Abstract
Ewing sarcoma shows considerable histologic overlap with other round cell tumors. NKX2-2, a homeodomain transcription factor involved in neuroendocrine/glial differentiation and a downstream target of EWSR1-FLI1, has been reported as an immunohistochemical marker for Ewing sarcoma. We assessed the specificity of NKX2-2 for Ewing sarcoma compared with other round cell malignant neoplasms and other soft tissue tumors with EWSR1 translocations. We evaluated whole-tissue sections from 270 cases: 40 Ewing sarcomas (4 with atypical/large cell features), 20 CIC-DUX4 sarcomas, 5 BCOR-CCNB3 sarcomas, 9 unclassified round cell sarcomas, 10 poorly differentiated synovial sarcomas, 10 lymphoblastic lymphomas, 10 alveolar rhabdomyosarcomas, 10 embryonal rhabdomyosarcomas, 10 Merkel cell carcinomas, 10 small cell carcinomas, 20 melanomas, 5 NUT midline carcinomas, 10 Wilms tumors, 10 neuroblastomas, 10 olfactory neuroblastomas, 12 mesenchymal chondrosarcomas, 10 angiomatoid fibrous histiocytomas, 10 clear cell sarcomas, 5 gastrointestinal clear cell sarcoma-like tumors, 5 desmoplastic small round cell tumors, 10 extraskeletal myxoid chondrosarcomas, 10 soft tissue and cutaneous myoepitheliomas, and 19 myoepithelial carcinomas. NKX2-2 positivity was defined as moderate-to-strong nuclear immunoreactivity in at least 5% of cells. NKX2-2 was positive in 37/40 (93%) Ewing sarcomas, including all atypical Ewing sarcomas and cases with known EWSR1-FLI1 or EWSR1-ERG fusion; 85% of Ewing sarcomas showed diffuse (>50%) staining. NKX2-2 was positive in 9 (75%) mesenchymal chondrosarcomas, 8 (80%) olfactory neuroblastomas, 1 CIC-DUX4 sarcoma, 1 poorly differentiated synovial sarcoma, 1 neuroblastoma, 2 unclassified round cell sarcomas, and 3 small cell carcinomas; all other EWSR1-associated tumors were negative for NKX2-2, apart from 1 desmoplastic small round cell tumor, 1 myoepithelioma, and 1 myoepithelial carcinoma. In summary, NKX2-2 is a sensitive but imperfectly specific marker for Ewing sarcoma. Nonetheless, NKX2-2 may be helpful to distinguish Ewing sarcoma from some histologic mimics including CIC-DUX4 and BCOR-CCNB3 sarcomas. Most other EWSR1-associated soft tissue tumors are negative for NKX2-2.
Similar content being viewed by others
Main
Diagnosing Ewing sarcoma can be difficult and often requires immunohistochemical and molecular techniques. Morphologically, Ewing sarcoma belongs to the family of ‘small round blue cell tumors’ with a broad differential diagnosis, including other sarcomas (rhabdomyosarcomas, synovial sarcoma, and desmoplastic small round cell tumor), carcinomas (small cell and Merkel cell), lymphoblastic lymphomas, melanoma, and pediatric tumors (neuroblastoma and Wilms tumor). Unlike other round cell sarcomas, Ewing sarcoma is highly sensitive to specialized chemotherapy regimens;1 accurate diagnosis is therefore critical. In the context of appropriate histomorphology, the gold standard for diagnosing Ewing sarcoma involves demonstrating the presence of a specific fusion gene via reverse-transcription PCR (RT-PCR) or fluorescence in situ hybridization (FISH), using break-apart probes for EWSR1. However, EWSR1 rearrangements with fusion to other partners are common in mesenchymal neoplasms, such as EWSR1-NR4A3 in extraskeletal myxoid chondrosarcoma, EWSR1-DDIT3 in myxoid liposarcoma, EWSR1-WT1 in desmoplastic small round cell tumor, EWSR1-ATF1 or EWSR1-CREB1 in clear cell sarcoma, angiomatoid fibrous histiocytoma, and clear cell sarcoma-like tumor of the gastrointestinal tract (also known as gastrointestinal neuroectodermal tumor), and EWSR1-POU5F1, EWSR1-ZNF444, or EWSR1-PBX1 in a subset of myoepithelial neoplasms.2, 3 A positive FISH result for EWSR1 rearrangement is thus not specific for Ewing sarcoma.
