Abstract
A distinct subset of round cell sarcomas harbors capicua transcriptional repressor (CIC) rearrangement. Diagnosing these sarcomas can be difficult owing to their resemblance to Ewing sarcoma and other 'small round blue cell tumors'; molecular techniques are generally required. Recent gene expression studies of CIC-rearranged sarcomas identified the upregulation of ETV4. We assessed the sensitivity and specificity of ETV4 and WT1 immunohistochemistry for CIC-rearranged sarcoma. We evaluated whole-tissue sections from 40 CIC-rearranged sarcomas, 40 Ewing sarcomas, 4 BCOR-CCNB3 sarcomas, 6 unclassified round cell sarcomas, and 150 histologic mimics. Moderate-to-strong nuclear immunoreactivity for ETV4 in at least 50% of cells was observed in 36 (90%) CIC-rearranged sarcomas and 10 (5%) other tumors, including 5 unclassified round cell sarcomas, 2 Wilms tumors, and 1 each desmoplastic small round cell tumor, melanoma, and small cell carcinoma. Thirty-eight (95%) CIC-rearranged sarcomas showed nuclear staining for WT1, and 34 (85%) were positive for both ETV4 and WT1. Of 182 other tumors evaluated, 34 (19%) showed nuclear WT1 positivity, including all Wilms tumors and desmoplastic small round cell tumors, 5 unclassified round cell sarcomas, and a subset of lymphoblastic lymphomas, rhabdomyosarcomas, mesenchymal chondrosarcomas, carcinomas, and melanomas. In summary, diffuse moderate-to-strong ETV4 expression is present in most CIC-rearranged sarcomas and unclassified round cell sarcomas. More limited expression is seen in small subsets of various other round cell neoplasms. Nuclear WT1 expression is also present in most CIC-rearranged sarcomas and unclassified round cell sarcomas, along with Wilms tumors and desmoplastic small round cell tumors, and subsets of various histologic mimics. The sensitivity and specificity of diffuse ETV4 expression for CIC-rearranged sarcomas are 90% and 95%, respectively, whereas the sensitivity and specificity of WT1 are 95% and 81%, respectively. Diffuse ETV4 along with at least focal WT1 expression is helpful to distinguish CIC-rearranged sarcoma from Ewing sarcoma and other histologic mimics.
Similar content being viewed by others
Main
A distinct subset of round cell sarcomas that resemble Ewing sarcoma is negative for Ewing sarcoma breakpoint region 1 (EWSR1) rearrangement and instead harbors recurrent capicua transcriptional repressor (CIC) rearrangement,1 predominantly characterized by gene fusion between CIC on 19q13 and one of the two DUX4 retrogenes on 4q35 or 10q26.2, 3 CIC-rearranged sarcomas often present at extraskeletal sites and are highly aggressive with poor prognosis.4, 5 Diagnosing CIC-rearranged sarcoma can be difficult because of its histologic overlap with other 'small round blue cell tumors,' including various sarcomas (Ewing sarcoma, BCOR-CCNB3 round cell sarcoma, synovial sarcoma, myxoid liposarcoma, rhabdomyosarcomas, and desmoplastic small round cell tumor), carcinomas (small cell, NUT midline, and Merkel cell), lymphoblastic lymphomas, melanomas, and pediatric tumors (neuroblastomas and Wilms tumor). The diagnosis of CIC-rearranged sarcomas relies on demonstrating the presence of characteristic karyotype such as t(4;19)(q35;q13.1) or t(10;19)(q26;q13.1), reverse-transcription PCR (RT-PCR), or fluorescence in situ hybridization (FISH). By immunohistochemistry, CIC-rearranged sarcomas show nuclear WT1, ERG, and FLI1 expression, along with variable CD99 immunoreactivity.4, 6, 7 These markers are not specific, however, and can be seen in various other round cell neoplasms.8, 9, 10, 11, 12, 13
ETV4 (ETS variant 4; also known as PEA3 or E1AF), a member of the PEA3 subgroup in the ETS transcription factor family, is involved in regulating early embryogenesis14 and branching morphogenesis of epithelial structures including ureteric buds and limb buds.15, 16 Somatic mutations in ETV4 have been implicated in the progression of breast, colonic, gastric, endometrial, and squamous cell carcinomas.15 Translocations involving ETV4 are present in <1% of Ewing sarcomas (EWSR1-ETV4) and 1–2% of prostatic adenocarcinomas (TMPRSS2-ETV4).15, 17, 18 Furthermore, ETV4 expression is upregulated in CIC-mutant 1p/19q-codeleted oligodendrogliomas.19 Recent gene expression studies in CIC-rearranged sarcomas identified upregulation of WT1 and ETV4,3, 7 along with ETV5 (ERM) and ETV1,1, 3, 7 which appear to distinguish CIC-rearranged sarcomas from Ewing sarcomas. A recent study suggested that immunohistochemistry for ETV4 is sensitive and specific for CIC-DUX4 sarcomas.20 This study aimed to validate the diagnostic utility of ETV4 as an immunohistochemical marker for CIC-rearranged sarcomas versus other round cell histologic mimics, in comparison with WT1.
