Abstract
To infer the histories of population admixture, one important challenge with methods based on the admixture linkage disequilibrium (ALD) is to remove the effect of source LD (SLD), which is directly inherited from source populations. In previous methods, only the decay curve of weighted LD between pairs of sites whose genetic distance were larger than a certain starting distance was fitted by single or multiple exponential functions, for the inference of recent single- or multiple-wave admixture. However, the effect of SLD has not been well defined and no tool has been developed to estimate the effect of SLD on weighted LD decay. In this study, we defined the SLD in the formularized weighted LD statistic under the two-way admixture model and proposed a polynomial spectrum (p-spectrum) to study the weighted SLD and weighted LD. We also found that reference populations could be used to reduce the SLD in weighted LD statistics. We further developed a method, iMAAPs, to infer multiple-wave admixture by fitting ALD using a p-spectrum. We evaluated the performance of iMAAPs under various admixture models in simulated data and applied iMAAPs to the analysis of genome-wide single nucleotide polymorphism data from the Human Genome Diversity Project and the HapMap Project. We showed that iMAAPs is a considerable improvement over other current methods and further facilitates the inference of histories of complex population admixtures.
Similar content being viewed by others
Introduction
The ‘Out of Africa’ human migrations resulted in population differentiation in different continents, while subsequent migrations over the past millennia led to gene flow among previously separated human sub-populations. As a consequence, admixed populations came into being when previously mutually isolated populations met and intermarried. Population admixture has received a great deal of attention recently. Many studies based on genome-wide data have shown that gene flow is common among inter-continental and intra-continental populations, and that population admixture often leads to extended linkage disequilibrium (LD), which can greatly facilitate the mapping of human disease genes (McKeigue, 2005; Reich and Patterson, 2005; Smith and O’Brien, 2005).
High levels of LD are produced by admixture at loci that have different allele frequencies among the involved populations (Nei and Li, 1973). Because of recombination, this particular type of admixture LD (ALD) decays as a function of time since admixture. Consequently, it is possible to infer population admixture by modeling the dynamic changes of ALD. Moorjani et al. (2011) proposed such an approach by aggregating pairwise LD measurements through a weighting scheme. Its software, ROLLOFF, was fully explained by Patterson et al. (2012) and further developed as ALDER by Loh et al. (2013) and Pickrell et al (2014). This ALD-based approach is particularly useful for admixture dating.
Under the hybrid isolation (HI) model, the expected value of LD decreases at a rate of 1−d (Chakraborty and Weiss, 1988; Pfaff et al., 2001), where d is the genetic distance (in Morgan) between two sites. In addition, after g generations, the LD decays to (1−d)g of its original value, assuming that the admixed population is engaged in random mating and has infinite effective population size (Hill and Robertson, 1966). Recently, Pickrell et al. (2014) considered the situation of multiple waves of admixture from different source populations and showed that LD comprised multiple exponential terms, each of which refers to a single admixture event (Pickrell et al., 2014). Zhou et al. (2017) confirmed the polynomial expression (taking e−ld as the approximation of (1−d)l) for each admixture wave and added the effect of source LD (SLD) from source populations into the LD’s expression under the general admixture model. Based on this LD framework, dating admixture becomes a problem of fitting the polynomial terms in the ALD decay.
When dating admixture in empirical populations, two major factors affect the accuracy of estimation: background LD (or SLD in the context of this work) and representative reference populations. Pickrell et al. (2014) presented a method based on weighted LD to deal with multiple-wave admixture. In their method, they used a starting distance strategy (abandon loci whose genetic distance is shorter than a certain distance) to reduce the bias caused by SLD and scanned global populations to determine the best pair of reference populations for each admixture wave. The key assumption of their method is that the only effect by different pairs of reference populations resulted from the relative value of exponential/polynomial coefficients of weighted LD decay. However, they neither validated this assumption nor considered the possible effect from SLD.
Here we introduced the polynomial spectrum (p-spectrum), the fitting results with polynomial functions, to reveal the polynomial property of the weighted LD decay. With a simulated admixed population, we confirmed that the weighted LD decay curves with different pairs of source populations had similar p-spectrums and also found that a starting distance strategy could only partly reduce the SLD (Figure 3).
An alternative way to reduce SLD is to use ancestral source populations to estimate SLD (Zhou et al., 2017). Based on this idea, we developed a new approach to infer multiple-wave admixture, and implemented it in a method called iMAAPs, which infers multiple-wave admixture by fitting ALD using a p-spectrum. After evaluating this method under various admixture models, we applied it to the well-known admixed populations in Human Genome Diversity Project (HGDP; Rosenberg et al., 2002) and HapMap (The International HapMap Consortium, 2010) data, and demonstrated that this current study greatly facilitates the understanding of admixture history of human populations.
Materials and methods
Data sets
Data for simulation and empirical analysis were obtained from two public resources: the HGDP (Rosenberg et al., 2002) and the International HapMap Project phase III (The International HapMap Consortium, 2010). Data filtering was performed within each population with Plink (Wigginton et al., 2005): samples with missing rate >5% per individual, single-nucleotide polymorphisms with missing rate > 50% and single-nucleotide polymorphisms failing the Hardy–Weinberg equilibrium test (P-value <1 × 10−6) were permanently removed from subsequent analyses.
The abbreviation of populations used in this study are as follows: YRI, the Yoruba in Ibadan, Nigeria; LWK, Luhya in Webuye, Kenya; MKK, Maasai in Kinyawa, Kenya; ASW, African Ancestry in SW USA; CEU, US Utah residents with ancestry from northern and western Europe; TSI, Tuscans in Italy; MXL, Mexican Ancestry in LA, CA, USA; JPT (Japanese in Tokyo, Japan); CHB, Han Chinese in Beijing, China; and CHD, Chinese in metropolitan Denver, CO, USA. Haplotypes used as source populations in simulations were from 113 unrelated CEU individuals and 113 unrelated YRI individuals.
Simulations
To evaluate our method in dating admixture, we employed forward-time simulations to generate haplotypes under variant admixture scenarios: HI model, two-wave (TW) model (including the cases of one donor population and two donor populations for the second wave admixture) and the model of isolation after a period of continuous admixture. Our simulations were under the framework of a copying model, where new haplotypes are assembled from the segments of the source populations’ haplotypes generation by generation (Li and Stephens, 2003; Price et al., 2009). This has been used in previous work (Price et al., 2009). In our simulation, no mutation was considered when generating new haplotypes.
Under the HI model, admixture events were set as having occurred 20, 50, 100 and 200 generations ago. For the TW model, the simulated admixed population experienced two waves of admixture, which were at times of 100 and 20 generations ago, respectively, and was isolated in the other time. For the more recent admixture in the TW model, we simulated a scenario in which only one of the source populations donated genetic materials (TW-1 model) and the other scenario where both source populations provided gene flow (TW-2 model).
We also simulated admixed haplotypes in the scenarios of continuous migration, in which only gene flow from source populations to the admixed population was allowed and after that the admixed population was isolated outside the window of continuous migration. In our simulation, we used modified gradual admixture (Jin et al., 2012) and continuous gene flow (Pfaff et al., 2001) models to shape the gene flow in the migration window, which separately resulted in the gradual admixture-I model and continuous gene flow-I model. Under these two models, we set the window size of migration as 80 generations and the isolation duration as 20 generations for the long lasting migration; conversely, we set the window size of migration as 30 generations and the isolation duration as 70 generations for the short lasting migration.
Source populations also evolved in isolation so that both the reference populations and admixed population were of the same age. The sample sizes for both source populations and admixed populations were set as 5000. More details of simulation parameters are given in Supplementary Tables S1–S3.
Weighted LD statistic and its estimator under the two-way admixture model
Under the two-way admixture model (Figure 1), two source populations provide genetic materials to the newly formed admixed population. Following the notations of Zhou et al. (2017), the LD in the admixed population of (n+1)-st generation is composed of SLD and the admixture created LD:

where mi is the genetic proportion derived from the source population i, serving as the weight for linear combination of Di(x,y) (LD in source populations i) to form the SLD; δ12(x) is the allele frequency difference between population 1 and population 2 at site x; d is the genetic distance between site x and site y; and c(l) is a natural admixture indicator whose positive value means that admixture occurred at l generations ago. c(l) is defined as follows:

where wi(l) is the total genetic contribution from source population i in the admixed population B(l). In our notation system, ‘0’ in the subscript represents the admixed population and ‘1’ and ‘2’ represents the two source populations.
Using allele frequency difference δ12(x)δ12(y) as weight, the weighted LD statistic is defined as the average LD with the weight over a set holding pairs of single-nucleotide polymorphisms whose pairwise genetic distance is d (Loh et al., 2013):

where

and ɛ is a discretization parameter inducing a discretization on d. By summing over both sides of Equation (1) weighted by δ12(x)δ12(y) over the set S(d), we have

where

The estimators for the weighted LD statistic for the admixed population and source populations are given by Loh et al. (2013):

where  is the observed allele frequency difference and
is the observed allele frequency difference and  is the estimator of Di(x,y) on the modern data in population i. For the source populations,
is the estimator of Di(x,y) on the modern data in population i. For the source populations,  , and
, and  is a biased estimator when the same group of samples are used for calculating both the LD and its weight. Fortunately, there are two ways to eliminate the bias: (1) divide the target population into two groups, where one group is used for calculating the allele frequency difference, whereas the other is used for calculating the LD (Moorjani et al., 2011); (2) employ the unbiased statistics (Loh et al., 2013). In this study, we used the second method to correct the bias in the SLD estimation. Besides, F(d) can be independently estimated by
is a biased estimator when the same group of samples are used for calculating both the LD and its weight. Fortunately, there are two ways to eliminate the bias: (1) divide the target population into two groups, where one group is used for calculating the allele frequency difference, whereas the other is used for calculating the LD (Moorjani et al., 2011); (2) employ the unbiased statistics (Loh et al., 2013). In this study, we used the second method to correct the bias in the SLD estimation. Besides, F(d) can be independently estimated by