In conjunction with histomorphology and molecular techniques, immunohistochemical markers such as CD99 and FLI-1 have been used for the diagnosis of Ewing sarcoma,4, 5, 6 albeit with suboptimal specificities. CD99 (MIC2 gene product, recognized by the monoclonal antibody O13) is a surface glycoprotein in normal thymic T lymphocytes and an extremely sensitive marker for Ewing sarcoma; however, its expression is also observed in neuroendocrine carcinoma, lymphoma, synovial sarcoma, rhabdomyosarcoma, and mesenchymal chondrosarcoma.4, 5, 6 A member of the ETS family and a marker of endothelial differentiation, FLI-1 appears less sensitive for Ewing sarcoma than CD99 and is also expressed in various other round cell tumors including neuroblastoma, lymphoma, synovial sarcoma, rhabdomyosarcoma, desmoplastic small round cell tumor, Merkel cell carcinoma, and small cell carcinoma.7 NKX2-2, a homeodomain-containing transcription factor involved in neuronal development and glial/neuroendocrine differentiation,8, 9 is a downstream target of EWSR1-FLI1 oncogenic signaling in Ewing sarcoma.10 In recent times, NKX2-2 has been reported as a diagnostically useful immunohistochemical marker for Ewing sarcoma.11
In published studies on NKX2-2 expression in Ewing sarcoma,11, 12 only classic Ewing sarcomas with EWSR1-FLI1 rearrangement have been evaluated. It thus remains unclear whether NKX2-2 can be diagnostically useful in atypical Ewing sarcomas, Ewing sarcomas with other gene rearrangements such as EWSR1-ERG, and other Ewing sarcoma-like round cell sarcomas including those with CIC-DUX4 or BCOR-CCNB3 fusion. Although gene expression studies suggest that CIC-DUX4 and BCOR-CCNB3 round cell sarcomas do not express typical EWSR1-ETS target genes such as NKX2-2, data on protein expression of NKX2-2 in these tumors are lacking. Furthermore, it is unknown whether NKX2-2 is expressed in soft tissue tumors with EWSR1 fusion to other partners. This study aimed to evaluate the specificity of NKX2-2 as an immunohistochemical marker for Ewing sarcoma versus other round cell sarcomas, including those with CIC-DUX4 or BCOR-CCNB3 fusion, and other soft tissue tumors with EWSR1 rearrangement.
Materials and methods
Cases were retrieved from the surgical pathology and consultation files of Brigham and Women’s Hospital, Boston, MA, and from the consultation files of two of the authors (CDMF and JLH). Representative hematoxylin-and-eosin-stained slides were reviewed, with whole-tissue sections evaluated for NKX2-2 expression in 270 cases: 40 Ewing sarcomas, 20 CIC-DUX4 round cell sarcomas, 5 BCOR-CCNB3 round cell sarcomas, 10 poorly differentiated synovial sarcomas, 10 lymphoblastic lymphomas, 10 alveolar rhabdomyosarcomas, 10 embryonal rhabdomyosarcomas, 10 Merkel cell carcinomas, 10 small cell carcinomas, 20 melanomas, 5 NUT midline carcinomas, 10 Wilms tumors, 10 neuroblastomas, 10 olfactory neuroblastomas, 12 mesenchymal chondrosarcomas, 10 angiomatoid fibrous histiocytomas, 10 clear cell sarcomas, 5 gastrointestinal clear cell sarcoma-like tumors, 5 desmoplastic small round cell tumors, 10 extraskeletal myxoid chondrosarcomas, 10 myoepitheliomas (6 soft tissue and 4 cutaneous syncytial), 19 soft tissue myoepithelial carcinomas, and 9 unclassified round cell sarcomas. Unclassified round cell sarcomas were defined as those with no identifiable line of differentiation by immunohistochemistry and no EWSR1 or CIC gene rearrangements (see Results). Immunohistochemistry for CD99 was previously performed on 39 Ewing sarcomas, 18 CIC-DUX4 sarcomas, 4 BCOR-CCNB3 sarcomas, and 8 unclassified round cell sarcomas. Immunohistochemistry for WT-1 was previously performed in 18 CIC-DUX4 sarcomas.
