Abstract
Cryptochromes (CRYs) are photoreceptors that mediate light regulation of the circadian clock in plants and animals. Here we show that CRYs mediate blue-light regulation of N6-methyladenosine (m6A) modification of more than 10% of messenger RNAs in the Arabidopsis transcriptome, especially those regulated by the circadian clock. CRY2 interacts with three subunits of the METTL3/14-type N6-methyladenosine RNA methyltransferase (m6A writer): MTA, MTB and FIP37. Photo-excited CRY2 undergoes liquid–liquid phase separation (LLPS) to co-condense m6A writer proteins in vivo, without obviously altering the affinity between CRY2 and the writer proteins. mta and cry1cry2 mutants share common defects of a lengthened circadian period, reduced m6A RNA methylation and accelerated degradation of mRNA encoding the core component of the molecular oscillator circadian clock associated 1 (CCA1). These results argue for a photoregulatory mechanism by which light-induced phase separation of CRYs modulates m6A writer activity, mRNA methylation and abundance, and the circadian rhythms in plants.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 digital issues and online access to articles
$119.00 per year
only $9.92 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Data availability
All data supporting the findings of this study are available in the main text or the supplementary table. Biological materials used in this study are available from C.L. on reasonable request. The m6A-seq and RNA-seq data reported in this study have been deposited in the NCBI Gene Expression Omnibus under accession number GSE152466 which is fully available. The RNA-seq data for WT and cry1cry2 sample in the dark condition have been reported previously (GSE80350)26. Source data are provided with this paper.
Change history
22 October 2021
A Correction to this paper has been published: https://doi.org/10.1038/s41477-021-01027-4
References
Sanchez, S. E., Rugnone, M. L. & Kay, S. A. Light perception: a matter of time. Mol. Plant 13, 363–385 (2020).
Patke, A., Young, M. W. & Axelrod, S. Molecular mechanisms and physiological importance of circadian rhythms. Nat. Rev. Mol. Cell Biol. 21, 67–84 (2020).
Webb, A. A. R., Seki, M., Satake, A. & Caldana, C. Continuous dynamic adjustment of the plant circadian oscillator. Nat. Commun. 10, 550 (2019).
Wang, Q. & Lin, C. Mechanisms of cryptochrome-mediated photoresponses in plants. Annu. Rev. Plant Biol. 71, 103–129 (2020).
Cashmore, A. R. Cryptochromes: enabling plants and animals to determine circadian time. Cell 114, 537–543 (2003).
Sancar, A. Cryptochrome: the second photoactive pigment in the eye and its role in circadian photoreception. Annu. Rev. Biochem. 69, 31–67 (2000).
Bailey-Serres, J., Zhai, J. & Seki, M. The dynamic kaleidoscope of RNA biology in plants. Plant Physiol. 182, 1–9 (2020).
Parker, M. T. et al. Nanopore direct RNA sequencing maps the complexity of Arabidopsis mRNA processing and m(6)A modification. eLife; https://doi.org/10.7554/eLife.49658 (2020).
Wang, C. Y., Yeh, J. K., Shie, S. S., Hsieh, I. C. & Wen, M. S. Circadian rhythm of RNA N6-methyladenosine and the role of cryptochrome. Biochem. Biophys. Res. Commun. 465, 88–94 (2015).
Fustin, J. M. et al. RNA-methylation-dependent RNA processing controls the speed of the circadian clock. Cell 155, 793–806 (2013).
Anderson, S. J. et al. N6-Methyladenosine inhibits local ribonucleolytic cleavage to stabilize mRNAs in Arabidopsis. Cell Rep. 25, 1146–1157 e1143 (2018).
Wang, X. et al. N6-methyladenosine-dependent regulation of messenger RNA stability. Nature 505, 117–120 (2014).
Zaccara, S., Ries, R. J. & Jaffrey, S. R. Reading, writing and erasing mRNA methylation. Nat. Rev. Mol. Cell Biol. 20, 608–624 (2019).
