Abstract
Recently, an effective Bayesian shrinkage estimation method has been proposed for mapping QTL in inbred line crosses. However, with regard to outbred populations, such as half-sib populations with maternal information unavailable, it is not straightforward to utilize such a shrinkage estimation for QTL mapping. The reasons are: (1) the linkage phase of markers in the outbred population is usually unknown; and (2) only paternal genotypes can be used for inferring QTL genotypes of offspring. In this article, a novel Bayesian shrinkage method was proposed for mapping QTL under the half-sib design using a mixed model. A simulation study clearly demonstrated that the proposed method was powerful for detecting multiple QTL. In addition, we applied the proposed method to map QTL for economic traits in the Chinese dairy cattle population. Two or more novel QTL harbored in the chromosomal region were detected for each trait of interest, whereas only one QTL was found using traditional maximum likelihood analyses in our earlier studies. This further validated that our shrinkage estimation method could perform well in empirical data analyses and had practical significance in the field of linkage studies for outbred populations.
Similar content being viewed by others
Introduction
Many economically important traits and disease-resistant traits in animals are controlled by multiple genes, and the locations of these genes on the chromosomes are called quantitative-trait loci (QTL). With the development of molecular technology, these QTL can be localized and eventually the actual genes within these QTL can be cloned. Outbred populations are very ubiquitous in domestic animals; moreover, paternal half-sib families are quite often used for mapping QTL in such populations, in which the phenotypes of the offspring and genotypes of paternal parents and offspring are used in the analysis.
Numerous QTL mapping methods for half-sib design have been proposed. Georges et al. (1995) developed a maximum likelihood method for single-family analysis and implemented it to map QTL of milk production traits in the US Holstein population. The regression method of interval mapping proposed by Knott et al. (1996) is a common method used to map QTL in half-sib families, particularly in dairy cattle (Fulker and Cardon, 1994; Spelman et al., 1996; Zhang et al., 1998; Velmala et al., 1999; Heyen et al., 1999; de Koning et al., 2001; Nadesalingam et al., 2001; Ron et al., 2001; Plante et al., 2001; Freyer et al., 2002; Rodriguez-Zas et al., 2002; Viitala et al., 2003). Grignola et al. (1996a, 1996b) proposed a restricted maximum likelihood method and it was used by Zhang et al. (1998); Freyer et al. (2002) and Liu et al. (2004) to map QTL in Holstein populations in America, Germany and Canada, respectively. All these methods are based on models of a single QTL, and are hard to be extended to handle multiple QTL. If the trait is controlled by multiple QTL, the single QTL model-based estimation of QTL position and effect may be biased because of the presence of multiple linked QTL on the same chromosome. In the situation where the effects of two-linked QTL are in the opposite direction, the QTL effects may cancel out each other and none of them can be detected. On the other hand, if their effects are in the same direction, a ‘ghost’ QTL may be mapped between the two real QTLs. To overcome the above problems, Jansen (1993) and Zeng (1994) independently proposed a composite interval mapping (CIM) method. The major problem in the CIM method is that it is difficult to determine the number of markers as cofactors, because too many nuisance markers will decrease the detection power and too few markers cannot control the genetic background. Kao et al. (1999) proposed a multiple interval mapping (MIM) approach that took multiple QTL simultaneously into consideration. However, MIM only detects epistasis between main-effect QTL and cannot identify QTL with small effects. Furthermore, the CIM and MIM methods were originally developed for QTL mapping in inbred populations rather than outbred populations.
Recently, Bayesian approach has been developed for mapping multiple QTLs, in which the number of QTL is considered as a parameter to be estimated. Within the Bayesian multiple QTL mapping framework, several algorithms have been proposed, such as the reversible jump Markov chain Monte Carlo (RJMCMC) (Sillanpää and Arjas, 1998; Stephens and Fisch, 1998), the stochastic search variable selection (SSVS) (Yi, 2004) and the Bayesian shrinkage method (Xu, 2003; Wang et al., 2005; Xu, 2007). The key feature of the RJMCMC algorithm is that the number of QTL is treated as an unknown model parameter and is estimated through Bayesian model selection. A shortcoming of RJMCMC is that the Markov chain may converge slowly and have a poor mixing character due to model dimension changing with the number of QTL (Satagopan and Yandell, 1996; Yi and Xu, 2002; Liu et al., 2007; Yi et al., 2007). Compared with RJMCMC, SSVS and the Bayesian shrinkage estimation can overcome this issue to some extent. In SSVS, a previous mixture is adopted to explicitly make a probabilistic statement about the inclusion of a QTL, and the markers with significant effects can be identified as those with higher posterior probabilities involved in the model (Yi, 2004). In the Bayesian shrinkage analysis, each marker or marker interval is assumed to be associated with one QTL. If a marker or a marker interval is not associated with any QTL, the corresponding QTL effect will be shrunk toward zero. Accordingly, both SSVS and the Bayesian shrinkage estimation can largely avoid the problems existing in RJMCMC (Xu et al., 2005; Yang et al., 2006, 2007). A specific advantage of the Bayesian shrinkage estimation is that it can handle the situation where the number of unknown parameters is more than the number of observations. Recently, the Bayesian shrinkage estimation has been proposed to map multiple QTLs and epistatic QTLs in inbred line crosses (Xu and Jia, 2007). Compared with the Bayesian shrinkage method, SSVS is not optimal for QTL parameter estimation because the previous variance of QTL is ascertained arbitrarily to some extent (Wang et al., 2005). The results from simulations and real experiments also clearly showed that the Bayesian shrinkage estimation outperforms SSVS (Wang et al., 2005), that is, it could expedite the convergence process of the Markov chain and decrease the chance of missing QTL.
Although the Bayesian shrinkage method is effective for mapping QTL in inbred line crosses, it is not straightforward to be extended for use in outbred populations. The difficulty of mapping QTL in outbred populations is that the marker linkage phases are usually unknown and need to be inferred using the marker information of the offspring and their parents, and then the probability of each QTL genotype can be estimated using the reconstructed linkage phases. Motivated by the obvious advantages of the Bayesian shrinkage mapping over other existing methods, we proposed a novel Bayesian shrinkage method to map multiple QTLs in half-sib families in this study. Meanwhile, the performance of the proposed method was demonstrated using simulated data and a real data set of dairy cattle.
Methods
The multiple QTL model
A specific paternal half-sib population was taken into consideration in this study, where the maternal information on phenotypes and genotypes was not available. According to the feature of half-sib population, the phenotypic observation of each individual is modeled as:

where yi is the phenotypic observation of individual i, μ is the overall mean, gk is the substitution effect of QTL k, q is the number of QTL, xik is an indicator variable of QTL k with value 1 or −1 corresponding to the two paternal QTL alleles Q and q carried by individual i, uj is the residual polygenic effect following the distribution N(0, Aσ2u) with A being additive genetic relationship matrix and σu2 the variance of the polygenic effect, and ei is the residual error following the distribution N(0, I σe2).
The posterior and previous distributions
In multiple QTL mapping, the estimation of the number of QTL is an important aspect. In the Bayesian shrinkage analysis, the number of QTL can be estimated through shrinking the predefined maximum number of QTL to the true number of QTL in a natural way instead of estimating it directly. The parameters of direct interest in model (1) are the QTL position vector λ=(λ1λ2…λq)′ and the QTL effect vector g = (g1 g2...gq. The whole set of unknown parameters in model (1) is θ′ = (μ, λ ′, g ′, σ21,σ22,...,σ2q, X, u′, σ2u, σ2e), here X is the vector of the indicator variables of QTL genotypes of all individuals, and u is the vector of the residual polygenic effects of all individuals. The posterior distribution of θ given the observed data D is

where p(D∣θ) is the likelihood of the data given θ, and p(θ∣Θ) is the previous density of θ with hyper parameter vector Θ.
In this study, we assume that the overall mean μ has a uniform previous distribution, the QTL effect gk (k=1, 2,…, q) has a normal previous distribution gk ∼ N(0, σ2k) with p(σ2u) ∝ 1/σ2k, the residual polygenic effects have a multivariate normal previous distribution u ∼ N(0, A σ2u) with p(σ2u) ∝ 1/σ2u, the residual error effects follow a normal previous distribution e ∼ N(0, I σ2e) with p(σ2e) ∝ 1/σ2e, and the previous probability of the position of QTL k is p(λk) ∝ 1/dk with dk being the length of the marker interval that QTL k falls in.
For QTL mapping, there are two sources of observed data, the phenotypic data y and the marker data M, which are conditionally independent. Therefore, the likelihood in (2) can be written as

Assuming normal distribution of y, the likelihood of y given θ is

where N is the total number of individuals in all of the half-sib families.