Materials and methods
Cases were retrieved after approval of the Institution Review Board from the surgical pathology and consultation files of Brigham and Women's Hospital, Boston, MA, USA, and from the consultation files of two of the authors (CDMF and JLH). Representative hematoxylin and eosin-stained slides were reviewed. Whole-tissue sections were evaluated for ETV4 expression in 240 cases: 40 CIC-rearranged sarcomas, 40 Ewing sarcomas, 4 BCOR-CCNB3 round cell sarcomas, 6 unclassified round cell sarcomas, 10 poorly differentiated synovial sarcomas, 10 high-grade (round cell) myxoid liposarcomas, 10 lymphoblastic lymphomas, 10 alveolar rhabdomyosarcomas, 10 embryonal rhabdomyosarcomas, 10 mesenchymal chondrosarcomas, 10 Merkel cell carcinomas, 10 Wilms tumors, 10 neuroblastomas, 10 olfactory neuroblastomas, 5 NUT midline carcinomas, 5 desmoplastic small round cell tumors, 20 small cell carcinomas, and 20 melanomas. Immunohistochemistry for WT1 was performed on all 40 CIC-rearranged sarcomas and 182 other tumors (additional materials from 18 of the cases listed above were not available). Immunohistochemistry for CD99 was previously performed on 37 CIC-rearranged sarcomas, 39 Ewing sarcomas, 4 BCOR-CCNB3 sarcomas, and 6 unclassified round cell sarcomas. Immunohistochemistry for NKX2-2 was previously performed on 17 CIC-rearranged sarcomas, 38 Ewing sarcomas, 4 BCOR-CCNB3 sarcomas, and 5 unclassified round cell sarcomas.21
Genetic confirmation by FISH and/or RT-PCR was previously obtained in all CIC-rearranged sarcomas, Ewing sarcomas, poorly differentiated synovial sarcomas, high-grade myxoid liposarcomas, as well as 4 of 5 desmoplastic small round cell tumors and 2 of 4 BCOR-CCNB3 sarcomas (see below). Of the 40 CIC-rearranged sarcomas, 38 were confirmed by break-apart FISH for CIC rearrangement, with the remaining two cases confirmed by RT-PCR for CIC-DUX4 fusion. Of the 40 Ewing sarcomas, 36 were confirmed by FISH for EWSR1 rearrangement, with the remaining 4 cases confirmed by RT-PCR; in addition, 4 cases were confirmed to be negative for CIC rearrangement by FISH, and 4 cases underwent targeted next generation sequencing by OncoPanel (a massively parallel sequencing platform using Agilent SureSelect hybrid capture kit and Illumina HiSeq 2500 sequencer).22 EWSR1-FLI1 fusion was confirmed in seven cases: three by cytogenetics with t(11;22)(q24;q12), two by both RT-PCR and OncoPanel, one by RT-PCR alone, and one by OncoPanel alone. EWSR1-ERG fusion was confirmed in three cases: one by cytogenetics with t(21;22)(q22;q12), one by RT-PCR, and one by OncoPanel. Of the four BCOR-CCNB3 sarcomas, two were confirmed by RT-PCR (one additionally confirmed by FISH), and two were confirmed by immunohistochemistry for CCNB3.23 All 10 poorly differentiated synovial sarcomas were confirmed by FISH for SS18 rearrangement. All 10 high-grade myxoid liposarcomas were confirmed by FISH for DDIT3 (CHOP) rearrangement, with one case showing concurrent EWSR1 rearrangement and no FUS rearrangement consistent with EWSR1-DDIT3 fusion. Of five desmoplastic small round cell tumors, four were confirmed by FISH for EWSR1 gene rearrangement.
