Abstract
Solid-pseudopapillary neoplasms are rare, but are distinctive pancreatic tumors of low-malignant potential. While the histogenesis of these tumors is unclear, they are often associated with gain-of-function mutations in the catenin (cadherin-associated protein), beta 1 (88 kDa), or CTNNB1 gene, resulting in nuclear accumulation of CTNNB1. CTNNB1 is a central component of the Wnt signaling pathway and mediates gene expression through the lymphoid enhancer-binding factor 1 (LEF1) /T-cell factor transcription complex. Although LEF1 has a pivotal role in the transactivation of Wnt/CTNNB1 responsive genes, the status of LEF1 in solid-pseudopapillary neoplasms and other pancreatic tumors has not been examined. We analyzed both LEF1 and CTNNB1 in a large cohort of pancreatic tumors (n=155). In all cases of solid-pseudopapillary neoplasms including surgical resections (n=27) and cytologic samples (n=8) had strong and diffuse nuclear labeling for both LEF1 and CTNNB1. The surrounding uninvolved pancreatic parenchyma was devoid of any LEF1 staining. All resection and cytologic specimens from well-differentiated pancreatic neuroendocrine tumors (n=44; n=29, respectively), high-grade pancreatic neuroendocrine carcinomas (n=2; n=1), pancreatic ductal adenocarcinomas (n=25; n=12), and acinar cell carcinomas (n=9; n=2) studied were negative for both nuclear LEF1 and CTNNB1. However, nuclear LEF1 and CTNNB1 were detected in all four resected pancreatoblastomas (no cytologic specimens were available for immunolabeling), but primarily centered around and within squamoid corpuscles. In summary, abnormal CTNNB1 accumulation was accompanied by nuclear LEF1 overexpression in both solid-pseudopapillary neoplasms and pancreatoblastomas. But, in contrast to pancreatoblastomas, a diffuse, nuclear labeling was observed in solid-pseudopapillary neoplasms and further implicates the CTNNB1/LEF1 transcriptional complex in the development of solid-pseudopapillary neoplasms. In addition, as part of an immunohistochemical panel, LEF1 can be a useful ancillary stain in the diagnosis of solid-pseudopapillary neoplasms.
Similar content being viewed by others
Introduction
First described by Franz in 1959, solid-pseudopapillary neoplasms are distinctive, but are rare pancreatic tumors.1 Patients are typically young females within their 20s and 30s and often present with nonspecific abdominal symptoms.2, 3, 4 Solid-pseudopapillary neoplasms can occur throughout the pancreas and generally form a solitary, well-demarcated mass with an average diameter between 5 and 10 cm.5 Despite their large size, the majority of these tumors behave indolently and, upon resection, are associated with a favorable prognosis.6, 7 As their name would imply, solid-pseudopapillary neoplasms are histologically composed of poorly cohesive, monomorphic cells forming a heterogenous growth pattern of solid, pseudopapillary and hemorrhagic, pseudocystic structures. While over 700 cases have been described, their cell of origin, direction of differentiation, and pathogenesis remain elusive.8
Recent whole-exomic sequencing of neoplastic pancreatic cysts has uncovered a relative paucity of genetic alterations in solid-pseudopapillary neoplasms.9 In fact, the only recurrent mutations identified were those located in CTNNB1, and thus, implicating CTNNB1 in the pathogenesis of these tumors. CTNNB1 is a key mediator of the Wnt signal transduction pathway. Under normal conditions, CTNNB1 is primarily located at the cytoplasmic plasma cell membrane and constantly targeted for degradation by phosphorylation. The continual elimination of CTNNB1 prevents it from reaching the nucleus. Missense mutations within exon 3 of CTNNB1, as those found in solid-pseudopapillary neoplasms, inhibit phosphorylation, resulting in cytoplasmic and eventual nuclear accumulation of CTNNB1.10, 11, 12 Upon nuclear translocation, mutant CTNNB1 interacts with the DNA-bound lymphoid enhancer-binding factor 1/T-cell factor (LEF1/TCF) transcriptional complex.13, 14 By itself, LEF1 has no transcriptional activation potential, but in association with CTNNB1, LEF1 transactivates a host of Wnt-responsive genes including those that regulate cell cycle and survival.15 Consistent with these observations, the presence of nuclear CTNNB1 and overexpression of its transcriptional targets (eg cyclin D1) are characteristic findings and useful markers in the diagnosis of solid-pseudopapillary neoplasms.10, 16
Although LEF1 has a crucial role in Wnt/CTNNB1 signaling, the status of LEF1 in solid-pseudopapillary neoplasms has not been examined. We, therefore, analyzed both LEF1 and CTNNB1 protein expression by immunohistochemistry in a large cohort of solid-pseudopapillary neoplasms that included surgical resections and cytology specimens. Furthermore, we compared these findings to other pancreatic tumors, which may histologically and cytologically mimic solid-pseudopapillary neoplasms.
