Abstract
Static fermentation of amidepsine-producing fungus Humicola sp. FO-2942 led to the production of six new amidepsines, including a new type of glycosylated congener. Non-glycosylated amidepsine J inhibited both human diacylglycerol acyltransferases 1 (DGAT1) and DGAT2 with the same IC50 value of 40 μM, whereas glycosylated amidepsines F to I showed very weak inhibitory activity against DGAT1 and DGAT2.
Similar content being viewed by others
Introduction
Diacylglycerol acyltransferase (DGAT) catalyzes the esterification of diacylglycerol with long-chain acyl-CoA to synthesize triacylglycerol. DGAT is actually two isozymes, DGAT1 and DGAT2,1, 2 and is considered a potential target for the treatment of obesity or metabolic disorder.1, 2, 3 Accordingly, researchers have focused on the discovery and development of DGAT inhibitors.4, 5
Amidepsines, gyropholic acid derivatives produced by Humicola sp. FO-29426, 7 and Humicola sp. FO-5962,8 showed inhibitory activity against DGAT. Recently, we completed the total synthesis of amidepsine B (2), which consists of gyropholic acid and alanine moieties, and elucidated its absolute stereochemistry.9
To further study the biological characteristics of amidepsines, a large amount of the sample is required, but the original productivity of amidepsine by fermentation of strain FO-2942 was very low (0.1–0.6 μg ml−1); therefore, we started to optimize the fermentation conditions to increase the productivity of amidepsines. We found that the production of amidepsines A (1) to D (3) increased 60- to 480-fold when the strain was cultured under static conditions. Furthermore, six new amidepsine congeners (6–11), including a new type of amidepsines F (6) to I (9) having a sugar moiety (Figure 1), were also isolated from culture broth obtained under this condition. In this study, the fermentation, isolation, structure elucidation and biological properties of new amidepsines are described.
Results
Effects of culture conditions on the production of amidepsines
The efficient culture conditions for amidepsine production were examined. The productivity of amidepsines was monitored by HPLC and compared with that obtained under original condition a.1 As shown in Table 1, the production of 1–4 increased 2.6- to 11-fold when fermented under culture condition b. Note that production markedly increased 65- to 480-fold under static culture condition 3. Furthermore, new peaks (6–9, 10 and 11 in Figure 2a), having a UV spectrum similar to that of amidepsines, were found by HPLC analysis (Figure 2a), leading to the isolation of new amidepsines described below.
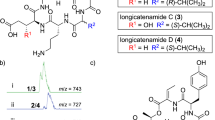
HPLC profiles of amidepsines. (a) HPLC profile of ethyl acetate extracts of the broth obtained from Humicola sp. FO-2942. One μg of ethyl acetate extracts (1 mg ml−1 of methanol) was subjected to HPLC under the conditions as described in ‘Methods.’ (b) Chromatographic profile of 1–4 and 10–12 by analytical HPLC. (c) Chromatographic profile of 6–9 by analytical HPLC.
The time course of amidepsine production under culture condition c is shown in Figure 3. The production of all amidepsines started on day 3. The concentration of 1, the most abundant component, peaked (144 μg ml−1) on day 9. The production of nonglycosylated amidepsines also peaked on day 9, whereas the production of glycosylated 6–9 peaked on day 5. Compound 6 showed the highest concentration (50 μg ml−1) among glycosylated amidepsines.
Isolation of amidepsines F (6) to I (9)
Compounds 6–9 and gyrophoric acid were isolated together with 1–4 from culture broth obtained under condition c. The whole broth (5.0 l) was extracted with an equal volume of ethyl acetate. The extracts were concentrated in vacuo to dryness to yield a brown oily material (3.8 g). The material was dissolved in CH3OH (400 ml) and extracted with hexane. The methanol layer was concentrated in vacuo to dryness to yield a yellow powder (1.2 g). The material was dissolved in CHCl3, applied on a silica gel column (40 g), and eluted stepwise with 100:0, 98:2, 95:5, 9:1, 8:2 and 0:100 (v/v) of CHCl3-CH3OH solvents (400 ml for each solvent).
The 9:1 fraction (CHCl3-CH3OH), which contained 1–4 and 10–12 (Figure 2b), was concentrated to give a red-brown oily material (340 mg). The oil (50 mg) was finally purified with preparative HPLC (column, PEGASIL ODS (Senshu Scientific, Tokyo, Japan), 20 × 250 mm; solvent, a 60-min linear gradient from 50 to 80% CH3CN; detection, UV at 210 nm; flow rate, 8.0 ml min−1). Under these conditions, 1–4 and 10–12 were eluted as peaks with retention times of 33, 44, 49, 52, 46, 55 and 45 min, respectively. Each fraction was concentrated in vacuo to dryness to give pure 1 (9.4 mg), 2 (4.1 mg), 3 (2.0 mg), 4 (2.0 mg), 10 (3.8 mg), 11 (2.4 mg) and 12 (3.7 mg) as colorless materials.