Here we separated F(d) from the coefficients of polynomial functions to avoid its possible influence on polynomial fitting in our later discussion.
Factorizing weighted LD statistic with polynomial functions
Based on the formula of weighted LD statistics in the admixed population (Equation (2)), admixture events are recorded in the polynomial function  , where a positive value of c(l) indicates the admixture at l generations ago. Therefore, the direct way to date the admixture is to determine the positive value of c(l). However, two possible risks may affect the results when fitting a0(d) with polynomial functions:
, where a positive value of c(l) indicates the admixture at l generations ago. Therefore, the direct way to date the admixture is to determine the positive value of c(l). However, two possible risks may affect the results when fitting a0(d) with polynomial functions:  , which represents the SLD, and F(d), which is a decaying function as d increases (Supplementary Figure S3). Inspired by the Weierstrass approximation theorem, which says continuous curves can be approximated by polynomial functions, we used polynomial functions to approximate
, which represents the SLD, and F(d), which is a decaying function as d increases (Supplementary Figure S3). Inspired by the Weierstrass approximation theorem, which says continuous curves can be approximated by polynomial functions, we used polynomial functions to approximate  and F(d) to explore the possible interaction between them. In fact, by fitting the decay curve with polynomial function ∑l∈Sg b(l)(1−d)l, we obtained the spectrum of b(l) values on set Sg, which we defined as the p-spectrum (Figure 2). In the polynomial fitting, b(l) must be non-negative and Sg is a finite set holding the candidate time points for the possible admixture signals. This numeric method to generate the p-spectrum is illustrated in the Appendix. Replacing ai(d),i=1,2, and F(d) with polynomial functions
and F(d) to explore the possible interaction between them. In fact, by fitting the decay curve with polynomial function ∑l∈Sg b(l)(1−d)l, we obtained the spectrum of b(l) values on set Sg, which we defined as the p-spectrum (Figure 2). In the polynomial fitting, b(l) must be non-negative and Sg is a finite set holding the candidate time points for the possible admixture signals. This numeric method to generate the p-spectrum is illustrated in the Appendix. Replacing ai(d),i=1,2, and F(d) with polynomial functions
P-spectrum for  in a simulated admixed population. The observed weighted LD decay (gray points in top right) are fitted by hundreds of polynomial functions (gray curves in the bottom panel, each curve connecting to the position l represents the decay of the function (1−d)l, with the d value ranging from 0 to 0.7 Morgan), and a few of them whose coefficients are positive (highlighted in heat color). The amplitudes for each positive coefficient are plotted along the value of l (generations ago) in the top left.
in a simulated admixed population. The observed weighted LD decay (gray points in top right) are fitted by hundreds of polynomial functions (gray curves in the bottom panel, each curve connecting to the position l represents the decay of the function (1−d)l, with the d value ranging from 0 to 0.7 Morgan), and a few of them whose coefficients are positive (highlighted in heat color). The amplitudes for each positive coefficient are plotted along the value of l (generations ago) in the top left.

a0(d) turns to be

This expression of a0(d) tells us the linear combination of ai(d),i=1,2, would bring in false-positive admixture signals, whereas F(d) has the potential to destroy the admixture time inference when we try to fit a0(d) directly with polynomial functions. Therefore, it is essential to evaluate the effect of ai(d),i=1,2 and F(d). Fortunately, ai(d),i=1,2 and F(d) could be estimated with the source populations for effect evaluation.
To evaluate the effects of ai(d),i=1,2 and F(d), we simulated a 100-generation-old admixed population under the HI model, and the simulated admixed population was initiated with the haplotype of YRI and CEU of the proportion 50%:50%. Derived populations from YRI and CEU were also generated, separately. Based on the simulated genotype data in both source populations and the admixed population, both  and
and  could be calculated through a Fast Fourier Transform algorithm, which can increase computational efficiency (Loh et al., 2013). Then, the p-spectrum was constructed on a time set ranging from 0 to 2000 generations accordingly. In the spectrum of
could be calculated through a Fast Fourier Transform algorithm, which can increase computational efficiency (Loh et al., 2013). Then, the p-spectrum was constructed on a time set ranging from 0 to 2000 generations accordingly. In the spectrum of  , we found three bunches of signals: 2 sharp bunches appeared around 100 and 1250 generations, and 1 flat bulb lay around 180 generations (Figure 2). In both
, we found three bunches of signals: 2 sharp bunches appeared around 100 and 1250 generations, and 1 flat bulb lay around 180 generations (Figure 2). In both  and
and  spectrums, we found signals around 1250 generations and signals close to 250 generations (Supplementary Figures S1 and S2). In addition, in the p-spectrum of
spectrums, we found signals around 1250 generations and signals close to 250 generations (Supplementary Figures S1 and S2). In addition, in the p-spectrum of  , we found only a strong peak at time 0 and two weak signal peaks over 250 generations ago, which explained the sharp decay in its decay curve (Supplementary Figure S3), suggesting that we must consider this effect to precisely resolve admixture. In the time spectrum of
, we found only a strong peak at time 0 and two weak signal peaks over 250 generations ago, which explained the sharp decay in its decay curve (Supplementary Figure S3), suggesting that we must consider this effect to precisely resolve admixture. In the time spectrum of  , signals around 100 could be explained easily by the designed admixture and both signals around 1250 and signals around 180 were probably introduced by the SLD. To test this explanation, we directly constructed z(d) from Equation (2) as
, signals around 100 could be explained easily by the designed admixture and both signals around 1250 and signals around 180 were probably introduced by the SLD. To test this explanation, we directly constructed z(d) from Equation (2) as