Materials and methods
Study Cohort
Study approval was obtained from the University of Pittsburgh and Johns Hopkins Hospital Institutional Review Boards. Resection specimens to include hematoxylin and eosin slides, and formalin-fixed and paraffin-embedded tissues blocks from 27 solid-pseudopapillary neoplasms, 44 well-differentiated pancreatic neuroendocrine tumors, 2 high-grade pancreatic neuroendocrine carcinomas, 25 pancreatic ductal adenocarcinomas, and 4 acinar cell carcinomas were identified and retrieved from the surgical pathology archives at the University of Pittsburgh Medical Center, Department of Pathology. In addition, 5 acinar cell carcinomas and 4 pancreatoblastomas were collected from the surgical pathology archives at Johns Hopkins Hospital, Department of Pathology.
For each resection specimen, the surgical pathology files were also queried for the availability of corresponding preoperative cytology cell blocks. In total, 8 solid-pseudopapillary neoplasms, 29 well-differentiated pancreatic neuroendocrine tumors, 1 high-grade pancreatic neuroendocrine carcinoma, 12 pancreatic ductal adenocarcinoma, and 2 acinar cell carcinomas had sufficient material for ancillary immunohistochemical labeling. In each case, all slides and available immunohistochemical stains were reviewed to confirm the diagnosis. Patient demographic data and gross findings were also recorded.
Immunohistochemistry
Immunohistochemical labeling was performed using 4-μm-thick sections from formalin-fixed and paraffin-embedded tissues. Slides were deparaffinized with serial xylene treatments and subjected to antigen retrieval by heated treatment in citrate solution (pH 6.0). All immunohistochemical labeling was performed on the automated Ventana Benchmark XT system using the biotin-free Ventana OptiView DAB IHC Detection Kit (Ventana Medical Systems, Tucson, AZ, USA). The following antibodies were applied at indicated dilutions: LEF1 (rabbit monoclonal, dilution 1:10, Epitomics, Burlingame, CA, USA), CTNNB1 (monoclonal mouse, dilution 1:250, Dako, Carpinteria, CA, USA), E-cadherin (mouse monoclonal, dilution 1:25, Dako, Carpinteria, CA, USA), CD99 (mouse monoclonal, dilution 1:75, Dako, Carpinteria, CA, USA), and CD10 (rabbit monoclonal, Ventana Medical Systems, Tucson, AZ, USA). Appropriate positive and negative controls were included with each immunolabeling procedure.
Results
The clinicopathologic features of the study cohort are summarized in Table 1. Immunohistochemical staining for LEF1 and CTNNB1 was performed on a total of 27 solid-pseudopapillary neoplasms, 44 well-differentiated pancreatic neuroendocrine tumors, 2 high-grade pancreatic neuroendocrine carcinomas, 25 pancreatic ductal adenocarcinomas, 9 acinar cell carcinomas, and 4 pancreatoblastomas.