Genetic confirmation by FISH and/or RT-PCR was previously obtained in all Ewing sarcomas, CIC-DUX4 sarcomas, poorly differentiated synovial sarcomas, and four of five BCOR-CCNB3 sarcomas (see below and Acknowledgments). Of the 40 Ewing sarcoma cases, 35 were confirmed by FISH for EWSR1 rearrangement and 5 were confirmed by RT-PCR. EWSR1-FLI1 fusion was confirmed in seven cases: four by cytogenetics with t(11;22)(q24;q12), two by RT-PCR, and one by OncoPanel (a massively parallel sequencing platform using Agilent SureSelect hybrid capture kit and Illumina HiSeq 2500 sequencer)13 and was previously published.14 EWSR1-ERG fusion was confirmed in three cases: one by cytogenetics with t(21;22)(q22;q12), one by RT-PCR, and one by OncoPanel. All 20 CIC-DUX4 sarcomas were confirmed by FISH and/or RT-PCR. Of the five BCOR-CCNB3 sarcomas, four were previously confirmed by RT-PCR, with the remaining case confirmed by immunohistochemistry for CCNB3.15 All 10 poorly differentiated synovial sarcomas were confirmed by FISH for SS18 rearrangement. Genetic confirmation by FISH for EWSR1 gene rearrangement and/or RT-PCR was previously obtained in all angiomatoid fibrous histiocytomas, gastrointestinal clear cell sarcoma-like tumors, and extraskeletal myxoid chondrosarcomas, as well as five clear cell sarcomas and four desmoplastic small round cell tumors. Of the 19 myoepithelial carcinomas, 7 were previously evaluated by FISH for EWSR1 rearrangement, which was present in 2 cases and absent in 5 cases.
Immunohistochemistry for NKX2-2 was performed on 4-μm-thick formalin-fixed paraffin-embedded whole-tissue sections following pressure cooker antigen retrieval (0.01 M citrate buffer pH 6.0) using a mouse anti-NKX2-2 monoclonal antibody (1:25 dilution, 40 min incubation, clone 74.5A5; Developmental Studies Hybridoma Bank, Iowa City, IA). Immunohistochemistry for CD99 was performed using a mouse anti-CD99 antibody (1:150 dilution, 40 min incubation, clone O13; BioLegend, Dedham, MA). Immunohistochemistry for WT-1 was performed using a mouse anti-WT1 monoclonal antibody (1:50 dilution, 40 min incubation, clone 6F-H2; Dako, Carpinteria, CA). Appropriate positive and negative controls were used throughout the study. The extent of immunoreactivity was graded according to the percentage of cells with nuclear staining (0, <5%; 1+, 5–25%; 2+, 25–50%; 3+, 50–75%; or 4+, 75–100%) and the intensity of the staining (weak, moderate, or strong). NKX2-2 positivity was defined as moderate or strong nuclear immunoreactivity in at least 5% of cells.
Results
Clinicopathologic Characteristics of the Ewing Sarcoma and Other Round Cell Sarcoma Study Groups
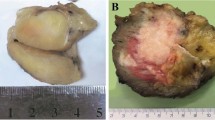
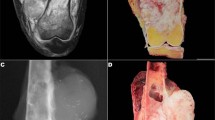
The Ewing sarcoma cases affected 21 males and 19 females, with a median age of 30 years (age range, 10–76 years). The anatomic distribution of primary Ewing sarcomas was wide and included lower limb (n=11), head and neck (n=7), upper limb (n=6), thoracic cavity (n=6), abdominopelvic cavity (n=6), retroperitoneum (n=2), trunk (n=1), and paravertebral region (n=1). The Ewing sarcoma study group typically involved soft tissue (n=18) or bone (n=10), rarely the lung (n=4), skin (n=2), brain (n=2), submandibular gland (n=2), oral cavity (n=1), or the uterus (n=1). Thirty-five primary Ewing sarcomas were included, along with two locally recurrent Ewing sarcomas and three metastatic tumors (in the lung or liver). Of the 40 Ewing sarcoma cases, 36 demonstrated classic histomorphology characterized by uniform, small, round-to-ovoid, primitive-appearing cells in a solid or vaguely lobular architecture (Figures 1a and 2a). The remaining four Ewing sarcoma cases were classified as ‘atypical,’ characterized by large-cell cytomorphology with vesicular chromatin and prominent nucleoli (Figure 2c). CD99 expression was observed in all 39 cases tested: 36 cases (92%) diffuse or multifocal and 3 cases (8%) focal and weak. As described above, all Ewing sarcomas had confirmed EWSR1 rearrangements.
The CIC-DUX4 sarcomas affected 12 male and 8 female patients, with a median age of 34 years (age range: 6 months to 81 years). The anatomic distribution of the primary tumors included abdominopelvic cavity (n=6), head and neck (n=4), lower limb (n=4), thoracic cavity (n=3), paravertebral region (n=2), and upper limb (n=1). Primary CIC-DUX4 sarcomas were typically extraosseous (n=18) rather than osseous (n=2); 18 primary and 2 metastatic cases were included. Histologic features of CIC-DUX4 sarcomas included small and round-to-ovoid or focally spindled cells, with vesicular chromatin, conspicuous nucleoli, and palely amphophilic cytoplasm (Figure 3a and c). As compared with Ewing sarcoma, CIC-DUX4 sarcomas displayed more variable cytomorphology and intratumoral heterogeneity. CD99 expression was present in 15 of 18 cases (83%) and ranged from patchy, mixed cytoplasmic and membranous (n=12) to a diffuse, predominantly membranous (n=3) staining pattern; nuclear WT-1 expression was present in 16 of 18 cases (89%).