Shen, L., Liang, Z., Wong, C. E. & Yu, H. Messenger RNA modifications in plants. Trends Plant Sci. 24, 328–341 (2019).
Zhao, B. S., Roundtree, I. A. & He, C. Post-transcriptional gene regulation by mRNA modifications. Nat. Rev. Mol. Cell Biol. 18, 31–42 (2017).
Dominissini, D., Moshitch-Moshkovitz, S., Salmon-Divon, M., Amariglio, N. & Rechavi, G. Transcriptome-wide mapping of N(6)-methyladenosine by m(6)A-seq based on immunocapturing and massively parallel sequencing. Nat. Protoc. 8, 176–189 (2013).
Wang, Z. Y. & Tobin, E. M. Constitutive expression of the CIRCADIAN CLOCK ASSOCIATED 1 (CCA1) gene disrupts circadian rhythms and suppresses its own expression. Cell 93, 1207–1217 (1998).
Liu, Q. et al. Molecular basis for blue light-dependent phosphorylation of Arabidopsis cryptochrome 2. Nat. Commun. 8, 15234 (2017).
Arribas-Hernandez, L. & Brodersen, P. Occurrence and functions of m(6)A and other covalent modifications in plant mRNA. Plant Physiol. 182, 79–96 (2020).
Ruzicka, K. et al. Identification of factors required for m6A mRNA methylation in Arabidopsis reveals a role for the conserved E3 ubiquitin ligase HAKAI. New Phytol. 215, 157–172 (2017).
Zhong, S. et al. MTA is an Arabidopsis messenger RNA adenosine methylase and interacts with a homolog of a sex-specific splicing factor. Plant Cell 20, 1278–1288 (2008).
Chen, Y. et al. Regulation of Arabidopsis photoreceptor CRY2 by two distinct E3 ubiquitin ligases. Nat. Commun. 12, 2155 (2021).
Yu, X. et al. Arabidopsis cryptochrome 2 completes its posttranslational life cycle in the nucleus. Plant Cell 19, 3146–3156 (2007).
Liu, Q. et al. Photooligomerization determines photosensitivity and photoreactivity of plant cryptochromes. Mol. Plant 13, 398–413 (2020).
Liu, H. et al. Photoexcited CRY2 interacts with CIB1 to regulate transcription and floral initiation in Arabidopsis. Science 322, 1535–1539 (2008).
Wang, Q. et al. Photoactivation and inactivation of Arabidopsis cryptochrome 2. Science 354, 343–347 (2016).
Yu, X. et al. Formation of nuclear bodies of Arabidopsis CRY2 in response to blue light is associated with its blue light-dependent degradation. Plant Cell 21, 118–130 (2009).
Mas, P., Devlin, P. F., Panda, S. & Kay, S. A. Functional interaction of phytochrome B and cryptochrome 2. Nature 408, 207–211 (2000).
Sabari, B. R. et al. Coactivator condensation at super-enhancers links phase separation and gene control. Science https://doi.org/10.1126/science.aar3958 (2018).
Bracha, D. et al. Mapping local and global liquid phase behavior in living cells using photo-oligomerizable seeds. Cell 175, 1467–1480 e1413 (2018).
Shin, Y. et al. Spatiotemporal control of intracellular phase transitions using light-activated optodroplets. Cell 168, 159–171 e114 (2017).
Dignon, G. L., Best, R. B. & Mittal, J. Biomolecular phase separation: from molecular driving forces to macroscopic properties. Annu. Rev. Phys. Chem. 71, 53–75 (2020).
Alberti, S., Gladfelter, A. & Mittag, T. Considerations and challenges in studying liquid–liquid phase separation and biomolecular condensates. Cell 176, 419–434 (2019).
Wang, Q. et al. The blue light-dependent phosphorylation of the CCE domain determines the photosensitivity of Arabidopsis CRY2. Mol. Plant 8, 631–643 (2015).
Shalitin, D. et al. Regulation of Arabidopsis cryptochrome 2 by blue-light-dependent phosphorylation. Nature 417, 763–767 (2002).