where Xi• = (xi1 xi2…xiq)′. Both p(M, Xi•∣λ) and p(Xi•∣λ) can be derived from a Markov model under the assumption of no segregation interference. For ease of illustration, three neighboring markers are used as an example to explain the specific expression of p(M, Xi•∣λ) and p(Xi•∣λ) as below:

where p (mi1)=1/2
The full conditional posterior distributions and MCMC sampling
To implement the Bayesian estimation via the MCMC algorithm, the full conditional posterior distributions of each parameter need to be derived from the joint posterior density by fixing all other parameters. Based on these full conditional posterior distributions, the MCMC can be implemented with the following steps:
-
1)
Initialize all variables with some legal values, which can be randomly assigned within their respective sample spaces;
-
2)
Update the overall mean μ using A (1);
-
3)
Update the QTL effects gk (k=1, 2…, q) using A (2);
-
4)
Update the residual polygenic effects ui (i=1, 2…, N) using A(3);
-
5)
Update the QTL variances σk2 (k=1, 2…, q) using A (4);
-
6)
Update the residual polygenic variance σu2 using A (5);
-
7)
Update the residual error variance σe2 using A (6);
-
8)
Impute the genotypes of missing markers using the approach originally proposed by Wang et al. (2005). The genotypes of missing markers are sampled sequentially along the genome individual by individual in each family because the missing marker information of different individuals is independent.
-
9)
Update the QTL genotype indicator variables xik (i=1, 2…, N) and QTL positions λk (k=1, 2…, q);
-
10)
Repeat steps (2)–(9) until the Markov chain reaches a desirable length. The details of A (1)–A (6) are given in the Appendix.
The detailed algorithm for step (9) is described below. To sample the QTL genotypes, the marker linkage phases should be available. In the outbred population, the marker linkage phases are usually unknown, so they have to be inferred according to the marker information of the offspring and their parents. Sampling QTL genotype posterior samples can be achieved according to the QTL Identity-By-Descent (IBD) probabilities among two generations of the half-sib families. The approach of inferring QTL IBD probability was presented by Haley and Knott (1992) and Knott et al. (1996). Three steps are involved in this approach: (1) infer the marker–marker linkage phases within each family according to marker genotypes of all members in the family; (2) infer the marker–QTL linkage phases given the position of the putative QTL and the inferred marker–marker linkage phases and (3) calculate the QTL genotypic transmission probabilities in the two generations of half-sib families using the inferred marker-QTL linkage phases. The conditional probabilities of QTL genotypes depend on the alleles inherited at the two nearest informative markers flanking the QTL and the recombination rates between the markers and the QTL. As the conditional probabilities sum to unity, only the probability for the first sire gamete need to be calculated. For any position, the flanking markers used to calculate these probabilities will vary from sire to sire and from progeny to progeny within a sire. It should be noted that for some individuals, a chosen position may be outside the last informative marker in the linkage group. In such cases the conditional probabilities depend on the single nearest informative marker. For an extreme situation where all markers in a linkage group are non-informative, the probabilities are set to be 0.5 for both gametes at all positions within the linkage group.
In step (9), QTL genotypes and QTL positions are updated jointly using information of all families simultaneously. As the genotype of QTL depends on the QTL position, so {λk, X•k} are sampled jointly using the Metropolis–Hastings algorithm. Each locus is sampled from a variable interval (Wang et al., 2005; Zhang and Xu, 2005) between the positions of the adjacent QTL. First a novel position is proposed, and then the QTL genotypes are sampled with the conditional probabilities of the QTL genotypes calculated at this position. The proposed QTL position is accepted with probability of min(1,α) (see also Wang et al., 2005; Zhang and Xu, 2005) with

where the superscripts (*) and (0) denote the new and the old sampled values, respectively.
The first term in (5) is the posterior ratio of the new to the old positions and can be calculated as

where P(λ(*)k) and P(λ(0)k) are the previous probabilities of the new and the old position, respectively, and P(λ(*)k)/P(λ(0)k) = 1 under uniform previous distribution.
The second term in (5) q(λ(0)k)/q(λ(*)k) = 1 is the proposal ratio of the new to old position and the third term  is the proposal ratio of the QTL genotypes corresponding to the new and old positions.
is the proposal ratio of the QTL genotypes corresponding to the new and old positions.
In the above MCMC process, the shrinkage for QTL parameters is achieved through step (5). Specifically, in the absence of a QTL, the estimate of the posterior mean for QTL effect inclines to zero so that the sampled observations of QTL effect are close to zero, whereas a ‘true’ QTL with large effects are estimated with virtually no shrinkage. To achieve this objective under the framework of Bayesian shrinkage estimation, we allow each QTL effect to have its own variance parameter with a specific previous distribution so that the variance can be estimated through the combined information of the observed data and its previous distribution under the framework of Bayesian estimation.
Post-MCMC analysis
In the Bayesian shrinkage analyses, QTL detection at a given position can be performed through the estimated QTL effect at that position weighted by the relative frequency of that position hit by the QTL (Wang et al., 2005). Although this method can give a clear signal for detecting QTL, it cannot provide a threshold level for declaring the significance of the existence of QTL. To overcome this limitation, a t-statistic is constructed for declaring the significance of the QTL, which is a special case of the Z-statistic that has been used in multivariate mapping of QTL (Yang and Xu, 2007). Specifically, we first divide the genome into m bins. For each bin, the t-statistic can be expressed as