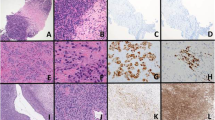
CIC-DUX4 sarcoma of the perineum composed of small ovoid cells with vesicular chromatin, small nucleoli, and amphophilic cytoplasm (a); NKX2-2 is negative (b). CIC-DUX4 sarcoma in the posterior mediastinum composed of ovoid-to-short spindle cells (c) with focal nuclear immunoreactivity for NKX2-2 (d). BCOR-CCNB3 sarcoma consisting of round and spindle cells with scant myxoid stroma (e) and showing an absence of NKX2-2 immunoreactivity (f).
The BCOR-CCNB3 sarcomas affected five male patients, with a median age of 12 years (age range: 5–44 years). Three tumors arose in the bone (sacrum, femur, and skull base) and two in soft tissue (pelvis and thigh). Histologic features of BCOR-CCNB3 sarcomas included small and round or ovoid to focally spindled cells (Figure 3e), reminiscent of Ewing sarcoma or CIC-DUX4 sarcoma. CD99 and WT-1 expression was absent in all cases tested (five and four cases, respectively).
The nine unclassified round cell sarcomas affected six males and three females, with a median age of 25 years (age range: 3–54 years), all extraosseous with primary sites including lower limb (n=4), chest wall (n=2), head and neck (n=1), abdominal cavity (n=1), and paravertebral region (n=1). CD99 expression was observed in all eight cases tested (diffuse, membranous in one, multifocal in five, and weak, multifocal in two) and nuclear WT-1 staining was present in five of eight cases. All nine cases were genetically confirmed to lack EWSR1 and CIC gene rearrangement; BCOR and FUS gene rearrangements were confirmed to be absent in eight and seven cases, respectively.
NKX2-2 Expression in Ewing Sarcomas, Non-Ewing Sarcoma Round Cell Malignant Neoplasms, and EWSR1-Associated Soft Tissue Tumors
The results of immunohistochemistry for NKX2-2 are summarized in Table 1. As we have noted weak nuclear staining with a wide range of antibodies directed against transcription factors in clinical practice (which we interpret as a nonspecific finding in most instances), we scored weak nuclear staining as negative for this study. NKX2-2 was positive in 37 of 40 Ewing sarcoma cases (93%), including all 4 cases of ‘atypical’ Ewing sarcoma with large-cell cytomorphology (Figure 2d), all 7 cases with known EWSR1-FLI1 fusion (Figure 1d), and all 3 cases with known EWSR1-ERG fusion (Figure 2b). Among the 40 Ewing sarcoma cases, the extent of NKX2-2 immunoreactivity was as follows: 4+ in 31, 3+ in 4, 2+ in 2, 1+ in 2, and 0 in 1 (2 of these cases showed weak staining and were considered negative). The intensity of staining was as follows: strong in 23, moderate in 14, and weak in 3. Thirty-four (85%) cases showed diffuse (>50%) and moderate-to-strong staining.
NKX2-2 was positive in 28 of 230 (12%) non-Ewing sarcoma cases, including 9 of 12 (75%) mesenchymal chondrosarcomas (4+ in 7, 3+ in 1, and 2+ in 1, strong in 2 and moderate in 7), 8 of 10 (80%) olfactory neuroblastomas (4+ in 5 and 3+ in 3, strong in 4 and moderate in 4), 1 of 20 (5%) CIC-DUX4 sarcoma (4+ moderate), 1 of 10 (10%) poorly differentiated synovial sarcoma (2+ moderate), 2 of 9 (22%) unclassified round cell sarcomas (4+ moderate and 3+ strong), 1 of 10 (10%) neuroblastoma (4+ strong), 3 of 10 (30%) small cell carcinomas (2+ strong, 1+ moderate, and 1+ moderate), 1 of 5 (20%) desmoplastic small round cell tumors (2+ moderate), 1 of 10 (10%) myoepitheliomas (2+ moderate), and 1 of 19 (5%) myoepithelial carcinomas (4+ moderate). All lymphoblastic lymphomas, alveolar rhabdomyosarcomas, embryonal rhabdomyosarcomas, NUT midline carcinomas, Wilms tumors, Merkel cell carcinomas, and melanomas were negative for NKX2-2. Of the 28 NKX2-2-positive non-Ewing sarcoma cases, 21 showed diffuse, moderate-to-strong staining, including 8 mesenchymal chondrosarcomas, 8 olfactory neuroblastomas, 1 CIC-DUX4 sarcoma, 2 unclassified round cell sarcomas, 1 neuroblastoma, and 1 myoepithelial carcinoma (Figures 4,5,6).