Devlin, P. F. & Kay, S. A. Cryptochromes are required for phytochrome signaling to the circadian clock but not for rhythmicity. Plant Cell 12, 2499–2510 (2000).
Somers, D. E., Devlin, P. F. & Kay, S. A. Phytochromes and cryptochromes in the entrainment of the Arabidopsis circadian clock. Science 282, 1488–1490 (1998).
Bodi, Z. et al. Adenosine methylation in Arabidopsis mRNA is associated with the 3ʹ end and reduced levels cause developmental defects. Front. Plant Sci. 3, 48 (2012).
Guo, H., Yang, H., Mockler, T. C. & Lin, C. Regulation of flowering time by Arabidopsis photoreceptors. Science 279, 1360–1363 (1998).
Legris, M., Ince, Y. C. & Fankhauser, C. Molecular mechanisms underlying phytochrome-controlled morphogenesis in plants. Nat. Commun. 10, 5219 (2019).
Leivar, P. & Quail, P. H. PIFs: pivotal components in a cellular signaling hub. Trends Plant Sci. 16, 19–28 (2011).
Bhat, S. S. et al. mRNA adenosine methylase (MTA) deposits m(6)A on pri-miRNAs to modulate miRNA biogenesis in Arabidopsis thaliana. Proc. Natl Acad. Sci. USA 117, 21785–21795 (2020).
Aschoff, J. Exogenous and endogenous components in circadian rhythms. Cold Spring Harb. Symp. Quant. Biol. 25, 11–28 (1960).
Jang, G. J., Yang, J. Y., Hsieh, H. L. & Wu, S. H. Processing bodies control the selective translation for optimal development of Arabidopsis young seedlings. Proc. Natl Acad. Sci. USA 116, 6451–6456 (2019).
Godoy Herz, M. A. et al. Light regulates plant alternative splicing through the control of transcriptional elongation. Mol. Cell 73, 1066–1074 e1063 (2019).
Wu, S. H. Gene expression regulation in photomorphogenesis from the perspective of the central dogma. Annu. Rev. Plant Biol. 65, 311–333 (2014).
Paik, I., Yang, S. & Choi, G. Phytochrome regulates translation of mRNA in the cytosol. Proc. Natl Acad. Sci. USA 109, 1335–1340 (2012).
Juntawong, P. & Bailey-Serres, J. Dynamic light regulation of translation status in Arabidopsis thaliana. Front. Plant Sci. 3, 66 (2012).
Wang, X. et al. SKIP is a component of the spliceosome linking alternative splicing and the circadian clock in Arabidopsis. Plant Cell 24, 3278–3295 (2012).
Mockler, T. C., Guo, H., Yang, H., Duong, H. & Lin, C. Antagonistic actions of Arabidopsis cryptochromes and phytochrome B in the regulation of floral induction. Development 126, 2073–2082 (1999).
Harmoko, R. et al. RNA-dependent RNA polymerase 6 is required for efficient hpRNA-induced gene silencing in plants. Mol. Cells 35, 202–209 (2013).
Clough, S. J. Floral dip: agrobacterium-mediated germ line transformation. Methods Mol. Biol. 286, 91–102 (2005).
Wu, F. H. et al. Tape-Arabidopsis Sandwich – a simpler Arabidopsis protoplast isolation method. Plant Methods 5, 16 (2009).
Schindelin, J. et al. Fiji: an open-source platform for biological-image analysis. Nat. Methods 9, 676–682 (2012).
Fang, X. et al. Arabidopsis FLL2 promotes liquid–liquid phase separation of polyadenylation complexes. Nature 569, 265–269 (2019).
Rapsomaniki, M. A. et al. easyFRAP: an interactive, easy-to-use tool for qualitative and quantitative analysis of FRAP data. Bioinformatics 28, 1800–1801 (2012).
Tseng, Q. et al. A new micropatterning method of soft substrates reveals that different tumorigenic signals can promote or reduce cell contraction levels. Lab Chip 11, 2231–2240 (2011).
Lund, F. W. et al. SpatTrack: an imaging toolbox for analysis of vesicle motility and distribution in living cells. Traffic 15, 1406–1429 (2014).