The CH3OH fraction containing 6–9 (Figure 2c) was concentrated to yield oily material (147.0 mg), which was subjected to ODS column chromatography (10 g), and eluted stepwise with 5:95, 1:9, 2:8, 1:1 and 0:100 (v/v) of CH3CN–H2O solvents (50 ml for each solvent). The 1:1 fraction was concentrated to give a brown material (66.9 mg) that was dissolved in CH3OH (3.3 ml) and the soluble supernatant was purified by HPLC (column, PEGASIL ODS, 20 × 250 mm; solvent, 35% CH3CN; UV at 210 nm; flow rate, 9 ml min−1). Under these conditions, 6–9 were eluted as peaks with retention times of 45, 61, 80 and 113 min, respectively. Each fraction was concentrated in vacuo to dryness to yield pure 6 (2.4 mg), 7 (4.2 mg), 8 (4.8 mg) and 9 (5.3 mg) as colorless materials.
Physicochemical properties of amidepsines F (6) to K (11)
Physicochemical properties of 6–11 are summarized in Table 2. They showed similar absorption maxima at 203–213 and 251–281 nm in the UV spectra, suggesting the presence of the same chromophore in their structures. Compounds 6–10 had optical rotations, whereas 11 had no optical rotation, suggesting the achiral structure of 11.
Structures of amidepsines J (10) and K (11)
Comparison of spectral data between 10 and 3 indicated that amidepsine J (C31H33NO11) is larger by a CH2 unit than 3 (Table 2), and that the methoxy proton (δ 3.83) was observed from the 1H-NMR spectra for 10 (Table 3). In heteronuclear multiple bond correlation (HMBC) experiments (Figure 4a), long-range coupling was observed from a methoxy proton to C-19 (δ 158.3), indicating that 10 has a methoxy group instead of hydroxyl group C-19 of 3. Thus, 10 is elucidated as 19-methoxy-3 (Figure 1).
Comparison of the spectral data of 4 and 11 indicated that 11 is smaller by a CH2 unit than 4 (Table 2), and that the 19-OH (δ 10.37) proton was observed for 11 from NMR spectra (Table 3). The structure was confirmed by HMBC experiments, as shown in Figure 4b, in which long-range couplings were observed from 19-OH (δ 10.37) to C-18 (δ 106.7), C-19 (δ 159.1) and C-20 (δ 99.0), indicating that 11 has a hydroxyl group instead of the methoxy group at C-19 of 4. Thus, 11 is elucidated as 19-hydroxy-4 (Figure 1).
Structure elucidation of amidepsines G (7)
The molecular formula of 7 was determined to be C34H37NO16 on the basis of HRFAB-MS measurements (Table 2), indicating that 7 is larger by a C6H10O5 unit than 2. Further, 1D NMR spectral data and the physicochemical properties showed that 7 has the same skeleton as 2. The 13C NMR spectrum of 7 exhibited six signals of a sugar moiety at δ 62.0, 70.5, 74.0, 78.5, 78.7 and 102.5. These data, together with one anomeric proton signal at δ 5.05 (C-1′), were indicative of the presence of a sugar moiety, which was confirmed by 1H-1H COSY and HMBC experiments (Figure 4c). The sugar moiety was assigned as glucose by 3JHH coupling constants between C-1′ and C-2′ (7.2 Hz), C-2′ and C-3′ (9.0 Hz), C-3′ and C-4′ (8.4 Hz), and C-4′ and C-5′ (9.6 Hz), as shown in Figure 5. The connectivity of the glucose moiety was determined by observing the correlation between the anomeric proton signal (δ 5.05) and the corresponding carbon signal C-11 (δ 156.4) in the HMBC experiment to form a glycoside bond (Figure 4c). Thus, the structure of 7 was elucidated as shown in Figure 1.
Structures of amidepsines F (6), H (8) and I (9)
Comparison of the spectral data between 6 and 7 indicated that 6 is larger by a CH2 unit than 7 (Table 2), and that the methoxy proton (δ 3.88) was observed for 6 in the 1H NMR spectra (Table 3). In HMBC experiments (Figure 4e), long-range coupling was observed from the methoxy proton to C-19 (δ 160.2), indicating that 6 has a methoxy group instead of the hydroxy group at C-19 of 7. Thus, the structure of 6 was determined as 19-methoxy-7 (Figure 1).