Poorly differentiated synovial sarcoma consisting of sheets of round cells with vesicular chromatin and small nucleoli (a), and showing focal weak-to-moderate immunoreactivity for NKX2-2 (b). Small cell carcinoma (c) with focal moderate-to-strong immunoreactivity for NKX2-2 (d). Thymic neuroblastoma consisting of small round cells embedded within a fibrillary matrix (e); immunohistochemistry for NKX2-2 shows diffuse, moderate-to-strong nuclear staining (f).
Among EWSR1-associated soft tissue tumors, all angiomatoid fibrous histiocytomas, clear cell sarcomas, gastrointestinal clear cell sarcoma-like tumors, and extraskeletal myxoid chondrosarcomas were negative for NKX2-2. Focal-to-diffuse moderate staining for NKX2-2 was present only in 1 desmoplastic small round cell tumor (2+ moderate), 1 myoepithelioma (2+ moderate), and 1 myoepithelial carcinoma (4+ moderate) (Figure 6). Thus, aside from Ewing sarcoma, most EWSR1-associated soft tissue tumors were negative for NKX2-2.
Discussion
Ewing sarcoma is the second most common sarcoma in children and young adults; older adults may be less often affected. Ewing sarcoma most often presents in the bone, although a large subset of cases arises primarily at extraskeletal sites. Rare histologic variants of Ewing sarcoma include atypical ‘large-cell,’ ‘vascular-like,’ ‘adamantinoma-like,’ spindle cell and sclerosing.4, 5, 16 Ewing sarcoma is characterized by reciprocal chromosomal translocation and gene rearrangement involving the Ewing sarcoma breakpoint region 1 (EWSR1) at 22q12 and FLI1 at 11q24 in 85–90% of cases.17 This rearrangement creates the oncogenic transcription factor EWSR1-FLI1, whereby the transactivation domain of EWSR1, a ubiquitously expressed member of unknown function in the TET family, is fused to the DNA-binding domain of FLI1, a member in the ETS family.18 The remainder of Ewing sarcoma cases are characterized by alternate fusion partners, with FLI1 substituted by ERG in 5–10% of cases, rarely by other ETS members (FEV, E1AF, ETV1, and ETV4) or non-ETS genes (POU5F1, PATZ1, and NFATc2), or with EWSR1 substituted by FUS.2, 3 Ewing sarcoma and its histologic variants (including primitive neuroectodermal tumor) are currently classified by the World Health Organization as a single histopathologic entity.19
The chimeric transcription factor EWSR1-FLI1 in Ewing sarcoma drives oncogenic signaling of downstream targets including NKX2-2. Since its identification in gene expression studies,10 NKX2-2 has been described as a diagnostic immunohistochemical marker for Ewing sarcoma,11 in a similar manner as DOG1 (ANO1) for gastrointestinal stromal tumor,20 TLE1 for synovial sarcoma,21 and MUC4 for low-grade fibromyxoid sarcoma.22 Two prior studies demonstrated NKX2-2 positivity in 80–93% of Ewing sarcomas;11, 12 however, only cases with classic morphology and EWSR1-FLI1 rearrangements were evaluated. In this study, we examined 40 genetically confirmed Ewing sarcomas, including 4 with atypical large-cell cytomorphology, 4 from uncommon locations (cutaneous and uterine), and 3 with confirmed EWSR1-ERG rearrangements. NKX2-2 was positive in 37 (93%) Ewing sarcoma cases, including all atypical Ewing sarcomas and tumors with known EWSR1-FLI1 or EWSR1-ERG fusion. Moreover, diffuse (>50%) and moderate-to-strong staining was present in 34 (85%) cases. The sensitivity of NKX2-2 for Ewing sarcoma in this study was thus comparable to prior studies. Although the three Ewing sarcomas negative for NKX2-2 were confirmed to harbor EWSR1 rearrangements by FISH, they were not tested by other molecular methods; the fusion partners are therefore unknown.