Ries, R. J. et al. m(6)A enhances the phase separation potential of mRNA. Nature 571, 424–428 (2019).
Zielinski, T., Moore, A. M., Troup, E., Halliday, K. J. & Millar, A. J. Strengths and limitations of period estimation methods for circadian data. PLoS ONE 9, e96462 (2014).
Moore, A., Zielinski, T. & Millar, A. J. in Plant Circadian Networks: Methods and Protocols (ed. Staiger, D.) 13–44 (Springer, 2014).
Hutchison, A. L. et al. Improved statistical methods enable greater sensitivity in rhythm detection for genome-wide data. PLoS Comput. Biol. 11, e1004094 (2015).
Chantarachot, T. et al. DHH1/DDX6-like RNA helicases maintain ephemeral half-lives of stress-response mRNAs. Nat. Plants 6, 675–685 (2020).
Sorenson, R. S., Deshotel, M. J., Johnson, K., Adler, F. R. & Sieburth, L. E. Arabidopsis mRNA decay landscape arises from specialized RNA decay substrates, decapping-mediated feedback, and redundancy. Proc. Natl Acad. Sci. USA 115, E1485–E1494 (2018).
Pfaffl, M. W. A new mathematical model for relative quantification in real-time RT–PCR. Nucleic Acids Res. 29, e45 (2001).
Trcek, T., Larson, D. R., Moldon, A., Query, C. C. & Singer, R. H. Single-molecule mRNA decay measurements reveal promoter-regulated mRNA stability in yeast. Cell 147, 1484–1497 (2011).
Kim, D. et al. TopHat2: accurate alignment of transcriptomes in the presence of insertions, deletions and gene fusions. Genome Biol. 14, R36 (2013).
Mortazavi, A., Williams, B. A., McCue, K., Schaeffer, L. & Wold, B. Mapping and quantifying mammalian transcriptomes by RNA-Seq. Nat. Methods 5, 621–628 (2008).
Trapnell, C. et al. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 7, 562–578 (2012).
Luo, G. Z. et al. Unique features of the m6A methylome in Arabidopsis thaliana. Nat. Commun. 5, 5630 (2014).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Yang, Y., Li, Y., Sancar, A. & Oztas, O. The circadian clock shapes the Arabidopsis transcriptome by regulating alternative splicing and alternative polyadenylation. J. Biol. Chem. 295, 7608–7619 (2020).
Romanowski, A., Schlaen, R. G., Perez-Santangelo, S., Mancini, E. & Yanovsky, M. J. Global transcriptome analysis reveals circadian control of splicing events in Arabidopsis thaliana. Plant J. 103, 889–902 (2020).
Li, S. et al. CGDB: a database of circadian genes in eukaryotes. Nucleic Acids Res. 45, D397–D403 (2017).
Covington, M. F., Maloof, J. N., Straume, M., Kay, S. A. & Harmer, S. L. Global transcriptome analysis reveals circadian regulation of key pathways in plant growth and development. Genome Biol. 9, R130 (2008).
Acknowledgements
We thank J. Bailey-Serres for research discussions and manuscript editing, R. G. Fray for providing the ABI3::MTA/mta line, R. McClung and X. Xu for providing pCCA1::LUC and pTOC1::LUC reporter constructs and reporter lines, the UCLA-FAFU Joint Research Center on Plant Proteomics and the UCLA-MCDB/BSCRC Microscopy Core for institutional support. Works in the authors’ laboratories are supported in part by the National Natural Science Foundation of China (31970265 to Q.W.), Natural Science Foundation of Fujian Province (2019J06014 to Q.W.), the National Institutes of Health (R01GM056265 to C.L.) and UCLA Sol Leshin Programme (to C.L.).
Author information
Authors and Affiliations
Contributions
C.L. and X.W. designed the study and wrote the paper. L.G. performed bioinformatics analyses. X.W., B.C., Y.C., M.M., M.Z., E.N. and Q.W. performed the experiments and analysed the data. Q.W. contributed materials/analysis tools.