The molecular formula of 9 is C36H40O17N, indicating that it is larger by a C2H2O unit than 7. The 1H-NMR spectrum of 9 was almost identical to that of 7 except for the proton signal H-2′(δ 5.06). In the HMBC spectrum of 9 (Figure 4d), the correlation between H3-8′(δ 2.00) and H-2′ to C-7′(δ 171.9) indicated that an O-acetyl group was connected to the C-2′ of 7. Thus, the structure of 9 was determined as shown in Figure 1.
Comparison of the spectral data between 8 and 9 indicated that 8 is larger by a CH2 unit than 9 (Table 2) and that the methoxy proton (δ 3.84) was observed for 8 (Table 3). In HMBC experiments (Figure 4f), long-range coupling was observed from methoxy proton to C-19 (δ 160.2), indicating that 8 has a methoxy group instead of the hydroxy group at C-19 of 9; therefore, the structure of 8 was determined as 19-methoxy-9 (Figure 1).
Effects of amidepsines on human DGAT1 and DGAT2 activities
Effects of 9–11 on human DGAT1 and DGAT2 activity (IC50 value) are summarized in Table 4. Glycosylated 6–9 showed very weak or almost no inhibitory activity against human DGAT1 and DGAT2; however, nonglycosylated 10 inhibited both isozymes with the same IC50 value of 40 μM, indicating that the potency is analogous to that of 1, 2 and 4.
Discussion
In this study, we found that when Humicola sp. FO-2942 was cultured under static conditions, the production of 1–4 increased markedly about 65- to 480-fold. At the same time, new 6–11 were isolated from the culture broth. From the structure elucidation, 6–9 were a new type of amidepsines having a sugar moiety at C-11. Interestingly, amidepsine E (5) produced by Humicola sp. FO-59693 was not detected in the culture of strain FO-2942.
Glycosylated 6–9 were produced only under static conditions and their yields peaked on day 5 after inoculation; however, the production of nonglycosylated amidepsines continued to increase until day 9. It is easy to speculate that amidepsine A/B is glycosylated to produce 6/7, and then acetylated at position 2 of the sugar residue to produce 8/9; therefore, glycosylation and acetylation occurred at a specific time (until day 5) after inoculation.
Compounds 1–4 are dual DGAT inhibitors with similar IC50 values for DGAT1 and DGAT2.7 Nonglycosylated 10 also has a similar characteristic. On the other hand, glycosylated 6–9 showed very weak or almost no activity against the two isozymes, indicating that the presence of a sugar moiety at C-11 is not compatible with DGAT inhibition.
Methods
Materials
Strain and culture conditions. Fungal strain Humicola sp. FO-2942 was used for the production of amidepsines.1, 2 Strain FO-2942 was grown and maintained on an agar slant containing 0.1% glycerol, 0.08% KH2PO4, 0.02% K2HPO4, 0.02% MgSO4–7H2O, 0.02% KCl, 0.2% NaNO3, 0.02% yeast extract and 1.5% agar (pH 6.0). A loopful of spore of strain FO-2942 was inoculated into seed medium consisting of 2.0% glucose, 0.5% polypeptone, 0.05% MgSO4–7H2O, 0.2% yeast extract, 0.1% KH2PO4 and 0.1% agar (pH 6.0) in a 500 ml Erlenmeyer flask. The inoculated flasks were shaken on a rotary shaker for 4 days at 27 °C to obtain the seed culture, which was transferred to production medium consisting of 2.0% sucrose, 1.0% glucose, 1.0% corn steep liquor, 0.5% meat extract, 0.1% KH2PO4, 0.05% MgSO4 7H2O, 1.0 ml of trace elements contained in g l−1: 1.0 FeSO4–7H2O, 1.0 MnCl2–4H2O, 1.0 ZnSO4–7H2O, 1.0 CuSO4–5H2O and 1.0 CoCl2–2H2O, CaCO3 0.3% and 0.1% agar (pH 6.0) and the fermentation was carried out under the following culture conditions: (a) jar fermentation: the seed culture (500 ml) was transferred to the production medium (30 l) in a 50-l jar fermenter. The fermentation was carried out at 27 °C for 3 days as previously described.1 (b) Shaking and static culture: the seed culture (1 ml) was transferred into a 500-ml Erlenmeyer flask containing the production medium (100 ml). The first culture was carried out at 27 °C for 3 days on a rotary shaker (200 r.p.m.), and then the second culture was carried out at 27 °C for an additional 7 days under static conditions. (c) Static culture: the seed culture (1 ml) was transferred to the production medium (100 ml) in a 500-ml Erlenmeyer flask. The culture was carried out at 27 °C for 10 days under static conditions. Compounds 6–11 were obtained under culture condition c.