Among all 27 (100%) solid-pseudopapillary neoplasms, a strong and diffuse nuclear staining for LEF1 was detected in virtually all neoplastic cells (Figures 1d and 2b). The interspersed foamy macrophages, endothelial cells lining the delicate, branching tumor vessels, and both intracellular and extracellular hyaline globules were negative. The surrounding uninvolved pancreatic parenchyma including the exocrine (acinar and ductal epithelium) and endocrine (islets of Langerhans) components was also devoid of any nuclear labeling (Figure 1c). Consistent with previous reports, LEF1 nuclear staining was positive in paracortical T-cells within lymph nodes present in the peripancreatic soft tissue. For comparison, immunohistochemical stains for CTNNB1, E-cadherin, CD10, and CD99 were also assessed in all 27 solid-pseudopapillary neoplasms (Figures 2a–f). Both nuclear and cytoplasmic accumulation of CTNNB1 (Figures 1f and 2c) and loss of E-cadherin (Figure 2d) were identified in all (100%) cases. In addition, all solid-pseudopapillary neoplasms showed membranous and cytoplasmic immunoreactivity for CD10 (Figure 2e). However, the immunolabeling varied among the tumors with 19 (70%) showing diffuse staining, while the remaining 8 (30%) were patchy in distribution. CD99 was also positive in all cases with a paranuclear dot-like pattern (Figure 2f), but, similar to CD10, which was patchy in 12 (44%) and diffuse in 15 (56%) cases.
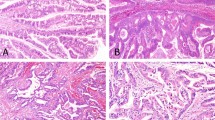
Immunohistochemical staining for LEF1 and CTNNB1 in normal pancreatic parenchyma (a, H&E, × 200) and solid-pseudopapillary neoplasms (b, H&E, × 200). The normal pancreatic parenchyma including endocrine and exocrine components is negative for LEF1 (c, × 200) and shows a membranous staining pattern for CTNNB1 (d, × 200). In contrast, a strong and diffuse nuclear labeling for LEF1 (e, × 200) and abnormal nuclear and cytoplasmic CTNNB1 accumulation (f, × 200) are observed with solid-pseudopapillary neoplasms.
At high magnification, solid-pseudopapillary neoplasms are composed of loosely cohesive cells surrounded by delicate capillaries and interspersed macrophages (a, H&E, × 400). Immunohistochemical stains for LEF1 are diffusely positive within the tumor nuclei, but negative within endothelial cells and macrophages (b, × 400). CTNNB1 shows a membranous staining in both the neoplastic and nonneoplastic components of the tumor; however, nuclear and cytoplasmic accumulation is only seen in the neoplastic cells (c, × 400). In comparison, membranous labeling for E-cadherin is lost within the tumor cells, while retained in the intervening macrophages. Patchy CD10 (e, × 400) and paranuclear dot-like CD99 (f, × 600) staining were also seen within the tumors.
In 8 of 27 solid-pseudopapillary neoplasms, correlative preoperative cytopathology cell block material was available for immunohistochemical staining. In all 8 (100%) cases, the neoplastic cells showed a uniformly, strong nuclear staining for LEF1 (Figure 3b). Although nuclear and cytoplasmic CTNNB1 was identified in all cases; in three specimens, the cell membranes of >50% of the neoplastic cells were disrupted, but the nuclei remained intact. Consequently, immunohistochemical staining for CTNNB1 showed both a nuclear and a dispersed paranuclear pattern of staining (Figure 3c). Similar to the surgical resection material, membranous E-cadherin was absent in all cases. In addition, CD10 was positive in six of eight (75%) specimens, but negative in the remaining two. In these two cases, corresponding surgical specimens showed patchy CD10 immunolabeling. A paranuclear dot-like pattern for CD99 was identified in five of eight (63%) cases (Figure 3d). Of the three CD99-negative cases, corresponding resection specimens were patchy for CD99.