Corresponding authors
Ethics declarations
Competing interests
The authors declare potential competing interests.
Additional information
Peer review information Nature Plants thanks Peter Quail and the other, anonymous, reviewer(s) for their contribution to the peer review of this work.
Publisher’s note Springer Nature remains neutral with regard to jurisdictional claims in published maps and institutional affiliations.
Extended data
Extended Data Fig. 1 CRY-mediated photoresponsive epitranscriptomic changes in plants.
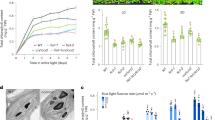
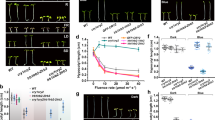
a, The heatmaps showing relative m6A abundance for all genes (epitranscriptome, top) or clock-controlled genes (CCG, bottom) in wild type (WT) and cry1cry2 mutants grown in the dark (D) or blue light conditions (B). b, Genomic visualization of m6A density maps of core clock genes detected in m6A-seq. CRY1 and CRY2 detected only in WT but not cry1cry2 samples are shown as the negative control. c, Hierarchical clustering analysis of different m6A-seq samples indicates no unusual sample variances. d, m6A abundance of individual sites of selected genes was analyzed by m6A-IP-qPCR. The relative m6A level in each gene was calculated by normalizing the m6A-IP to input signals. Data are shown as Mean ± SD from 3 independent experiments. D, dark; B, blue light (30 μmol m-2 s-1).
Extended Data Fig. 2 Epitranscriptomes of selected GO group of genes.
a–j, The scatter plots showing photoresponsive changes of m6A abundance and RNA abundance (left), and metagene profiles showing photoresponsive changes of m6A density (right) of mRNAs categorized as the TAIR10 GO groups.
Extended Data Fig. 3 CRYs interacts with subunits of m6A mRNA methyltransferase in human cells.
a-c, co-IP assays showing interactions between CRY2 and MTA (a), MTB (b) and FIP37 (c) in HEK293T cells which were illuminated with blue light (BL; 100 μmol m-2 s-1) for indicated time (0-60 min). d, co-IP assay showing the interaction of CRY2D387A mutant and MTA in HEK293T cells with or without blue light irradiation. e, co-IP assay showing interactions between MTA and CRY1 in HEK293T cells. BL, blue light (100 μmol m-2 s-1). Three independent experiments are performed for a, two independent experiments for b-e, showing similar results.
Extended Data Fig. 4 Photoresponsive condensation of the CRY2-MTA complex in Arabidopsis.
a, Time-lapse images showing the partial colocalization of MTA and CRY2 in CRY2 photobodies over the time of blue laser illumination in Arabidopsis protoplasts. Scale bar= 2 μm. Two independent experiments are performed showing similar results. b, Quantification of partition coefficient of CRY2-YFP and MTA-DsRed in the assay shown in (a). The data is presented as mean ± SD (n = 15 independent measurements from 5 nuclei). c, Fluorescence profiles of CRY2-YFP and MTA-DsRed over the white line shown in (a). The arrow heads indicate the locations of photobodies. d, Quantification of partition coefficient of CRY2D387A-YFP and MTA-DsRed in the assay shown in (a). The data is presented as mean ± SD (n = 15 independent measurements from 5 nuclei). e, Fluorescence profiles of CRY2D387A-YFP and MTA-DsRed over the white line shown in (a). f, FRAP of CRY2/MTA BiFC photobodies in protoplasts. White circle indicates the region for photobleaching. Quantification of FRAP is shown in Fig. 2m. Scale bar= 2 μm. g, Negative controls of BiFC assays. Plasmids expressing mRFP are used to monitor protoplasts transformation efficiency. The arrow in the cell indicates the location of nucleus. Scale bar = 5μm. Two independent experiments are performed showing similar results. h, m6A-IP-qPCR assay showing the relative abundance of m6A on the 3’UTR of CCA1 mRNA in etiolated wild-type seedlings treated with mock (H2O) or 1,6-hexanediol (10%) in response to blue light. The data is shown as Mean ± SD (n = 6 technical replicates from 2 biological repeats). i, m6A-IP-qPCR assay showing the relative abundance of m6A on the 3’UTR of CCA1 mRNA in wild-type (WT) and indicated genotypes grown in the Dark or Blue light (30 µmol m-2 s-1). The data is shown as Mean ± SD (n = 6 technical replicates from 2 biological repeats).