Distinct subsets of undifferentiated round cell sarcomas with morphologic resemblance to Ewing sarcoma have recently been shown to harbor gene rearrangements leading to CIC-DUX4 or BCOR-CCNB3 fusion. Gene fusion between CIC (encoding a high-mobility group box transcription factor) on 19q13 and one of the two DUX4 retrogenes (encoding a double-homeobox transcription factor) on 4q35 or 10q26 was reported initially in a Ewing sarcoma-like round cell sarcoma23 and subsequently in 19–68% of EWSR1-negative round cell sarcomas.24, 25 Highly aggressive in children and adults, CIC-DUX4-associated round cell sarcomas display small to intermediate-sized, round-to-spindled cells with pale cytoplasm, vesicular chromatin, prominent nucleoli, and more cytologic variability as compared with Ewing sarcoma.24, 25, 26 Despite morphologic overlap with Ewing sarcoma, CIC-DUX4 sarcomas show a distinct immunoprofile, with variable CD99 staining25 but consistent nuclear WT1 expression,27 and a distinct gene expression signature, with upregulation of WT1 and ETS transcription factors including ETV4, ETV5, and ETV1.23, 27 An additional 4–12% of EWSR1-negative round cell sarcomas harbor a recurrent paracentric inversion of chromosome X, leading to gene fusion between BCOR (encoding the BCL6 co-repressor) and CCNB3 (encoding the testis-specific cyclin B3).15, 28 Predominantly arising in the bone in young children, BCOR-CCNB3-associated round cell sarcomas show spindled-to-round cell morphology, with positive immunohistochemistry for CCNB3 in most cases.15, 28, 29 Despite histologic overlap with Ewing sarcoma, BCOR-CCNB3 sarcomas do not show an EWSR1-ETS expression signature and thus appear biologically distinct from Ewing sarcoma.15 This is the first study, to our knowledge, that compared NKX2-2 expression in Ewing sarcomas to Ewing sarcoma-like round cell sarcomas with CIC-DUX4 and BCOR-CCNB3 fusions. We observed NKX2-2 expression in only 1 of 20 CIC-DUX4 sarcomas and none of 5 BCOR-CCNB3 sarcomas evaluated. NKX2-2 may be a useful adjunct to separate Ewing sarcoma from CIC-DUX4 and BCOR-CCNB3 sarcomas.
NKX2-2 is sensitive but imperfectly specific for Ewing sarcoma among most histologic mimics, and diagnostic pitfalls remain. In this study, NKX2-2 expression was absent in all lymphoblastic lymphomas, alveolar and embryonal rhabdomyosarcomas, Merkel cell carcinomas, NUT midline carcinomas, melanomas, and Wilms tumors. Focal NKX2-2 staining was noted in one poorly differentiated synovial sarcoma and three small cell carcinomas. Diffuse NKX2-2 staining was noted in 1 neuroblastoma, 8 of 10 (80%) olfactory neuroblastomas, and 9 of 12 (75%) mesenchymal chondrosarcomas. Given that EWSR1-FLI1 promotes NKX2-2 expression by upregulating GLI1,30 NKX2-2 positivity in olfactory neuroblastomas may be explained by tumoral activation of sonic hedgehog signaling.31 Mesenchymal chondrosarcomas, characterized by HEY1-NCOA2 gene fusion,32 demonstrated immunoreactivity for NKX2-2 in 33% of cases in a prior study11 and, in this study, 75% of cases, showing diffuse moderate staining in most cases. The mechanism underlying NKX2-2 expression in mesenchymal chondrosarcomas is unknown. Although a combination of CD99 and NKX2-2 has been suggested to be highly specific for the diagnosis of Ewing sarcoma,12 this would not exclude mesenchymal chondrosarcomas, which are often positive for both markers.
Despite the known connection between EWSR1-FLI1 and NKX2-2 in Ewing sarcoma, this is the first study, to our knowledge, that evaluated NKX2-2 expression in other neoplasms driven by EWSR1 rearrangements, including Ewing sarcomas with EWSR1-ERG and other soft tissue tumors with EWSR1 fusions. The presence of NKX2-2 expression in all three Ewing sarcomas with genetically confirmed EWSR1-ERG rearrangements suggests that similar to EWSR1-FLI1, rearrangements involving other ETS members such as ERG may drive oncogenic signaling with downstream targets including NKX2-2. Notably, most other EWSR1 fusion-driven soft tissue tumors were negative for NKX2-2, except one desmoplastic small round cell tumor and two soft tissue myoepithelial tumors. Apart from EWSR1, whether other TET members such as FUS may drive NKX2-2 expression is currently unknown. Interestingly, NKX2-2 positivity was noted in two unclassified round cell sarcomas genetically confirmed to lack EWSR1, CIC, and BCOR gene rearrangements (one of which also lacks FUS gene rearrangement), suggesting that NKX2-2 expression may be promoted by other as yet unidentified gene rearrangements.