Cytologic cell blocks from solid-pseudopapillary neoplasms often show small, uniform, individual cells with bland nuclei (a, H&E × 400). Both LEF1 (b, × 400) and CTNNB1 immunohistochemical stains demonstrate nuclear positivity. However, during cell block preparation, the tumor cell membranes may be disrupted. This results in a dispersed cytoplasmic staining of CTNNB1, which can impede nuclear interpretation (c, × 400). Similarly, paranuclear dot-like CD99 staining can be difficult to identify (d, × 600) and in some cases be negative.
The 44 well-differentiated pancreatic neuroendocrine tumors, 2 high-grade pancreatic neuroendocrine carcinomas, 25 pancreatic ductal adenocarcinomas, and 9 acinar cell carcinomas were negative for LEF1 (Figure 4). In addition, immunohistochemical labeling for CTNNB1 in these neoplasms revealed a membranous and faint cytoplasmic labeling, but no nuclear or cytoplasmic accumulation. However, nuclear LEF1 and CTNNB1 staining was detected in all four pancreatoblastomas with expression ranging from 10 to 40% of the neoplastic cells. LEF1 staining was primarily centered around and within squamoid corpuscles including neoplastic cells with optically clear nuclei. Preoperative cytopathology cell blocks were available for 29 of 44 well-differentiated pancreatic neuroendocrine tumors, 1 of 2 high-grade pancreatic neuroendocrine carcinomas, 12 of 25 pancreatic ductal adenocarcinomas, and 2 of 9 acinar cell carcinomas. In all cases, both LEF1 and nuclear CTNNB1 were negative. No pancreatoblastoma cell blocks were available for immunohistochemical labeling.
Well-differentiated pancreatic neuroendocrine tumors (a (H&E) and b (immunohistochemical stains), × 200), acinar cell carcinomas (c and d), and pancreatic ductal adenocarcinomas (g and h) were negative for LEF1 (left panel) and showed membranous staining for CTNNB1 (right panel). In contrast, nuclear LEF1 and CTNNB1 staining was detected in pancreatoblastomas, but primarily centered around and within squamoid corpuscles including tumor cells with optically clear nuclei (e and f).
Discussion
While the cell of origin and direction of differentiation of solid-pseudopapillary neoplasms has been a subject of much debate and speculation; recent molecular, genomic, and animal studies have shed light on the pathogenesis of these tumors. In fact, compelling evidence suggest that aberrant Wnt/CTNNB1 is a driver of tumorigenesis. Solid-pseudopapillary neoplasms almost universally harbor gain-of-function mutations in CTNNB1, and mutations in this gene are frequently the only mutation identified after assessment of all coding sequences.9, 10, 12 In mouse models, conditional activation of CTNNB1 in the pancreas results in the formation large tumors that histologically resemble solid-pseudopapillary neoplasms.17 Moreover, many of the CTNNB1/LEF1-responsive genes are known oncogenes, such as cyclin D1 and c-myc.16, 18 Interestingly, some of these responsive genes are located on chromosome 11q22-24 and thus thought to be a region of potential significance for tumorigenesis.19 As described here, LEF1 is diffusely overexpressed in solid-pseudopapillary neoplasms and directly correlates with nuclear and cytoplasmic CTNNB1. This further implicates the CTNNB1/LEF1 transcriptional complex in tumor development.
LEF1 is one of the four members of the LEF1/TCF family of high mobility-group transcription factors.20 Although well-described as a regulator of the Wnt/CTNNB1 signaling pathway, LEF1 was originally identified as a driver of T-cell antigen receptor alpha-chain expression and as being independent from its involvement with Wnt/CTNNB1 signaling.21, 22 Besides CTNNB1, LEF1 cooperates with a diverse number of transcription factors and consequently associated with many signaling pathways including TGFβ and Notch.23, 24 Additionally, under certain circumstances LEF1 can function as a transcriptional repressor.25 Consequently, LEF1 may also be involved in other aspects of tumor biology.