which can be estimated with the simulated admixed population and derived source populations by

In the p-spectrum of  , the relative strength of noise-like signals outside the bunch of signals around 100 generations became much weaker than that in the p-spectrum of
, the relative strength of noise-like signals outside the bunch of signals around 100 generations became much weaker than that in the p-spectrum of  (Supplementary Figure S4). This result confirms our explanation of the p-spectrum of
(Supplementary Figure S4). This result confirms our explanation of the p-spectrum of  , and indicates that source populations can be used to reduce the effect of SLD so that
, and indicates that source populations can be used to reduce the effect of SLD so that  can be used for admixture time inference. In the next section, we discuss how to use the p-spectrum of
can be used for admixture time inference. In the next section, we discuss how to use the p-spectrum of  for admixture dating.
for admixture dating.
Time inference for multiple-wave admixture
As c(l) is a natural indicator of admixture events, the natural extension for the p-spectrum of  is to infer the admixture time. In empirical populations, both the
is to infer the admixture time. In empirical populations, both the  and
and  can be calculated based on the genotype data of the admixed population and reference populations. Meanwhile, the population admixture proportions were estimated by
can be calculated based on the genotype data of the admixed population and reference populations. Meanwhile, the population admixture proportions were estimated by

Then,  could be calculated so as its p-spectrum {c(l)}l∈Sg.
could be calculated so as its p-spectrum {c(l)}l∈Sg.
Next, we dated the admixture and evaluated the existence for each admixture wave using a Jackknife-based method. Suppose we have 22 autosomes for the target admixed population and each chromosome is excluded one at each time to calculate the decay curve of  (Loh et al., 2013). This means when chromosome i is excluded, the remaining 21 chromosomes are used to calculate
(Loh et al., 2013). This means when chromosome i is excluded, the remaining 21 chromosomes are used to calculate  and the p-spectrum {ci(l)}l∈Sg. Then the P-values are attained on each l with a one-sided t-test by testing whether the mean value for the set
and the p-spectrum {ci(l)}l∈Sg. Then the P-values are attained on each l with a one-sided t-test by testing whether the mean value for the set  is bigger than 0. We used cm(l), the median of {ci(l)}i=1,…,22, as the summary p-spectrum for the target population. We could also use the mean value to construct cm(l), but it would lead to more false admixture signals. Based on the summary p-spectrum, l with positive values of cm(l) were gathered as the candidate admixture time points, and then they were clustered into groups as different waves of admixture, Sg(k) for the kth admixture wave. Once these time points were grouped, the mean and variance of the time for that wave of admixture could be calculated by
is bigger than 0. We used cm(l), the median of {ci(l)}i=1,…,22, as the summary p-spectrum for the target population. We could also use the mean value to construct cm(l), but it would lead to more false admixture signals. Based on the summary p-spectrum, l with positive values of cm(l) were gathered as the candidate admixture time points, and then they were clustered into groups as different waves of admixture, Sg(k) for the kth admixture wave. Once these time points were grouped, the mean and variance of the time for that wave of admixture could be calculated by