In summary, NKX2-2 is a sensitive but imperfectly specific immunohistochemical marker for Ewing sarcoma, as several other round cell malignant neoplasms commonly express this transcription factor, most notably mesenchymal chondrosarcoma and olfactory neuroblastoma. Nonetheless, NKX2-2 may be helpful in distinguishing Ewing sarcoma from some histologic mimics including CIC-DUX4 and BCOR-CCNB3 sarcomas. NKX2-2 is also positive in atypical Ewing sarcoma and Ewing sarcoma with EWSR1-ERG gene rearrangements. Most other EWSR1-associated soft tissue tumors are negative for NKX2-2.
References
Grier HE, Krailo MD, Tarbell NJ et al. Addition of ifosfamide and etoposide to standard chemotherapy for Ewing's sarcoma and primitive neuroectodermal tumor of bone. N Engl J Med 2003;348:694–701.
Lessnick SL, Ladanyi M . Molecular pathogenesis of Ewing sarcoma: new therapeutic and transcriptional targets. Annu Rev Pathol 2012;7:145–159.
Romeo S, Dei Tos AP . Soft tissue tumors associated with EWSR1 translocation. Virchows Arch 2010;456:219–234.
Folpe AL, Goldblum JR, Rubin BP et al. Morphologic and immunophenotypic diversity in Ewing family tumors: a study of 66 genetically confirmed cases. Am J Surg Pathol 2005;29:1025–1033.
Llombart-Bosch A, Machado I, Navarro S et al. Histological heterogeneity of Ewing's sarcoma/PNET: an immunohistochemical analysis of 415 genetically confirmed cases with clinical support. Virchows Arch 2009;455:397–411.
Weidner N, Tjoe J . Immunohistochemical profile of monoclonal antibody O13: antibody that recognizes glycoprotein p30/32MIC2 and is useful in diagnosing Ewing’s sarcoma and peripheral neuroepithelioma. Am J Surg Pathol 1994;18:486–494.
Mhawech-Fauceglia P, Herrmann FR, Bshara W et al. Friend leukaemia integration-1 expression in malignant and benign tumours: a multiple tumour tissue microarray analysis using polyclonal antibody. J Clin Pathol 2007;60:694–700.
Briscoe J, Sussel L, Serup P et al. Homeobox gene Nkx2.2 and specification of neuronal identity by graded Sonic hedgehog signalling. Nature 1999;398:622–627.
Wang YC, Gallego-Arteche E, Iezza G et al. Homeodomain transcription factor NKX2.2 functions in immature cells to control enteroendocrine differentiation and is expressed in gastrointestinal neuroendocrine tumors. Endocr Relat Cancer 2009;16:267–279.
Smith R, Owen LA, Trem DJ et al. Expression profiling of EWS/FLI identifies NKX2.2 as a critical target gene in Ewing's sarcoma. Cancer Cell 2006;9:405–416.
Yoshida A, Sekine S, Tsuta K et al. NKX2.2 is a useful immunohistochemical marker for Ewing sarcoma. Am J Surg Pathol 2012;36:993–999.
Shibuya R, Matsuyama A, Nakamoto M et al. The combination of CD99 and NKX2.2, a transcriptional target of EWSR1-FLI1, is highly specific for the diagnosis of Ewing sarcoma. Virchows Arch 2014;465:599–605.
Wagle N, Berger MF, Davis MJ et al. High-throughput detection of actionable genomic alterations in clinical tumor samples by targeted, massively parallel sequencing. Cancer Discov 2012;2:82–93.
Doyle LA, Wong KK, Bueno R et al. Ewing sarcoma mimicking atypical carcinoid tumor: detection of unexpected genomic alterations demonstrates the use of next generation sequencing as a diagnostic tool. Cancer Genet 2014;207:335–339.
Pierron G, Tirode F, Lucchesi C et al. A new subtype of bone sarcoma defined by BCOR-CCNB3 gene fusion. Nat Genet 2012;44:461–466.
Bridge JA, Fidler ME, Neff JR et al. Adamantinoma-like Ewing’s sarcoma: genomic confirmation, phenotypic drift. Am J Surg Pathol 1999;23:159–165.
Delattre O, Zucman J, Plougastel B et al. Gene fusion with an ETS DNA-binding domain caused by chromosome translocation in human tumours. Nature 1992;359:162–165.
May WA, Gishizky ML, Lessnick SL et al. Ewing sarcoma 11;22 translocation produces a chimeric transcription factor that requires the DNA-binding domain encoded by FLI1 for transformation. Proc Natl Acad Sci USA 1993;90:5752–5756.