During embryogenesis LEF1 is widely expressed in developing tissues, but restricted to hair follicle bulbs and pre-B and -T lymphocytes in adulthood.26, 27 In contrast, overexpression of LEF1 has been detected in several malignancies including colorectal, breast, prostate, ovarian, oropharyngeal, leukemia, and lymphoma.28, 29, 30, 31, 32, 33, 34 A number of mechanisms or combination thereof may be responsible for elevated LEF1 in tumors. The LEF1 promoter contains numerous consensus LEF1/TCF-DNA binding sequences. In vitro reporter studies have shown that expression of both CTNNB1 and LEF1 results in transactivation of the LEF1 promoter and indicates a positive feedback loop for Wnt/CTNNB1 signaling.35 This would certainly explain the increase in LEF1 protein in solid-pseudopapillary neoplasms as compared with the normal pancreas. Nevertheless, gene expression profiling of solid-pseudopapillary neoplasms have failed to identify elevated LEF1 mRNA levels.16, 18 Interestingly, the mRNA of another LEF1/TCF family member, TCF1/TCF7, is shown to be upregulated in solid-pseudopapillary neoplasms.25 But, in contrast to LEF1, TCF1/TCF7 is thought to be a tumor-suppressor gene. Disruption of TCF1/TCF7 in mouse models results in enhanced activity of other LEF1/TCF proteins and increased susceptibility toward developing gastrointestinal neoplasms and other tumors.36, 37 Another possibility for elevated LEF1 in solid-pseudopapillary neoplasms may be an as yet unidentified post-translational modification extending its protein half-life. Alternatively, increased LEF1 may simply reflect stabilization of the CTNNB1/LEF1 transactivation complex.
Pancreatoblastomas are also known to harbor CTNNB1 mutations and express nuclear CTNNB1.38, 39 Thus, it not surprising that LEF1 would demonstrate a similar pattern of staining. However, expression of both nuclear CTNNB1 and LEF1 were predominantly confined to squamoid corpuscles. Similar to solid-pseudopapillary neoplasms, the histogenesis of squamous corpuscles remains unknown, but hypothesized to represent a peculiar growth pattern.40 Of note, squamoid corpuscles contain optically clear nuclei that are rich in biotin, which commonly show false-positive immunolabeling using the avidin–biotin peroxidase complex methodology.41 As we employed an indirect, biotin-free antibody system, the presence of abundant endogenous biotin should not be an issue. Regardless, squamoid corpuscles are typically a minor component of pancreatoblastomas and the CTNNB1/LEF1 transcriptional complex is unlikely to has a significant role in their tumorigenesis.
Although a comprehensive cohort of pancreatic tumors has not been assessed, the strong and diffuse staining pattern of LEF1 has a high sensitivity and specificity for solid-pseudopapillary neoplasms. Owing to overlapping histologic features in both cytologic and surgical resection specimens, the distinction between solid-pseudopapillary neoplasms and other tumors of the pancreas can be diagnostically challenging. Thus, LEF1 may serve as a useful ancillary tool. Currently available immunohistochemical stains for CD10, CD99, E-cadherin, and CTNNB1 have proven to be reliable markers for solid-pseudopapillary neoplasms, but in some instances may be of limited utility. Both CD10 and CD99 can be negative in cytologic cell blocks and patchy in corresponding resection material. Further, CD10 positivity can be observed in pancreatic tumors that often mimic solid-pseudopapillary neoplasms.42 In addition, tumor cell membranes may be disrupted during cell block preparation, which would preclude proper evaluation of membranous stains. This can be particularly problematic when evaluating for loss of E-cadherin and redistribution of CTNNB1. Many of these issues can be negated using LEF1; however, a note of caution should be taken when evaluating lymph nodes. As mentioned previously, LEF1 is expressed in paracortical T lymphocytes. Hence, LEF1 should be interpreted as part of a panel of immunohistochemical stains.