General experimental procedures
Kieselgel 60 (Merck KGaA, Darmstadt, Germany) and SSC-ODS-7515-12 (Senshu Scientific) were used for silica gel and octadecyl silyl (ODS) column chromatography, respectively. HPLC was carried out using a LaChrom Elite HTA system (Hitachi High-Technologies, Tokyo, Japan). To determine the amount of amidepsine in culture broth, ethyl acetate extracts were dissolved in methanol and analyzed by LC-UV (HP1100 systems; Agilent Technologies, Santa Clara, CA, USA) under the following conditions; Pegasil ODS/5 mm column (4.6 × 250 mm; Senshu Scientific), a 30-min linear gradient from 5% CH3CN/0.05% H3PO4 to 100% CH3CN/0.05% H3PO4, 0.7 ml min−1, and UV at 210 and 260 nm. Compounds 1–4 and 6–12 were eluted as peaks with retention times of 24.3, 24.8, 26.5, 27.3, 20.3, 20.7, 21.1, 21.6, 25.4, 28.4 and 25.0 min, respectively (Figure 2a).
Ultraviolet spectra were recorded on a spectrophotometer (DU640; Beckman Coulter, Fullerton, CA, USA). IR spectra were recorded on a Fourier transform infrared spectrometer (FT-710; HORIBA, Kyoto, Japan). Optical rotations were measured with a digital polarimeter (DIP-1000; JASCO, Tokyo, Japan). EI-MS spectra and HREI-MS spectra were recorded on a mass spectrometer (JMS-AX505HA; JEOL, Tokyo, Japan). Various NMR spectra were collected with a spectrometer (XL-400; Varian, Palo Alto, CA, USA).
Assay for diacylglycerol acyltransferase
The activity of DGAT1 and DGAT2 was measured in an enzyme assay using microsomal fractions prepared from Saccharomyces cerevisiae expressing human DGAT1 and DGAT2, as previously reported.4, 7
References
Tomoda, H. & Ōmura, S. Potential therapeutics for obesity and atherosclerosis: inhibitors of neutral lipid metabolism from microorganisms. Pharmacol. Ther. 115, 375–389 (2007).
Matsuda, D. & Tomoda, H. DGAT inhibitors for obesity. Curr. Opin. Investig. Drugs 8, 836–841 (2007).
Yen, C. L., Stone, S. J., Koliwad, S., Harris, C. & Farese, R. V. Jr. DGAT enzymes and triacylglycerol biosynthesis. J. Lipid Res. 49, 2283–2301 (2008).
Inokoshi, J. et al. Expression of two human acyl-CoA:diacylglycerol acyltransferase isozymes in yeast and selectivity of microbial inhibitors toward the isozymes. J. Antibiot. 62, 51–54 (2009).
Zammit, V. A., Buckett, L. K., Turnbull, A. V., Wure, H. & Proven, A. Diacylglycerol acyltransferases: potential roles as pharmacological targets. Pharmacol. Ther. 118, 295–302 (2008).
Tomoda, H. et al. Amidepsines, inhibitors of diacylglycerol acyltransferase produced by Humicola sp. FO-2942. I. Production, isolation and biological properties. J. Antibiot. 48, 937–941 (1995).
Tomoda, H., Tabata, N., Ito, M. & Ōmura, S. Amidepsines, inhibitors of diacylglycerol acyltransferase produced by Humicola sp. FO-2942. II. Structure elucidation of amidepsines A, B and C. J. Antibiot. 48, 942–947 (1995).
Tomoda, H. et al. Amidepsine E, an inhibitor of diacylglycerol acyltransferase produced by Humicola sp. FO-5969. J. Antibiot. 49, 929–931 (1996).
Nagamitsu, T. et al. Total synthesis of amidepsine B and revision of its absolute configuration. J. Antibiot. 62, 69–74 (2009).
Acknowledgements
We express our thanks to Ms N Sato for NMR experiments, and to Ms T Sakabe and Ms A Nakagawa for mass spectra.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Inokoshi, J., Takagi, Y., Uchida, R. et al. Production of a new type of amidepsine with a sugar moiety by static fermentation of Humicola sp. FO-2942. J Antibiot 63, 9–16 (2010). https://doi.org/10.1038/ja.2009.110
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ja.2009.110
Keywords
This article is cited by
-
Natural Products of the Fungal Genus Humicola: Diversity, Biological Activity, and Industrial Importance
Current Microbiology (2021)