Immunohistochemistry for ETV4 was performed on 4 μm-thick formalin-fixed paraffin-embedded whole-tissue sections following pressure cooker antigen retrieval (Target Retrieval Solution, pH 6.1; Dako, Carpinteria, CA, USA) using a mouse anti-PEA3 (ETV4) monoclonal antibody (1:50 dilution; 40 min incubation; clone 16/sc-113; Santa Cruz Biotechnology, Santa Cruz, CA, USA) and the Novolink Polymer Detection System (Leica, Buffalo Grove, IL, USA). The latter detection system was used after preliminary testing with the EnVision+ kit (Dako) showed at most weak staining, even at low primary antibody dilution. Immunohistochemistry for WT1 was performed using a mouse anti-WT1 monoclonal antibody (1:50 dilution; 40 min incubation; clone 6F-H2; Dako), CD99 using a mouse anti-CD99 antibody (1:150 dilution; 40 min incubation; clone O13; BioLegend, Dedham, MA, USA), and NKX2-2 using a mouse anti-NKX2-2 monoclonal antibody (1:25 dilution; 40 min incubation; clone 74.5A5; Developmental Studies Hybridoma Bank, Iowa City, IA, USA), as previously described.21 Appropriate positive and negative controls were used throughout the study. The extent of immunoreactivity was graded according to the percentage of cells with nuclear staining (0, <5%; 1+, 5–25%; 2+, 25–50%; 3+, 50–75%; or 4+, 75–100%) and intensity (weak, moderate, or strong). ETV4 positivity was defined as moderate-to-strong nuclear immunoreactivity in at least 50% of cells. WT1 positivity was defined as moderate-to-strong nuclear immunoreactivity in at least 5% of cells.
Results
Clinicopathologic Characteristics of the CIC-Rearranged Sarcoma and Other Round Cell Sarcoma Study Groups
The CIC-rearranged sarcomas affected 21 males and 19 females, with a median age of 32 years (age range, 0.5–81 years). The anatomic distribution of the primary CIC-rearranged sarcomas included lower limb (n=13), abdominopelvic cavity (n=12), thoracic cavity (n=5), upper limb (n=3), retroperitoneum (n=3), paravertebral region (n=2), and head and neck (n=2). Thirty-eight primary tumors were included, along with one locally recurrent and one metastatic case (in the lung); the study group typically involved soft tissue (n=37), rarely bone (n=1), lung (n=1), or colon (n=1). CIC-rearranged sarcomas displayed a wide histologic spectrum. The tumors predominantly showed a diffuse or vaguely lobular architecture, with small to intermediate-sized, round-to ovoid and occasionally spindled cells with vesicular chromatin and prominent nucleoli; the cytoplasm could be moderate or scant, eosinophilic and occasionally clear or amphophilic (Figure 1a and c). Geographic necrosis, hemorrhage, and focal sclerotic stroma were commonly seen, along with frequent mitoses and single-cell apoptosis. Two CIC-rearranged sarcomas showed a reticular or pseudoalveolar architecture within a myxoid stroma (Figure 1e), mimicking extraskeletal myxoid chondrosarcoma or myoepithelial carcinoma.4, 6 Immunohistochemical expression of CD99 was present in 32 of 37 cases (87%), with staining pattern ranging from focal (n=5) or multifocal (n=24), mixed membranous and cytoplasmic, to diffuse and predominantly membranous (n=3). All 17 cases tested were negative for NKX2-2 expression.
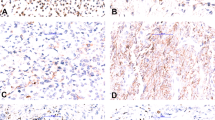
CIC-rearranged sarcoma in the gluteal region, composed of ovoid cells with amphophilic cytoplasm and prominent nucleoli (a), showing diffuse ETV4 staining (b). CIC-rearranged sarcoma in the soft tissue of the posterior neck, composed of sheets and vague nests of epithelioid cells with clear cytoplasm and small nucleoli (c), showing diffuse nuclear immunoreactivity for ETV4 (d). CIC-rearranged sarcoma in the buttock, characterized by round-to epithelioid cells with hyperchromatic nuclei in a reticular architecture within a myxoid stroma (e), mimicking extraskeletal myxoid chondrosarcoma or a myoepithelial tumor; diffuse strong nuclear ETV4 expression is present (f).
The Ewing sarcomas affected 22 males and 18 females, with a median age of 29 years (age range, 10–76 years). The anatomic distribution of primary Ewing sarcomas included lower limb (n=9), abdominopelvic cavity (n=9), head and neck (n=7), upper limb (n=6), thoracic cavity (n=6), retroperitoneum (n=2), and paravertebral region (n=1); this study group typically involved soft tissue (n=19) or bone (n=10), rarely lung (n=4), brain (n=2), submandibular gland (n=2), skin (n=1), oral cavity (n=1), or uterus (n=1), with 35 primary, 2 locally recurrent, and 3 metastatic cases (in the lung or liver). Ewing sarcoma showed classic histomorphology with uniform round cells in a solid or lobular arrangement (Figure 2a) in 36 cases, with the remainder 4 characterized by atypical cytomorphology with small nucleoli and small amounts of cytoplasm. CD99 expression was observed in all 39 cases tested, predominantly with diffuse (n=36) and rarely focal or weak (n=3) membranous staining patterns. NKX2-2 expression was present in 32 of 38 cases (84%).