In summary, abnormal nuclear and cytoplasmic β-catenin accumulation is accompanied by nuclear LEF1 overexpression in both solid-pseudopapillary neoplasms and pancreatoblastomas. However, in contrast to pancreatoblastomas, a diffuse, nuclear labeling was observed in solid-pseudopapillary neoplasms. While the mechanism for LEF1 overexpression in solid-pseudopapillary neoplasms remains unknown, it further implicates the CTNNB1/LEF1 transcriptional complex in tumor development. Furthermore, the absence of LEF1 in potential mimickers of solid-pseudopapillary neoplasms underscores it potential to be a useful ancillary stain.
References
Frantz VK . Tumors of the pancreas. Atlas of Tumor Pathology, section VII, Fascicles 27 and 28, Armed Forces Institute of Pathology: Washington, DC, 1959, pp 32–33.
Reddy S, Cameron JL, Scudiere J et al. Surgical management of solid-pseudopapillary neoplasms of the pancreas (Franz or Hamoudi tumors): a large single-institutional series. J Am Coll Surg 2009;208:950–957 discussion 957–959.
Butte JM, Brennan MF, Gonen M et al. Solid pseudopapillary tumors of the pancreas. Clinical features, surgical outcomes, and long-term survival in 45 consecutive patients from a single center. J Gastrointest Surg 2011;15:350–357.
Martin RC, Klimstra DS, Brennan MF et al. Solid-pseudopapillary tumor of the pancreas: a surgical enigma? Ann Surg Oncol 2002;9:35–40.
Hruban RH, Pitman MB, Klimstra DS . Solid-Pseudopapillary Neoplasms. Tumors of the pancreas. In Atlas of Tumor Pathology, 4th Series, Fascicle 6., American Registry of Pathology: Washington, DC, 2007, pp 231–250.
Kloppel G, Hruban RH, Klimstra DS et al. Solid-pseudopapillary neoplasm of the pancreas In: Bosman FT, Carneiro F, Hruban RH, Theise ND (eds). WHO Classification of Tumours of the Digestive System 4th edn International Agency for Research on Cancer: Lyon, France, 2010, pp 327–330.
Klimstra DS, Wenig BM, Heffess CS . Solid-pseudopapillary tumor of the pancreas: a typically cystic carcinoma of low malignant potential. Semin Diagn Pathol 2000;17:66–80.
Papavramidis T, Papavramidis S . Solid pseudopapillary tumors of the pancreas: review of 718 patients reported in English literature. J Am Coll Surg 2005;200:965–972.
Wu J, Jiao Y, Dal Molin M et al. Whole-exome sequencing of neoplastic cysts of the pancreas reveals recurrent mutations in components of ubiquitin-dependent pathways. Proc Natl Acad Sci USA 2011;108:21188–21193.
Abraham SC, Klimstra DS, Wilentz RE et al. Solid-pseudopapillary tumors of the pancreas are genetically distinct from pancreatic ductal adenocarcinomas and almost always harbor beta-catenin mutations. Am J Pathol 2002;160:1361–1369.
Tanaka Y, Notohara K, Kato K et al. Usefulness of beta-catenin immunostaining for the differential diagnosis of solid-pseudopapillary neoplasm of the pancreas. Am J Surg Pathol 2002;26:818–820.
Tanaka Y, Kato K, Notohara K et al. Frequent beta-catenin mutation and cytoplasmic/nuclear accumulation in pancreatic solid-pseudopapillary neoplasm. Cancer Res 2001;61:8401–8404.
Behrens J, von Kries JP, Kuhl M et al. Functional interaction of beta-catenin with the transcription factor LEF-1. Nature 1996;382:638–642.