Fletcher CDM, Bridge JA, Hogendoorn PCW et al. WHO Classification of Tumours of Soft Tissue and Bone 4th (edn). IARC Press: Lyon, 2013.
West RB, Corless CL, Chen X et al. The novel marker, DOG1, is expressed ubiquitously in gastrointestinal stromal tumors irrespective of KIT or PDGFRA mutation status. Am J Pathol 2004;165:107–113.
Terry J, Saito T, Subramanian S et al. TLE1 as a diagnostic immunohistochemical marker for synovial sarcoma emerging from gene expression profiling studies. Am J Surg Pathol 2007;31:240–246.
Doyle LA, Moller E, Dal Cin P et al. MUC4 is a highly sensitive and specific marker for low-grade fibromyxoid sarcoma. Am J Surg Pathol 2011;35:733–741.
Kawamura-Saito M, Yamazaki Y, Kaneko K et al. Fusion between CIC and DUX4 up-regulates PEA3 family genes in Ewing-like sarcomas with t(4;19)(q35;q13) translocation. Hum Mol Genet 2006;15:2125–2137.
Graham C, Chilton-MacNeill S, Zielenska M et al. The CIC-DUX4 fusion transcript is present in a subgroup of pediatric primitive round cell sarcomas. Hum Pathol 2012;43:180–189.
Italiano A, Sung YS, Zhang L et al. High prevalence of CIC fusion with double-homeobox (DUX4) transcription factors in EWSR1-negative undifferentiated small blue round cell sarcomas. Genes Chromosomes Cancer 2012;51:207–218.
Choi EY, Thomas DG, McHugh JB et al. Undifferentiated small round cell sarcoma with t(4;19)(q35;q13.1) CIC-DUX4 fusion: a novel highly aggressive soft tissue tumor with distinctive histopathology. Am J Surg Pathol 2013;37:1379–1386.
Specht K, Sung YS, Zhang L et al. Distinct transcriptional signature and immunoprofile of CIC-DUX4 fusion-positive round cell tumors compared to EWSR1-rearranged Ewing sarcomas: further evidence toward distinct pathologic entities. Genes Chromosomes Cancer 2014;53:622–633.
Peters TL, Kumar V, Polikepahad S et al. BCOR-CCNB3 fusions are frequent in undifferentiated sarcomas of male children. Mod Pathol 2014;28:575–586.
Puls F, Niblett A, Marland G et al. BCOR-CCNB3 (Ewing-like) sarcoma: a clinicopathologic analysis of 10 cases, in comparison with conventional Ewing sarcoma. Am J Surg Pathol 2014;38:1307–1318.
Zwerner JP, Joo J, Warner KL et al. The EWS/FLI1 oncogenic transcription factor deregulates GLI1. Oncogene 2008;27:3282–3291.
Mao L, Xia YP, Zhou YN et al. Activation of sonic hedgehog signaling pathway in olfactory neuroblastoma. Oncology 2009;77:231–243.
Wang L, Motoi T, Khanin R et al. Identification of a novel, recurrent HEY1-NCOA2 fusion in mesenchymal chondrosarcoma based on a genome-wide screen of exon-level expression data. Genes Chromosomes Cancer 2012;51:127–139.
Acknowledgements
We thank Cristina Antonescu (Memorial Sloan-Kettering Cancer Center, New York, NY) and Paola Dal Cin (Brigham and Women’s Hospital, Boston, MA) for performing FISH and RT-PCR studies during the work-up of the cases included in this study.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Hung, Y., Fletcher, C. & Hornick, J. Evaluation of NKX2-2 expression in round cell sarcomas and other tumors with EWSR1 rearrangement: imperfect specificity for Ewing sarcoma. Mod Pathol 29, 370–380 (2016). https://doi.org/10.1038/modpathol.2016.31
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.31
This article is cited by
-
Adamantinoma-like ewing sarcoma arising in the pancreatic tail: a case report of a rare entity and review of the literature
Diagnostic Pathology (2023)
-
The value of GLI1 and p16 immunohistochemistry in the premolecular screening for GLI1-altered mesenchymal neoplasms
Virchows Archiv (2023)
-
TrkC, a novel prognostic marker, induces and maintains cell survival and metastatic dissemination of Ewing sarcoma by inhibiting EWSR1-FLI1 degradation
Cell Death & Disease (2022)
-
A unique case of adamantinoma-like Ewing sarcoma in the calcaneus, exhibiting prominent squamous differentiation and displaying EWSR1 gene rearrangement
Skeletal Radiology (2022)
-
Does PAX7 and NKX2.2 immunoreactivity in Ewing sarcoma have prognostic significance?
Virchows Archiv (2022)