The BCOR-CCNB3 sarcomas affected four males, with a median age of 12 years (age range, 2–44 years) and included two osseous (femur and sacrum) and two extraosseous primary sites (neck and thigh). BCOR-CCNB3 sarcomas showed ovoid to focally spindled tumor cells in a myxoid or collagenous stroma (Figure 2c). CD99 and NKX2-2 expression were absent in all four cases tested.
The six unclassified round cell sarcomas affected three males and three females, with a median age of 20 years (age range, 13–34 years), all extraosseous and involving lower limb in two cases, chest wall, abdominopelvic cavity, tonsil, and the paravertebral region in one case each. All six cases showed diffuse sheets or vague lobules of round-to-ovoid and focally spindled tumor cells with eosinophilic or focally clear cytoplasm (Figure 3a and c). Of the six cases, all showed diffuse to multifocal membranous CD99 expression; none of the five tested cases were positive for NKX2-2. All six unclassified round cell sarcomas were genetically confirmed by FISH to lack EWSR1 and CIC gene rearrangement; BCOR, FUS, and ERG gene rearrangements were confirmed by FISH to be absent in five, four, and three cases, respectively.
An unclassified round cell sarcoma in the abdominal cavity of a 17-year-old woman (a), without CIC, EWSR1, or BCOR rearrangements, showing diffuse strong ETV4 expression (b). An unclassified round cell sarcoma in the supraclavicular soft tissue of a 34-year-old man (c), showing diffuse strong nuclear ETV4 expression (d).
ETV4 Expression in CIC-Rearranged Sarcomas and Other Round Cell Neoplasms
The results of ETV4 immunohistochemistry in CIC-rearranged sarcomas and other round cell tumors are summarized in Table 1. Diffuse moderate-to-strong nuclear ETV4 staining was present in 36 of 40 (90%) CIC-rearranged sarcomas (Figure 1b), with at least 1+ staining of any intensity in 38 of 40 (95%) cases. The intensity of ETV4 staining in the 40 CIC-rearranged sarcomas was as follows: strong in 29, moderate in 10, and weak in 1. In contrast, Ewing sarcomas showed at most focal weak-to-moderate ETV4 expression (1+ in 12, including moderate in 6 and weak in 6; 0 in 28). There was no ETV4 immunoreactivity in BCOR-CCNB3 sarcomas (Figure 2d), olfactory neuroblastomas, Merkel cell carcinomas, and NUT midline carcinomas (all 0). Of the 200 non-CIC-rearranged tumors, ETV4 staining was present in 30 (15%). Focal (5–50%) moderate nuclear ETV4 expression was present in 20 (10%) cases, including 6 (15%) Ewing sarcomas (Figure 2b), 6 melanomas, 3 Wilms tumors, and 1 each poorly differentiated synovial sarcoma, lymphoblastic lymphoma, alveolar rhabdomyosarcoma, embryonal rhabdomyosarcoma, and neuroblastoma. Diffuse moderate-to-strong nuclear ETV4 expression was present in 10 (5%) cases overall, including 5 unclassified round cell sarcomas (strong in three and moderate in two) (Figure 3), 2 Wilms tumors (1 moderate and 1 strong) (Figure 4f), 1 desmoplastic small round cell tumor (moderate) (Figure 5f), 1 melanoma (strong) (Figure 4b), and 1 small cell carcinoma (strong) (Figure 4d).
High-grade myxoid (round cell) liposarcoma (a) showing an absence of ETV4 staining (b). Poorly differentiated synovial sarcoma (c) showing focal weak ETV4 staining (d). Desmoplastic small round cell tumor (e) showing nests of round-to ovoid cells and multifocal weak-to-moderate ETV4 immunoreactivity (f).
Although most CIC-rearranged sarcomas showed diffuse strong nuclear ETV4 expression, focal weak-to-moderate ETV4 staining was present in a minor subset of various other round cell tumors. As antibodies directed against transcription factors occasionally show weak nuclear staining in clinical practice, which in most instances we interpret as nonspecific, ETV4 positivity was defined as moderate-to-strong nuclear staining in at least 50% of cells. With this stringent threshold, ETV4 immunohistochemistry was positive in 90% of CIC-rearranged sarcomas and negative in all Ewing sarcomas, BCOR-CCNB3 sarcomas, poorly differentiated synovial sarcomas, high-grade myxoid liposarcomas, lymphoblastic lymphomas, alveolar rhabdomyosarcomas, embryonal rhabdomyosarcomas, mesenchymal chondrosarcomas, Merkel cell carcinomas, neuroblastomas, olfactory neuroblastomas, and NUT midline carcinomas. Overall, ETV4 positivity was present in 5% of non-CIC-rearranged tumors, including 5 of 6 unclassified round cell sarcomas, 2 of 10 Wilms tumors, 1 of 5 desmoplastic small round cell tumors, and 1 of 20 small cell carcinomas and melanomas.