Aoki M, Hecht A, Kruse U et al. Nuclear endpoint of Wnt signaling: neoplastic transformation induced by transactivating lymphoid-enhancing factor 1. Proc Natl Acad Sci USA 1999;96:139–144.
Schepers A, Clevers H . Wnt signaling, stem cells, and cancer of the gastrointestinal tract. Cold Spring Harb Perspect Biol 2012;4:a007989.
Park M, Kim M, Hwang D et al. Characterization of gene expression and activated signaling pathways in solid-pseudopapillary neoplasm of pancreas. Mod Pathol advance online publication, 27 September 2013 doi:10.1038/modpathol.2013.154(e-pub ahead of print).
Heiser PW, Cano DA, Landsman L et al. Stabilization of beta-catenin induces pancreas tumor formation. Gastroenterology 2008;135:1288–1300.
Cavard C, Audebourg A, Letourneur F et al. Gene expression profiling provides insights into the pathways involved in solid pseudopapillary neoplasm of the pancreas. J Pathol 2009;218:201–209.
Tiemann K, Heitling U, Kosmahl M et al. Solid pseudopapillary neoplasms of the pancreas show an interruption of the Wnt-signaling pathway and express gene products of 11q. Mod Pathol 2007;20:955–960.
Eastman Q, Grosschedl R . Regulation of LEF-1/TCF transcription factors by Wnt and other signals. Curr Opin Cell Biol 1999;11:233–240.
Giese K, Amsterdam A, Grosschedl R . DNA-binding properties of the HMG domain of the lymphoid-specific transcriptional regulator LEF-1. Genes Dev 1991;5:2567–2578.
Travis A, Amsterdam A, Belanger C et al. LEF-1, a gene encoding a lymphoid-specific protein with an HMG domain, regulates T-cell receptor alpha enhancer function [corrected]. Genes Dev 1991;5:880–894.
Galceran J, Sustmann C, Hsu SC et al. LEF1-mediated regulation of Delta-like1 links Wnt and Notch signaling in somitogenesis. Genes Dev 2004;18:2718–2723.
Medici D, Hay ED, Goodenough DA . Cooperation between snail and LEF-1 transcription factors is essential for TGF-beta1-induced epithelial-mesenchymal transition. Mol Biol Cell 2006;17:1871–1879.
Levanon D, Goldstein RE, Bernstein Y et al. Transcriptional repression by AML1 and LEF-1 is mediated by the TLE/Groucho corepressors. Proc Natl Acad Sci USA 1998;95:11590–11595.
Reya T, O’Riordan M, Okamura R et al. Wnt signaling regulates B lymphocyte proliferation through a LEF-1 dependent mechanism. Immunity 2000;13:15–24.
Petropoulos K, Arseni N, Schessl C et al. A novel role for Lef-1, a central transcription mediator of Wnt signaling, in leukemogenesis. J Exp Med 2008;205:515–522.
Tandon B, Peterson L, Gao J et al. Nuclear overexpression of lymphoid-enhancer-binding factor 1 identifies chronic lymphocytic leukemia/small lymphocytic lymphoma in small B-cell lymphomas. Mod Pathol 2011;24:1433–1443.
Kriegl L, Horst D, Reiche JA et al. LEF-1 and TCF4 expression correlate inversely with survival in colorectal cancer. J Transl Med 2010;8:123.
Nguyen A, Rosner A, Milovanovic T et al. Wnt pathway component LEF1 mediates tumor cell invasion and is expressed in human and murine breast cancers lacking ErbB2 (her-2/neu) overexpression. Int J Oncol 2005;27:949–956.
Li Y, Wang L, Zhang M et al. LEF1 in androgen-independent prostate cancer: regulation of androgen receptor expression, prostate cancer growth, and invasion. Cancer Res 2009;69:3332–3338.
