Abstract
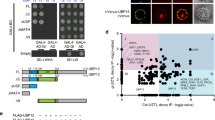
The underlying mechanism of circadian rhythmicity appears to be conserved among organisms, and is based on negative transcriptional feedback loops forming a cellular oscillator (or ‘clock’)1,2. Circadian changes in protein stability, phosphorylation and subcellular localization also contribute to the generation and maintenance of this clock1,2. In plants, several genes have been shown to be closely associated with the circadian system3,4. However, the molecular mechanisms proposed to regulate the plant clock are mostly based on regulation at the transcriptional level3,4. Here we provide genetic and molecular evidence for a role of ZEITLUPE (ZTL)5,6,7 in the targeted degradation of TIMING OF CAB EXPRESSION 1 (TOC1)8,9 in Arabidopsis thaliana (thale cress). The physical interaction of TOC1 with ZTL is abolished by the ztl-1 mutation, resulting in constitutive levels of TOC1 protein expression. The dark-dependent degradation of TOC1 protein requires functional ZTL, and is prevented by inhibiting the proteosome pathway. Our results show that the TOC1–ZTL interaction is important in the control of TOC1 protein stability, and is probably responsible for the regulation of circadian period by the clock.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Harmer, S. L., Panda, S. & Kay, S. A. Molecular bases of circadian rhythms. Annu. Rev. Cell Dev. Biol. 17, 215–253 (2001)
Young, M. W. & Kay, S. A. Time zones: A comparative genetics of circadian clocks. Nature Rev. Genet. 2, 702–715 (2001)
McClung, C. R. Circadian rhythms in plants. Annu. Rev. Plant Physiol. Plant Mol. Biol. 52, 139–162 (2001)
Yanovsky, M. J. & Kay, S. A. Signaling networks in the plant circadian system. Curr. Opin. Plant Biol. 4, 429–435 (2001)
Somers, D. E., Schultz, T. F., Milnamow, M. & Kay, S. A. ZEITLUPE encodes a novel clock-associated PAS protein from Arabidopsis. Cell 101, 319–329 (2000)
Jarillo, J. A. et al. An Arabidopsis circadian clock component interacts with both CRY1 and phyB. Nature 410, 487–490 (2001)
Kiyosue, T. & Wada, M. LKP1 (LOV kelch protein 1): a factor involved in the regulation of flowering time in Arabidopsis. Plant J. 23, 807–815 (2000)
Strayer, C. A. et al. Cloning of the Arabidopsis clock gene TOC1, an autoregulatory response regulator homolog. Science 289, 768–771 (2000)
Makino, S., Matsushika, A., Kojima, M., Yamashino, T. & Mizuno, T. The APRR1/TOC1 quintet implicated in circadian rhythms of Arabidopsis thaliana: Characterization with APRR1-overexpressing plants. Plant Cell Physiol. 43, 58–69 (2002)
Alabadí, D. et al. Reciprocal regulation between TOC1 and LHY/CCA1 within the Arabidopsis circadian clock. Science 293, 880–883 (2001)
Wang, Z. Y. & Tobin, E. M. Constitutive expression of the CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) gene disrupts circadian rhythms and suppresses its own expression. Cell 93, 1207–1217 (1998)
Schaffer, R. et al. The late elongated hypocotyl mutation of Arabidopsis disrupts circadian rhythms and the photoperiodic control of flowering. Cell 93, 1219–1229 (1998)
Más, P., Alabadi, D., Yanovsky, M. J., Oyama, T. & Kay, S. A. Dual role of TOC1 in the control of circadian and photomorphogenic responses in Arabidopsis. Plant Cell 15, 223–236 (2003)
Schultz, T. F., Kiyosue, T., Yanovsky, M., Wada, M. & Kay, S. A. A role for LKP2 in the circadian clock of Arabidopsis. Plant Cell 13, 2659–2670 (2001)
Cheng, P., He, Q., Yang, Y., Wang, L. & Liu, Y. Functional conservation of light, oxygen, or voltage domains in light sensing. Proc. Natl Acad. Sci. USA 100, 5938–5943 (2003)