WT1 Expression in CIC-Rearranged Sarcomas and Other Round Cell Neoplasms
The results of WT1 immunohistochemistry in CIC-rearranged sarcomas and other round cell tumors are summarized in Table 2. Of 40 CIC-rearranged sarcomas, 38 (95%) showed nuclear staining for WT1, including 31 (78%) with diffuse (>50%) moderate-to-strong reactivity; 34 tumors (85%) were positive for both ETV4 and WT1. Nuclear WT1 expression was absent in Ewing sarcomas, BCOR-CCNB3 sarcomas, poorly differentiated synovial sarcomas, high-grade myxoid liposarcomas, neuroblastomas, and olfactory neuroblastomas (Table 2). Of 182 non-CIC-rearranged tumors tested, 34 (19%) showed nuclear WT1 positivity, including all Wilms tumors and desmoplastic small round cell tumors, 5 of 6 unclassified round cell sarcomas, and a subset of lymphoblastic lymphomas, rhabdomyosarcomas, mesenchymal chondrosarcomas, carcinomas, and melanomas.
The sensitivity and specificity of diffuse ETV4 expression for CIC-rearranged sarcomas were 90% and 95%, respectively, whereas the sensitivity and specificity of nuclear WT1 were 95% and 81%, respectively. The specificity of expression of both markers was 96%.
Discussion
A primitive round cell sarcoma characterized by a recurrent chromosomal translocation t(4;19)(q35;q13.1) was first described in 1996 in the ankle of a 12-year-old boy, who developed bilateral pulmonary metastasis and died within 10 months.24 In 2006, two adult extraosseous round cell sarcomas with Ewing sarcoma-like morphology and t(4;19)(q35;q13.1) translocation were shown to harbor a fusion between CIC on 19q13 and DUX4 on 4q35, creating an oncogenic transcription factor CIC-DUX4.1 CIC, a human homolog of Drosophila capicua, encodes a high mobility group box transcription repressor in a receptor tyrosine kinase pathway.25 CIC is normally expressed in cerebellar granular neurons; its dysregulation is implicated in spinocerebellar ataxia type 1 neurodegeneration, and inactivating CIC mutations have been identified in oligodendrogliomas19 and breast cancer.26 DUX4 and its paralog, located in the D4Z4 macrosatellite repeat at 4q35 and 10q26, respectively,27 encode a double homeodomain transcription factor that is normally expressed in testis and is epigenetically suppressed in differentiated cells.28 Aberrant DUX4 expression due to D4Z4 contraction at 4q35 underlies facioscapulohumeral muscular dystrophy.29 As compared with wild-type CIC, the chimeric CIC-DUX4 enhances transcriptional activities of selected genes by >100 fold1 with activation of WT1 along with PEA3 subgroup of ETS transcription factors including ETV4, ETV1, and ETV5.1, 7 Since its initial description, CIC rearrangement has been documented in over 50 cases of pediatric and adult Ewing sarcoma-like round cell sarcomas1, 2, 3, 4, 5, 6, 7, 30, 31, 32, 33, 34 and accounts for up to 68% of EWSR1-negative round cell sarcomas.3
Although referred to as ‘Ewing-like sarcoma’1 and morphologically overlapping with Ewing sarcoma,35, 36 CIC-rearranged sarcoma shows distinct clinical, histologic, immunophenotypic, and molecular features: clinically, whereas CIC-rearranged sarcoma and Ewing sarcoma present with a wide age distribution in both sexes,4 osseous presentation is rare in CIC-rearranged sarcomas but common in Ewing sarcoma;4, 21 CIC-rearranged sarcoma is associated with chemoresistance and worse prognosis as compared to Ewing sarcoma.4, 5, 6 Histologically, unlike typical Ewing sarcoma with small uniform round cells, CIC-rearranged sarcoma displays variable cytomorphology and intratumoral heterogeneity, with ovoid to spindled and occasionally clear cells, vesicular chromatin, prominent nucleoli, and amphophilic cytoplasm. The architecture is usually diffuse or lobular, reminiscent of Ewing sarcomas; however, rare cases can show a reticular or pseudoalveolar patterns (Figure 1e), reminiscent of extraskeletal myxoid chondrosarcoma or myoepithelial carcinoma.4, 6 Immunophenotypically, CIC-rearranged sarcoma shows variable CD99 but