Huang FI, Chen YL, Chang CN et al. Hepatocyte growth factor activates Wnt pathway by transcriptional activation of LEF1 to facilitate tumor invasion. Carcinogenesis 2012;33:1142–1148.
Walther N, Ulrich A, Vockerodt M et al. Aberrant lymphocyte enhancer-binding factor 1 expression is characteristic for sporadic Burkitt’s lymphoma. Am J Pathol 2013;182:1092–1098.
Papagerakis P, Pannone G, Shabana AH et al. Aberrant beta-catenin and LEF1 expression may predict the clinical outcome for patients with oropharyngeal cancer. Int J Immunopathol Pharmacol 2012;25:135–146.
Filali M, Cheng N, Abbott D et al. Wnt-3A/beta-catenin signaling induces transcription from the LEF-1 promoter. J Biol Chem 2002;277:33398–33410.
Yu S, Zhou X, Steinke FC et al. The TCF-1 and LEF-1 transcription factors have cooperative and opposing roles in T cell development and malignancy. Immunity 2012;37:813–826.
Roose J, Huls G, van Beest M et al. Synergy between tumor suppressor APC and the beta-catenin-Tcf4 target Tcf1. Science 1999;285:1923–1926.
Tanaka Y, Kato K, Notohara K et al. Significance of aberrant (cytoplasmic/nuclear) expression of beta-catenin in pancreatoblastoma. J Pathol 2003;199:185–190.
Abraham SC, Wu TT, Hruban RH et al. Genetic and immunohistochemical analysis of pancreatic acinar cell carcinoma: frequent allelic loss on chromosome 11p and alterations in the APC/beta-catenin pathway. Am J Pathol 2002;160:953–962.
Klimstra DS, Wenig BM, Adair CF et al. Pancreatoblastoma. A clinicopathologic study and review of the literature. Am J Surg Pathol 1995;19:1371–1389.
Tanaka Y, Ijiri R, Yamanaka S et al. Pancreatoblastoma: optically clear nuclei in squamoid corpuscles are rich in biotin. Mod Pathol 1998;11:945–949.
Notohara K, Hamazaki S, Tsukayama C et al. Solid-pseudopapillary tumor of the pancreas: immunohistochemical localization of neuroendocrine markers and CD10. Am J Surg Pathol 2000;24:1361–1371.
Acknowledgements
We would like to thank Ms Robyn Roche and Ms Sandra Markowitz for their outstanding administrative assistance. In addition, we also thank Dr Bevan Tandon, Dr Steven H Swerdlow and Dr Samuel A Yousem for helpful comments and suggestions. Funding support: This project was supported in part by a grant from the National Pancreas Foundation, Western Pennsylvania Chapter (to AD Singhi), and P30CA006973.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Singhi, A., Lilo, M., Hruban, R. et al. Overexpression of Lymphoid Enhancer-Binding Factor 1 (LEF1) in solid-pseudopapillary neoplasms of the pancreas. Mod Pathol 27, 1355–1363 (2014). https://doi.org/10.1038/modpathol.2014.40
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/modpathol.2014.40
Keywords
This article is cited by
-
Case report: pseudoendocrine sarcoma, a clinicopathologic report of a newly described soft tissue neoplasm
Virchows Archiv (2023)
-
Immunohistochemical staining of LEF-1 is a useful marker for distinguishing WNT-activated medulloblastomas
Diagnostic Pathology (2022)
-
Clinicopathological features and prognostic significance of CTNNB1 mutation in low-grade, early-stage endometrial endometrioid carcinoma
Virchows Archiv (2021)
-
RETRACTED ARTICLE: A novel microRNA identified in hepatocellular carcinomas is responsive to LEF1 and facilitates proliferation and epithelial-mesenchymal transition via targeting of NFIX
Oncogenesis (2018)
-
LEF-1 is a Sensitive Marker of Cribriform Morular Variant of Papillary Thyroid Carcinoma
Head and Neck Pathology (2018)