Meanwhile, we used the minimum p-value on each time point in that group to measure the significance for each admixture wave. In this way, we could date multiple-wave admixture and measure the significance of each admixture wave. This algorithm was implemented in the method iMAAPs and available from http://www.picb.ac.cn/PGG/resource.php.
Results
Dating multiple-wave admixture with weighted LD statistics
There are two main difficulties for dating admixture in empirical analysis: reference population selection, and SLD reduction. To deal with these, Pickrell et al. (2014) claimed that different pairs of reference populations often have different relative values but always have the same sign of the coefficient of (1−d)l so that they can traverse all pairs of reference populations to test for the presence of possible admixture and estimate the time of each admixture wave. They used the LD whose pairwise genetic distance was longer than 0.5 cM, which was supposed to reduce the effect of SLD. Meanwhile, they also claimed that their algorithm (ALDER) was not very powerful in detecting multiple admixtures (Pickrell et al., 2014). Under our framework of weighted LD (Equation (2)), we confirmed that c(l) is an admixture determined constant and independent of the selection of reference populations; we pointed out that both the ai(d),i=1,2 and F(d) have the potential to affect the p-spectrum of a0(d), which directly affects the estimation of the coefficient of (1−d)l; we also noticed that the effect of SLD reduction with starting distance was not evaluated in the work by Pickrell et al. (2014), which may be the reason why their method is not powerful in detecting multiple admixtures. To verify our conjecture, we constructed a summary p-spectrum for weighted LD decay curves on a simulated admixed population with different pairs of reference populations.
A 100-generation-old admixed population was generated under the HI model, with YRI and CEU as source populations of admixture proportion 50:50%. A total of 55 pairs of HapMap populations (YRI, LWK, MKK, ASW, CEU, TSI, MXL, CHB, CHD, GIH and JPT) were used as references to calculate weighted LD a0(d) for further p-spectrum construction. In summary, p-spectrum with fully weighted LD decay (Figure 3a) for nearly all pairs of reference populations arose three main peaks around 100, 180, and 1250 generations. In the p-spectrum with weighted LD decay beginning at 0.5 cM (Figure 3b), the peaks around 180 and 1250 generations disappeared, but a new peak around 120 generations appeared, which was probably the remaining SLD and it may bias the time estimation of admixture. The remaining SLD should be the reason why ALDER did not work well for multiple waves of admixture (Pickrell et al., 2014). Meanwhile, we also observed that weighted LD decay with pairs of reference populations close to the true source populations had similar p-spectrums (Figure 3), which indicated that we could use populations not exact but similar to the source populations as references to construct the p-spectrum. This observation also supported the idea that using proper reference populations could increase the accuracy of ALDER in resolving weighted LD decay.
Summary p-spectrum for  with all pairs of populations from HapMap. Spectrum with true source populations (CEU and YRI) is in red lines; spectrum with selected pairs of reference populations (CEU, LWK; CEU, MKK; TSI, LWK; TSI, YRI; and TSI, MKK) are in black lines; spectrums with other pairs of reference populations are in gray lines. (a) Summary p-spectrum for full LD decay. (b) Summary spectrum for LD decay with a starting distance of 0.5 cM.
with all pairs of populations from HapMap. Spectrum with true source populations (CEU and YRI) is in red lines; spectrum with selected pairs of reference populations (CEU, LWK; CEU, MKK; TSI, LWK; TSI, YRI; and TSI, MKK) are in black lines; spectrums with other pairs of reference populations are in gray lines. (a) Summary p-spectrum for full LD decay. (b) Summary spectrum for LD decay with a starting distance of 0.5 cM.
Evaluation of iMAAPs
In our p-spectrum-based method iMAAPs, we used reference populations to estimate SLD and F(d), and separated their effect from the weighted LD of the admixed population. Thus, we could directly estimate the parameter c(l) and the number of admixture waves. A workable method in empirical admixture analysis should be robust to the proxy source populations. We have observed the robustness of the p-spectrum to different pairs of reference populations, and thus will evaluate the performance of iMAAPs to different reference pairs. Here, with the simulated 100-generation-old African European admixed population, generated by YRI and CEU, we showed that iMAAPs is very robust with African European pairs (YRI, CEU; LWK, CEU; MKK, CEU; LWK, TSI; YRI, TSI; MKK and TSI) as reference populations to infer admixture time (Supplementary Table S4).
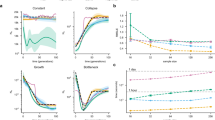
We also tested our method under various admixture models. iMAAPs were able to reconstruct the history of the admixture population well. For the one-wave and TW admixture models, iMAAPs gave times close to the true admixture; for the continuous migration models, it was able to place most of the signals in a particular migration time window (Figure 4 and Supplementary Figures S5–S13).
Performance of iMAAPs under various admixture models. The black vertical dashed lines represent the true simulated admixture time and gray areas represent the time window for continuous admixture. The summary p-spectrum of  for each simulated admixed population is plotted in heat color and the estimated admixture times are plotted in blue; points for the mean values and lines for the ranges of 3 s.d.
for each simulated admixed population is plotted in heat color and the estimated admixture times are plotted in blue; points for the mean values and lines for the ranges of 3 s.d.
Empirical analysis
This method was first applied to a few well-known admixed populations from available public databases: HGDP (Rosenberg et al., 2002) and HapMap Project phase III (The International HapMap Consortium, 2010). Our method is currently designed under the framework of two-way admixture and source populations or the populations similar to which are required in empirical analysis. Besides, two principles should be considered for interpretation:
(1) Existence of estimations for longer than 500 generations indicates that the SLD has not been well removed and thus some of the admixture signals, especially ancient signals, are probably generated by the SLD instead of the admixture.
(2) Existence of estimations close to generation 1, usually refers to 0 to 2, is always considered the result of the population substructure, not the admixture.
Based on these principles, we first analyzed three well-known admixed populations: African American (57 ASW individuals from HapMap), Mexican (86 MXL individuals from HapMap), and Uyghur (10 Uyghur individuals from HGDP). We also used ALDER to analyze these admixed populations (Supplementary Table S5). For each admixed population, we conducted three rounds of estimations. In the first round, we used all the populations in the full data set as the references to infer the admixture; in the second round, we used population pairs with the highest amplitude for each admixture wave in the first round as the reference populations to re-run ALDER; in the last round, we selected populations according to the admixture pattern based on the population inference in the first round. That is to say, if CEU and YRI were inferred as the best pair of populations to explain the admixture, then we selected all populations that could represent European and African ancestries as reference populations in the third run of ALDER. We believed this would increase estimation accuracy.
In our analysis with iMAAPs, reference populations were selected based on the results of ALDER’s inference on each admixture wave. CEU (n=113) and YRI (n=147) were chosen as the ancestral populations of ASW. YRI (n=147), TSI (n=102) and American Indian (7 Colombians, 14 Karitiana, 21 Maya, 14 Pimas and 8 Suruis from HGDP) were used as the ancestral populations of MXL. Basque (n=24), Sardinian (n=28), Japanese (n=28), Han (n=34) and French (n=28) were used as the ancestral populations of Uyghur.
The estimation of admixture time for ASW was 5.4±0.4 generations ago and the SLD was well reduced with YRI and CEU as reference populations (Supplementary Table S7 and Figure 5). Meanwhile, ALDER gave us two different results: 12.0±4.4 generations with all populations in HapMap as references; 6.3±3.3 and 77.0±65.9 generations with selected reference populations from HapMap (Supplementary Table S5). In this ALDER estimation, generation 6.3 was very close to our result, which can be interpreted as the admixture time of the population ASW. Furthermore, the result of generation 77.0 reflected the failure of SLD reduction with a starting distance of 0.5 cM.
Dating empirical admixture with iMAAPs. The summary p-spectrum of  for each simulated admixed population, calculated with the median, is plotted in heat color and the estimated admixture times are plotted in blue; points for the mean values and lines for the ranges of 3 s.d. The target admixed populations and reference populations are listed on the right.
for each simulated admixed population, calculated with the median, is plotted in heat color and the estimated admixture times are plotted in blue; points for the mean values and lines for the ranges of 3 s.d. The target admixed populations and reference populations are listed on the right.
MXL seemed to have experienced its main admixture 7.0±0.2 generations ago with TSI and American Indians as reference populations, and 8.2±0.4 generations ago with YRI and American Indians as reference populations (Supplementary Table S7). More admixture time points would be detected using the mean to construct a summary p-spectrum (Supplementary Table S6), which must be confirmed by further studies.
The Uyghur population has been reported to have a much longer admixture history than ASW and MXL (Xu and Jin, 2008; Xu et al., 2008; Qin et al., 2015). In the present study, admixture was found 33.3±0.5 generations ago with Han and French as reference populations. This admixture event has also been detected with Basque, Han, Sardinian and Japanese as reference populations, suggesting that the major admixture occurred around 825 years ago.
Loh et al. (2013) speculated that there could have been multiple waves of admixture in the history of MKK. Here, both our method and ALDER detected at least two waves of admixture (Supplementary Tables S5–S7 and Figure 5). We used reference pair of YRI and CEU, and pair of YRI and TSI to resolve the admixture of the MKK. With YRI and CEU as references, admixtures around 16.2 and 68.4 generations ago were detected; with YRI and TSI as references, admixtures around 17.9 and 70.3 generations ago were detected. However, both detections have estimations longer than 500 generations ago, indicating that we need to consider SLD when interpreting the time of admixture.
Discussion
Available methods based on ALD for admixture dating have shown their robustness in dealing with genotype data (Loh et al., 2013) and complicated admixture history (Pickrell et al., 2014). However, the effect of SLD in these methods has not been well defined and efficiently reduced, which may bias the estimation. In this study, we defined the SLD in the weighted LD statistic of the target admixed population under the two-way admixture model and introduced a p-spectrum to study the weighted LD decay for both the admixed population and source populations. We found that SLD tends to have higher degree of p-spectrum than the LD introduced by recent admixture, and that using the strategy relying on starting distance can partly reduce the effect of SLD (Figure 3). We also found that SLD can be well compensated by the LD of source populations (Supplementary Figures S5–S13), which motivated us to develop a new method, iMAAPs, to infer multiple-wave admixture. In this method, we used reference populations to estimate the SLD, and then reduced its effect and gave the accurate estimation.