strong nuclear WT1 expression and is negative for NKX2-2,4, 6, 7, 21 whereas Ewing sarcoma is diffusely and strongly positive for CD99 and NKX2-2 but negative for nuclear WT1 in most cases.4, 11, 13, 21, 37 FLI1 and ERG are positive in both CIC-rearranged and Ewing sarcomas and thus are not discriminatory.6, 7, 9, 10, 12 CIC-rearranged sarcoma rarely expresses desmin, keratin,3, 4, 5, 33 or S-100 protein,2, 3, 34 but often shows calretinin4 and diffuse MYC expression,6 which may be secondary to trisomy 8 with MYC amplification in subsets of cases.5 Genetically, Ewing sarcoma is characterized by fusion of EWSR1 on 22q12 (or rarely FUS) to FLI1 on 11q24 (or less frequently ERG or other ETS family members including FEV, ETV1, or E1AF/ETV4),38 whereas 70–80% of CIC-rearranged sarcomas published to date involves fusion between CIC and one of the two DUX4 retrogenes on 4q35 or 10q26.3, 7
This study describes ETV4 as a diagnostic immunohistochemical marker for CIC-rearranged sarcoma and compares its diagnostic performance with WT1. ETV4 immunohistochemistry is sensitive for CIC-rearranged sarcomas, with at least 1+ staining in 95% and diffuse moderate-to-strong expression in 90% of cases. As most tumors not associated with CIC rearrangement showed no to only focal immunoreactivity, diffuse ETV4 expression appears relatively specific for CIC-rearranged sarcomas among most histologic mimics including Ewing and BCOR-CCNB3 sarcomas. Nevertheless, diffuse moderate-to-strong ETV4 staining was noted in 5% of small cell carcinomas and melanomas, 2 of 10 Wilms tumors, 1 of 5 desmoplastic small round cell tumors, and 5 of 6 unclassified round cell sarcomas. Despite similar sensitivities of ETV4 and WT1 for CIC-rearranged sarcomas (90% and 95%, respectively), diffuse ETV4 expression is more specific than WT1 (95% versus 81%). We observed nuclear WT1 positivity in 19% of round cell malignant neoplasms other than CIC-rearranged sarcomas, including 100% of Wilms tumors and desmoplastic small round cell tumors, five of six unclassified round cell sarcomas, and a subset of lymphoblastic lymphomas, rhabdomyosarcomas, mesenchymal chondrosarcomas, carcinomas, and melanomas. The specificity of expression of both markers was 96%. Of note, as the five unclassified round cell sarcomas were histologically and immunophenotypically similar to CIC-rearranged sarcomas (that is, with expression of both ETV4 and WT1), we speculate that these cases may harbor cryptic CIC rearrangements that were not detected by conventional break-apart FISH.
In summary, ETV4 is a sensitive but not entirely specific immunohistochemical marker for CIC-rearranged sarcomas. Although weak ETV4 expression is present focally in small subsets of various other round cell tumors, diffuse moderate-to-strong ETV4 expression is 90% sensitive and 95% specific for CIC-rearranged sarcomas, with exceptional positive cases including occasional Wilms tumors and desmoplastic small round cell tumors, and rare small cell carcinomas and melanomas. ETV4 may thus be helpful to distinguish CIC-rearranged sarcomas from histologic mimics, such as Ewing sarcomas and those with BCOR-CCNB3 fusion, although we would expect the rare Ewing sarcomas with EWSR1-ETV4 fusion (<1% of all Ewing sarcomas) to express ETV4 as well. Most tumors not associated with CIC rearrangement are negative for ETV4. Although nuclear WT1 positivity is 95% sensitive for CIC-rearranged sarcoma, its specificity is only 81%. The combination of diffuse ETV4 and at least focal nuclear WT1 expression is helpful to distinguish CIC-rearranged sarcoma from histologic mimics.
References
Kawamura-Saito M, Yamazaki Y, Kaneko K et al. Fusion between CIC and DUX4 up-regulates PEA3 family genes in Ewing-like sarcomas with t(4;19)(q35;q13) translocation. Hum Mol Genet 2006;15:2125–2137.
Graham C, Chilton-MacNeill S, Zielenska M et al. The CIC-DUX4 fusion transcript is present in a subgroup of pediatric primitive round cell sarcomas. Hum Pathol 2012;43:180–189.
