medicine (2024)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.