Extended Data Fig. 5 LLPS of CRY2 in plant cells.
a, FRAP of CRY2 photobodies in CRY2-GFP transgenic plants, the quantification of FRAP is shown in Fig. 3c. Scale bar = 2μm. b, Time-lapse images showing light-elicited formation of CRY2-YFP photobodies in protoplasts. Light-insensitive CRY2D387A-YFP was used as the negative control. Quantification of partition coefficient for both CRY2-YFP and CRY2D387A-YFP fluorescence is shown in Fig. 3e. Scale bar = 2μm. c, FRAP of CRY2-YFP photobodies in Arabidopsis protoplasts. Protoplasts transfected with CRY2-YFP were illuminated with 488 nm laser for 30 s before FRAP. Time-lapse images before and after FRAP are shown on the left, white arrow heads point at the photobodies for photobleaching. Quantification of FRAP assay is shown on the right. Double exponential fit (the dark line) of normalized and averaged recovery curves is shown (n = 5 independent experiments; Mean ± SD). Scale bar = 2μm. d, Representative motion trajectories of CRY2-GFP photobodies in the live or fixed plant nucleus. The time-lapse images were taken every 3 s for 10 min. The dashed lines in the images are the outlines of the nucleus. Scale bar= 5 μm. e, The ensemble-averaged MSD (Mean Square Displacement) curves of tracked CRY2-GFP photobodies (as shown in d) in live or fixed samples. Shaded areas represent standard error (SE) of the mean (n = 4 and 3 independent cells for the live and fixed sample, respectively). Fitting of the MSD for live sample to the anomalous diffusion model yields the Diffusion constant (D), anomalous exponent (α; α < 1 indicates the subdiffusive motion of the particles) and the best fitted curve (black line) which are shown in the graph.
Extended Data Fig. 6 Wavelength specificity of LLPS of CRY2-DsRed in Arabidopsis protoplasts.
a, Time-lapse images showing CRY2-DsRed distribution pattern in the nucleus under different wavelengths in protoplasts. Protoplasts transiently expressing CRY2-DsRed were kept in the dark and illuminated with different lasers during imaging. Scale bar=2 µm. b, Quantification of PC of CRY2-DsRed under different wavelengths (n = 16 measurements for each time point from 3 nuclei; in each box, the center line denotes median value; boxes extend from the 25th to the 75th percentile of each group’s distribution of values; vertical extending lines denote the range from minima to maxima; the open circles denote the values ranging from minima to maxima).
Extended Data Fig. 7 CRY2 photobodies are reversible in response to light-dark cycles in Arabidopsis protoplasts.
Time-lapse images of CRY2-DsRed expressed in protoplasts. Blue triangle indicates that 488 nm laser was on during imaging. Quantification of this experiment is shown in Fig. 3d. Scale bar=2 µm.
Extended Data Fig. 8 PPK1-dependent phosphorylation is essential to maintain CRY2 photobodies in liquid phase.
a, Representative images showing the FRAP of CRY2 photobody in HEK293T cells with or without co-expression of PPK1 or PPK1D267N. The quantification of FRAP assays is shown in Fig. 3j. The dashed circle outlines the nucleus and the small solid circle inside the nucleus indicates the region for photobleaching. Scale bar = 2μm. b, The mobile fraction of CRY2 photobodies at 57 s after photobleaching (mean ± SD; n = 5 independent experiments). p value is calculated in two-tailed Students’ t-test.