Italiano A, Sung YS, Zhang L et al. High prevalence of CIC fusion with double-homeobox (DUX4) transcription factors in EWSR1-negative undifferentiated small blue round cell sarcomas. Genes Chromosomes Cancer 2012;51:207–218.
Yoshida A, Goto K, Kodaira M et al. CIC-rearranged sarcomas: a study of 20 cases and comparisons with Ewing sarcomas. Am J Surg Pathol 2016;40:313–323.
Choi EY, Thomas DG, McHugh JB et al. Undifferentiated small round cell sarcoma with t(4;19)(q35;q13.1) CIC-DUX4 fusion: a novel highly aggressive soft tissue tumor with distinctive histopathology. Am J Surg Pathol 2013;37:1379–1386.
Smith SC, Buehler D, Choi EY et al. CIC-DUX sarcomas demonstrate frequent MYC amplification and ETS-family transcription factor expression. Mod Pathol 2015;28:57–68.
Specht K, Sung YS, Zhang L et al. Distinct transcriptional signature and immunoprofile of CIC-DUX4 fusion-positive round cell tumors compared to EWSR1-rearranged Ewing sarcomas: further evidence toward distinct pathologic entities. Genes Chromosomes Cancer 2014;53:622–633.
Barnoud R, Sabourin JC, Pasquier D et al. Immunohistochemical expression of WT1 by desmoplastic small round cell tumor: a comparative study with other small round cell tumors. Am J Surg Pathol 2000;24:830–836.
Mhawech-Fauceglia P, Herrmann FR, Bshara W et al. Friend leukaemia integration-1 expression in malignant and benign tumours: a multiple tumour tissue microarray analysis using polyclonal antibody. J Clin Pathol 2007;60:694–700.
Miettinen M, Wang ZF, Paetau A et al. ERG transcription factor as an immunohistochemical marker for vascular endothelial tumors and prostatic carcinoma. Am J Surg Pathol 2011;35:432–441.
Nakatsuka S, Oji Y, Horiuchi T et al. Immunohistochemical detection of WT1 protein in a variety of cancer cells. Mod Pathol 2006;19:804–814.
Wang WL, Patel NR, Caragea M et al. Expression of ERG, an Ets family transcription factor, identifies ERG-rearranged Ewing sarcoma. Mod Pathol 2012;25:1378–1383.
Weidner N, Tjoe J . Immunohistochemical profile of monoclonal antibody O13: antibody that recognizes glycoprotein p30/32MIC2 and is useful in diagnosing Ewing's sarcoma and peripheral neuroepithelioma. Am J Surg Pathol 1994;18:486–494.
Xin JH, Cowie A, Lachance P et al. Molecular cloning and characterization of PEA3, a new member of the Ets oncogene family that is differentially expressed in mouse embryonic cells. Genes Dev 1992;6:481–496.
Oh S, Shin S, Janknecht R . ETV1, 4 and 5: an oncogenic subfamily of ETS transcription factors. Biochim Biophys Acta 2012;1826:1–12.
Lu BC, Cebrian C, Chi X et al. Etv4 and Etv5 are required downstream of GDNF and Ret for kidney branching morphogenesis. Nat Genet 2009;41:1295–1302.
Sankar S, Lessnick SL . Promiscuous partnerships in Ewing's sarcoma. Cancer Genet 2011;204:351–365.
Urano F, Umezawa A, Hong W et al. A novel chimera gene between EWS and E1A-F, encoding the adenovirus E1A enhancer-binding protein, in extraosseous Ewing's sarcoma. Biochem Biophys Res Commun 1996;219:608–612.
Padul V, Epari S, Moiyadi A et al. ETV/Pea3 family transcription factor-encoding genes are overexpressed in CIC-mutant oligodendrogliomas. Genes Chromosomes Cancer 2015;54:725–733.
Le Guellec S, Velasco V, Perot G et al. ETV4 is a useful marker for the diagnosis of CIC-DUX4 round cell sarcomas: a study of 110 cases including mimicking lesions. Mod Pathol 2016;29:21A.
Hung YP, Fletcher CD, Hornick JL . Evaluation of NKX2-2 expression in round cell sarcomas and other tumors with EWSR1 rearrangement: imperfect specificity for Ewing sarcoma. Mod Pathol 2016;29:370–380.
Wagle N, Berger MF, Davis MJ et al. High-throughput detection of actionable genomic alterations in clinical tumor samples by targeted, massively parallel sequencing. Cancer Discov 2012;2:82–93.
Pierron G, Tirode F, Lucchesi C et al. A new subtype of bone sarcoma defined by BCOR-CCNB3 gene fusion. Nat Genet 2012;44:461–466.