We applied both iMAAPs and ALDER to date several well-known admixed populations. When running ALDER, we conducted estimation in three rounds with different sets of reference populations. Based on ALDER’s assumption of the effect of different pairs of reference populations on weighted LD, time estimations in these three rounds should be very close to each other. However, for the population of ASW, the result of first round was different from the results of other rounds, indicating the potential risk for using ALDER with global references. In the estimations of the second and third round, we found signals around 6 and 70 generations ago. The signal around 70 generations was a false admixture signal caused by the remaining SLD, because iMAAPs only detected the signal close to the 6 and the SLD was well reduced. Furthermore, we found that signals were close to 1 in all these three rounds of estimation, which, as we suggested, should be interpreted as population substructure instead of admixture time. We also ran iMAAPs in populations of MXL, Uyghur and MKK. MXL seemed to experience admixture seven generations ago. For Uyghur, the major admixture occurred about 33 generations ago. This date was a little earlier than the date determined by ALDER, in which the most significant admixture was around 40 generations ago. This difference was probably caused by the remaining SLD with the starting distance strategy. Both ALDER and iMAAPs confirmed that the population MKK experienced multiple waves of admixture, which had been predicted in the work of Loh et al. (2013).
One of the fundamental ideas in this algorithm is to take advantage of proper representative reference populations to reduce the effect of SLD. As using improper reference populations may bias the final estimation, it is crucial to carefully select reference populations in empirical analysis. Another issue that should be noticed is that this algorithm might give multiple pulses of signals even though the true population history was a long lasting continuous admixture (Supplementary Figures S5-S8). Nevertheless, this work built the framework of weighted LD under two-way admixture and provided an alternative to reduce the effect of SLD in the estimation of admixture time.
References
Chakraborty R, Weiss KM . (1988). Admixture as a tool for finding linked genes and detecting that difference from allelic association between loci. Proc Natl Acad Sci USA 85: 9119–9123.
Hill WG, Robertson A . (1966). The effect of linkage on limits to artificial selection. Genet Res (Camb) 8: 269–294.
Jin W, Wang S, Wang H, Jin L, Xu S . (2012). Exploring population admixture dynamics via empirical and simulated genome-wide distribution of ancestral chromosomal segments. Am J Hum Genet 91: 849–862.
Li N, Stephens M . (2003). modeling linkage disequilibrium and identifying recombination hotspots using single-nucleotide polymorphism data. Genetics 165: 2213–2233.
Loh PR, Lipson M, Patterson N, Moorjani P, Pickrell JK, Reich D et al. (2013). Inferring admixture histories of human populations using linkage disequilibrium. Genetics 193: 1233–1254.
McKeigue PM . (2005). Prospects for admixture mapping of complex traits. Am J Hum Genet 76: 1–7.
Moorjani P, Patterson N, Hirschhorn JN, Keinan A, Hao L, Atzmon G et al. (2011). The history of African gene flow into Southern Europeans, Levantines, and Jews. PLoS Genet 7: e1001373.
Nei M, Li WH . (1973). Linkage disequilibrium in subdivided populations. Genetics 75: 213–219.
Patterson N, Moorjani P, Luo Y, Mallick S, Rohland N, Zhan Y et al. (2012). Ancient admixture in human history. Genetics 192: 1065–1093.
Pfaff CL, Parra EJ, Bonilla C, Hiester K, McKeigue PM, Kamboh MI et al. (2001). Population structure in admixed populations: effect of admixture dynamics on the pattern of linkage disequilibrium. Am J Hum Genet 68: 198–207.
Pickrell JK, Patterson N, Loh P-R, Lipson M, Berger B, Stoneking M et al. (2014). Ancient west Eurasian ancestry in southern and eastern Africa. Proc Natl Acad Sci USA 111: 2632–2637.
Price AL, Tandon A, Patterson N, Barnes KC, Rafaels N, Ruczinski I et al. (2009). Sensitive detection of chromosomal segments of distinct ancestry in admixed populations. PLoS Genet 5: e1000519.
Qin P, Zhou Y, Lou H, Lu D, Yang X, Wang Y et al. (2015). Quantitating and dating recent gene flow between European and East Asian populations. Sci Rep 5: 9500.
Reich D, Patterson N . (2005). Will admixture mapping work to find disease genes? Philos Trans R Soc Lond B Biol Sci 360: 1605–1607.
Rosenberg N a, Pritchard JK, Weber JL, Cann HM, Kidd KK, Zhivotovsky LA et al. (2002). Genetic structure of human populations. Science 298: 2381–2385.
Smith MW, O’Brien SJ . (2005). Mapping by admixture linkage disequilibrium: advances, limitations and guidelines. Nat Rev 6: 623–632.
The International HapMap Consortium. (2010). Integrating common and rare genetic variation in diverse human populations. Nature 467: 52–58.
Wigginton JE, Cutler DJ, Abecasis R . (2005). A note on exact tests of Hardy-Weinberg equilibrium. Am J Hum Genet 76: 887–893.
Xu S, Huang W, Qian J, Jin L . (2008). Analysis of genomic admixture in Uyghur and its implication in mapping strategy. Am J Hum Genet 82: 883–894.
Xu S, Jin L . (2008). A genome-wide analysis of admixture in Uyghurs and a high-density admixture map for disease-gene discovery. Am J Hum Genet 83: 322–336.
Zhou Y, Liu X, Xu S . (2017). Dissecting admixture linkage disequilibrium under a general model of population admixture, under review.
Acknowledgements
We are grateful to Dr Li Jin, Dr Yungang He, Dr Xiaoming Liu, Dr Minxian Wang, Dr Joshua Schraiber and one anonymous reviewer for their valuable comments. This work was supported by the Strategic Priority Research Program (XDB13040100) and Key Research Program of Frontier Sciences (QYZDJ-SSW-SYS009) of the Chinese Academy of Sciences (CAS), the National Natural Science Foundation of China (NSFC) grant (91331204), the National Science Fund for Distinguished Young Scholars (31525014), the Program of Shanghai Academic Research Leader (16XD1404700) and the National Key Research and Development Program (2016YFC0906403). SX is the Max-Planck Independent Research Group Leader and member of CAS Youth Innovation Promotion Association. SX also gratefully acknowledges the support of the National Program for Top-notch Young Innovative Talents of The ‘Wanren Jihua’ Project. Funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript. We thank LetPub (www.letpub.com) for its linguistic assistance during the preparation of this manuscript.
Author contributions
SX conceived and designed the study. YZ developed the methods with contribution from YY. YZ and KY developed computer tools and analyzed the data. SX, YZ and KY interpreted the data and wrote the paper, with contribution from YY, XN, PX and EX.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies this paper on Heredity website
Supplementary information
Appendix
Appendix
Construction of the P-spectrum for Decay Curve
The decay curve of z(d) can be fitted by a numerical routine known as the proximal gradient (Beck and Teboulle, 2009) to minimize the objective function to fit out the parameter c:

where  is a vector of z(d) with different d values.
is a vector of z(d) with different d values.  is the coefficient of the polynomial functions and all entries in vector c are non-negative. The
is the coefficient of the polynomial functions and all entries in vector c are non-negative. The  entry of the matrix As × (n+1) is Aij=(1−di)j, where
entry of the matrix As × (n+1) is Aij=(1−di)j, where  . In empirical analysis, the value of j ranges from 0 to thousands, say 0 to 2000 in our analysis, which would lead matrix Aij to be too large to compute efficiently. Thus, to increase computation efficiency, we used the set Sg, a subset containing time candidates sampled from the range of 0 to 2000, as the j’s value set. With this method, it was possible to find vector c and fitting curve Ac. Next, we constructed the p-spectrum from de-noising vector c.
. In empirical analysis, the value of j ranges from 0 to thousands, say 0 to 2000 in our analysis, which would lead matrix Aij to be too large to compute efficiently. Thus, to increase computation efficiency, we used the set Sg, a subset containing time candidates sampled from the range of 0 to 2000, as the j’s value set. With this method, it was possible to find vector c and fitting curve Ac. Next, we constructed the p-spectrum from de-noising vector c.
For vector c, each entry represents the magnitude of the signal and a large magnitude indicates that the signal contains information rather than noise. Based on this idea, only the top c(l) values that together composed 99.9% of Z were retained for p-spectrum construction. This means that we must determine the smallest set Ω⊆Sg subject to the condition

and then let c(l),  , be zero. In this way, we constructed the p-spectrum for the decay curve Z.
, be zero. In this way, we constructed the p-spectrum for the decay curve Z.
Clustering candidate time points
Suppose we have the candidate time points generated from summary p-spectrum as an increasing series {ti}i=1,2,…,I, then we say ti and ti+1 belong to the same cluster if only one of these two conditions stands:

or

Reference
Beck A, Teboulle M (2009). A Fast iterative shrinkage-thresholding algorithm for linear inverse problems. {SIAM} J Imaging Sci 2: 183–202.
Rights and permissions
About this article
Cite this article
Zhou, Y., Yuan, K., Yu, Y. et al. Inference of multiple-wave population admixture by modeling decay of linkage disequilibrium with polynomial functions. Heredity 118, 503–510 (2017). https://doi.org/10.1038/hdy.2017.5
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2017.5
This article is cited by
-
MultiWaver 2.0: modeling discrete and continuous gene flow to reconstruct complex population admixtures
European Journal of Human Genetics (2019)
-
Models, methods and tools for ancestry inference and admixture analysis
Quantitative Biology (2017)