Extended Data Fig. 9 C-terminus of CRY2 is necessary to maintain the photobodies in liquid phase in both human cells and Arabidopsis protoplasts.
a, Representative images showing the FRAP of CRY2, CRY2N509 (N509) and CRY2N489 (N489) photobodies in the presence of PPK1 in HEK293T cells. The dashed circle outlines the nucleus and the small solid circle inside the nucleus indicates the region for photobleaching. The quantification of FRAP assays is shown in Fig. 3m. Scale bar = 2μm. b, The mobile fraction of CRY2, N509 and N489 photobodies at 57 s after photobleaching in HEK293T cells (mean ± SD; n = 5 independent experiments). p value is calculated in two-tailed Students’ t-test. c, Images showing the FRAP of CRY2, N509 and N489 photobodies in Arabidopsis protoplasts. The circle inside the nucleus indicates the region for photobleaching. The quantification of FRAP assays is shown in Fig. 3n. Scale bar = 2μm. d, The mobile fraction of CRY2, N509 and N489 photobodies at 57 s after photobleaching in protoplasts (mean ± SD; n = 6 independent experiments). p value is calculated in two-tailed Students’ t-test.
Extended Data Fig. 10 Genotyping of ABI3::MTA/mta plants.
a, The schematic diagram of the T-DNA insertion mta mutant (SALK_074069). The exons (boxes), introns (lines) of the MTA gene and the T-DNA insert (opened triangle) are shown. The positions of primers used for genotyping the mutant locus are marked on MTA gene structure. The positions of RP (right primer) or LP (left primer) of the MTA gene, and primer LBb1.3 in the left border of T-DNA insert are indicated. b, Representative image of WT, mta (short for ABI3::MTA/mta) and cry1cry2 plants grown in long day conditions for 50 days. c, Genomic PCR of the MTA locus in WT and ABI3::MTA/mta plants using genomic DNA as the templates. The sizes of the PCR products were 965 bp (amplified with LP + RP) or 444-744 bp (amplified with LBb1.3+RP). Four plants (#1-#4) were randomly selected from a ABI3::MTA/mta population for genotyping. M, DNA marker.
Supplementary information
Supplementary Video 1
CRY2-YFP photobodies formation under blue light in the nucleus of Arabidopsis protoplast.
Supplementary Video 2
FRAP of CRY2-GFP photobodies in the nucleus of transgenic Arabidopsis plants.
Supplementary Video 3
Dynamics of CRY2-DsRed photobodies in response to light/dark cycle in the nucleus of Arabidopsis protoplast.
Supplementary Video 4
Movement of CRY2-GFP photobodies in the nucleus of transgenic Arabidopsis plants.
Supplementary Table 1
Epitranscriptome and transcriptome data.
Source data
Source Data Fig. 2
Statistical source data.
Source Data Fig. 2
Unprocessed western blots
Source Data Fig. 3
Statistical source data.
Source Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 1
Statistical source data.
Source Data Extended Data Fig. 3
Unprocessed western blots.
Source Data Extended Data Fig. 4
Statistical source data.
Source Data Extended Data Fig. 5
Statistical source data.
Source Data Extended Data Fig. 6
Statistical source data
Source Data Extended Data Fig. 8
Statistical source data
Source Data Extended Data Fig. 9
Statistical source data.
Rights and permissions
About this article
Cite this article
Wang, X., Jiang, B., Gu, L. et al. A photoregulatory mechanism of the circadian clock in Arabidopsis. Nat. Plants 7, 1397–1408 (2021). https://doi.org/10.1038/s41477-021-01002-z
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/s41477-021-01002-z
This article is cited by
-
Genome-wide identification of the N6-methyladenosine regulatory genes reveals NtFIP37B increases drought resistance of tobacco (Nicotiana tabacum L.)
BMC Plant Biology (2024)
-
Liquid–liquid phase separation of TZP promotes PPK-mediated phosphorylation of the phytochrome A photoreceptor
Nature Plants (2024)
-
NBR1-mediated selective autophagy of ARF7 modulates root branching
EMBO Reports (2024)
-
Photobody formation spatially segregates two opposing phytochrome B signaling actions of PIF5 degradation and stabilization
Nature Communications (2024)
-
Recent advances in the plant epitranscriptome
Genome Biology (2023)