Richkind KE, Romansky SG, Finklestein JZ . t(4;19)(q35;q13.1): a recurrent change in primitive mesenchymal tumors? Cancer Genet Cytogenet 1996;87:71–74.
Jimenez G, Guichet A, Ephrussi A et al. Relief of gene repression by torso RTK signaling: role of capicua in Drosophila terminal and dorsoventral patterning. Genes Dev 2000;14:224–231.
Jimenez G, Shvartsman SY, Paroush Z . The Capicua repressor—a general sensor of RTK signaling in development and disease. J Cell Sci 2012;125:1383–1391.
Lemmers RJ, van der Vliet PJ, van der Gaag KJ et al. Worldwide population analysis of the 4q and 10q subtelomeres identifies only four discrete interchromosomal sequence transfers in human evolution. Am J Hum Genet 2010;86:364–377.
Snider L, Geng LN, Lemmers RJ et al. Facioscapulohumeral dystrophy: incomplete suppression of a retrotransposed gene. PLoS Genet 2010;6:e1001181.
Lemmers RJ, van der Vliet PJ, Klooster R et al. A unifying genetic model for facioscapulohumeral muscular dystrophy. Science 2010;329:1650–1653.
Somers GR, Gupta AA, Doria AS et al. Pediatric undifferentiated sarcoma of the soft tissues: a clinicopathologic study. Pediatr Dev Pathol 2006;9:132–142.
Sugita S, Arai Y, Tonooka A et al. A novel CIC-FOXO4 gene fusion in undifferentiated small round cell sarcoma: a genetically distinct variant of Ewing-like sarcoma. Am J Surg Pathol 2014;38:1571–1576.
Gambarotti M, Benini S, Gamberi G et al. CIC-DUX4 Fusion-Positive Round Cell Sarcomas of soft tissue and bone: a single institution morphologic and molecular analysis of 7 cases. Histopathology; advance online publication, 15 April 2016; doi:10.1111/his.12985 [e-pub ahead of print].
Machado I, Cruz J, Lavernia J et al. Superficial EWSR1-negative undifferentiated small round cell sarcoma with CIC/DUX4 gene fusion: a new variant of Ewing-like tumors with locoregional lymph node metastasis. Virchows Arch 2013;463:837–842.
Somers GR, Shago M, Zielenska M et al. Primary subcutaneous primitive neuroectodermal tumor with aggressive behavior and an unusual karyotype: case report. Pediatr Dev Pathol 2004;7:538–545.
Antonescu C . Round cell sarcomas beyond Ewing: emerging entities. Histopathology 2014;64:26–37.
Marino-Enriquez A, Fletcher CD . Round cell sarcomas - biologically important refinements in subclassification. Int J Biochem Cell Biol 2014;53:493–504.
Yoshida A, Sekine S, Tsuta K et al. NKX2.2 is a useful immunohistochemical marker for Ewing sarcoma. Am J Surg Pathol 2012;36:993–999.
Romeo S, Dei Tos AP . Soft tissue tumors associated with EWSR1 translocation. Virchows Arch 2010;456:219–234.
Acknowledgements
We thank Cristina Antonescu (Memorial Sloan-Kettering Cancer Center, New York, NY, USA) and Paola Dal Cin (Brigham and Women's Hospital, Boston, MA, USA) for performing FISH and RT-PCR studies during the work-up of the cases included in this study. We thank Mei Zheng and the immunohistochemistry laboratory at the Brigham and Women's Hospital, Boston, MA, USA for technical support.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Hung, Y., Fletcher, C. & Hornick, J. Evaluation of ETV4 and WT1 expression in CIC-rearranged sarcomas and histologic mimics. Mod Pathol 29, 1324–1334 (2016). https://doi.org/10.1038/modpathol.2016.140
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2016.140
This article is cited by
-
Co-expression of ERG and CD31 in a subset of CIC-rearranged sarcoma: a potential diagnostic pitfall
Modern Pathology (2022)
-
Inactivation of the CIC-DUX4 oncogene through P300/CBP inhibition, a therapeutic approach for CIC-DUX4 sarcoma
Oncogenesis (2021)
-
Neues in der aktuellen WHO-Klassifikation (2020) für Weichgewebssarkome
Der Pathologe (2021)
-
Pancreaticoduodenectomy for a primary duodenal capicua transcriptional repressor (CIC) -rearranged sarcoma with severe bleeding: a case report
BMC Gastroenterology (2020)
-
NFATc2-rearranged sarcomas: clinicopathologic, molecular, and cytogenetic study of 7 cases with evidence of AGGRECAN as a novel diagnostic marker
Modern Pathology (2020)