Briggs, W. R. & Christie, J. M. Phototropins 1 and 2: Versatile plant blue-light receptors. Trends Plant Sci. 7, 201–210 (2002)
Deshaies, R. J. SCF and cullin/RING H2-based ubiquitin ligases. Annu. Rev. Cell Dev. Biol. 15, 435–467 (1999)
Risseeuw, E. P. et al. Protein interaction analysis of SCF ubiquitin E3 ligase subunits from Arabidopsis. Plant J. 34, 753–767 (2003)
Taylor, B. L. & Zhulin, I. B. PAS Domains: Internal sensors of oxygen, redox potential, and light. Microbiol. Mol. Biol. Rev. 63, 479–506 (1999)
Froehlich, A. C., Liu, Y., Loros, J. J. & Dunlap, J. C. White Collar-1, a circadian blue light photoreceptor, binding to the frequency promoter. Science 297, 815–819 (2002)
Plautz, J. D. et al. Quantitative analysis of Drosophila period gene transcription in living animals. J. Biol. Rhythms 12, 204–217 (1997)
Millar, A. J., Straume, M., Chory, J., Chua, N. H. & Kay, S. A. The regulation of circadian period by phototransduction pathways in Arabidopsis. Science 267, 1163–1166 (1995)
Kim, W.-Y., Geng, R. & Somers, D. E. Circadian phase-specific degradation of the F-box protein ZTL is mediated by the proteasome. Proc. Natl Acad. Sci. USA 100, 4933–4938 (2003)
Somers, D. E., Devlin, P. F. & Kay, S. A. Phytochromes and cryptochromes in the entrainment of the Arabidopsis circadian clock. Science 282, 1488–1490 (1998)
Más, P., Devlin, P. F., Panda, S. & Kay, S. A. Functional interaction of phytochrome B and cryptochrome 2. Nature 408, 207–211 (2000)
Yagita, K. et al. Nucleocytoplasmic shuttling and mCRY-dependent inhibition of ubiquitylation of the mPER2 clock protein. EMBO J. 21, 1301–1314 (2002)
Ko, H. W., Jiang, J. & Edery, I. Role for Slimb in the degradation of Drosophila Period protein phosphorylated by Doubletime. Nature 420, 673–678 (2002)
Grima, B. et al. The F-box protein slimb controls the levels of clock proteins period and timeless. Nature 420, 178–182 (2002)
Millar, A. J., Carre, I. A., Strayer, C. A., Chua, N. H. & Kay, S. A. Circadian clock mutants in Arabidopsis identified by luciferase imaging. Science 267, 1161–1163 (1995)
Somers, D. E., Webb, A. A. R., Pearson, M. & Kay, S. A. The short-period mutant toc1–1, alters circadian clock regulation of multiple outputs throughout development in Arabidopsis thaliana. Development 125, 485–494 (1998)
Acknowledgements
We thank H. Tran, T. Imaizumi and S. Hazen for critical reading of the manuscript. This research was supported by the NIH (S.A.K.) and the NSF (D.E.S).
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare that they have no competing financial interests.
Rights and permissions
About this article
Cite this article
Más, P., Kim, WY., Somers, D. et al. Targeted degradation of TOC1 by ZTL modulates circadian function in Arabidopsis thaliana. Nature 426, 567–570 (2003). https://doi.org/10.1038/nature02163
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/nature02163
This article is cited by
-
Flowering in sugarcane-insights from the grasses
3 Biotech (2023)
-
Noise reduction by upstream open reading frames
Nature Plants (2022)
-
Molecular insights into the circadian clock in marine diatoms
Acta Oceanologica Sinica (2022)
-
Variations in Circadian Clock Organization & Function: A Journey from Ancient to Recent
Planta (2022)
-
Environment-mediated mutagenetic interference on genetic stabilization and circadian rhythm in plants
Cellular and Molecular Life Sciences (2022)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.