where Nsam is the number of the poster samples, β(ξl) and s(ξl) are the average and s.d. of the QTL effect at position ξl, respectively. Under the null hypothesis of no existence of any QTL, t(ξl) follows a standard normal distribution. The critical value is 1.96 at the significant level of 0.05, and 2.58 at the significant level of 0.01 for declaring statistical significance at position ξl.
Application
Simulation study
A half-sib population with 20 sire families and 800 offspring in total was simulated here. A chromosomal region with the length of 250 cM was simulated, which was covered by 26 evenly placed markers (10 cM per marker interval), each with five alleles. With the assumption of linkage equilibrium among all loci and equal allele frequencies in the parental population, all parents’ genotypes at each marker locus were generated independently from the five alleles with equal probability. The genotypes of the offspring were generated based on Mendel's independent segregation rule and recombination rates (calculated using the Haldane's mapping function) between two adjacent loci according to the marker linkage phases in the parents.
The quantitative trait was assumed to be controlled by five QTL. The positions and effects of these QTL are given in Table 1. The overall mean μ was set to be 0. The residual polygenic variance and the residual error variance were set to be 0.1 and 0.2, respectively. The phenotypic values of the trait were simulated according to model (1).
In the MCMC analysis, the initial values of all unknown parameters were randomly assigned within the respective sample spaces. A single long chain with 55 000 cycles was generated. The burn-in period was set to be 5000 cycles, and one sample value was taken in every 20 cycles from the remaining 50 000 cycles. The total number of samples used for the post-MCMC analysis was 50 000/20=2500. The simulation experiment was replicated 10 times.
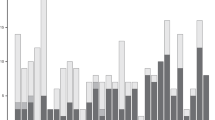
As the results of the 10 replicates were very similar to each other, results for only one replicate were presented here. Figures 1 and 2 show the profiles of the weighted QTL effects and the QTL intensities, respectively. Here, the definition of QTL intensity was similar to that of Sillanpää and Arjas (1998) and Yi and Xu (2000), that is, the frequency of hits by the QTL in a sufficient small interval (for example, 1 cM) around a particular position against that of the whole region covered by the markers. As expected, both profiles show five sharp peaks at the positions of the five simulated QTLs. The role of shrinkage is obvious: effects of intervals without QTL shrunk to zero and thus no variable selection was needed. The t-statistic profile (Figure 3) also shows significant signals at the positions of the simulated QTL. All of the five peaks of the t-statistic profile exceed the critical value (t=1.96) and no peaks elsewhere are higher than the critical value. The estimated QTL positions and effects (posterior means) from the 10 replicates are summarized in Table 1, showing that the Bayesian shrinkage analysis provides quite accurate estimates of QTL positions and effects. For further investigating the performance of the proposed method, we set the effects of the simulated QTL as zero, and ran the MCMC again. It turned out that the estimates of the QTL effects were close to zero (Figures 4 and 5), indicating that the Bayesian shrinkage estimation method did not tend to produce ‘pseudo QTL’.
For further validation of our proposed Bayesian shrinkage method, we compared our method with the common RJMCMC approach. Specifically, the RJMCMC method was employed to analyze the same simulated data as used for our method. In the RJMCMC process, the complete length of the MCMC chain was 205 000, the burn-in length was 5000 and the thinning interval was 20. The parameter estimates are summarized in Table 1 and the position estimates for the second simulated QTL from the 10 replicates by the two methods are also presented in Figure 6. These results show that there are no obvious differences in the estimates of QTL effects, their s.d. and QTL positions between the two methods. However, the s.d. of QTL position from our method are much smaller than that of the RJMCMC method, indicating that our method tends to give a more precise estimate for QTL position. In addition, although RJMCMC can detect similar QTL signals as those detected by our method, the estimate of the number of QTL (estimated as the posterior mode) given by RJMCMC is obviously biased. Specifically, the estimated number of QTL by RJMCMC from the 10 replicates all equal three, whereas the simulated ‘true’ number of QTL is five.
Real data analysis
The data came from a Chinese Hostein population with a daughter design and has been used by Chen et al. (2006) for mapping QTL for milk production traits using the regression method of interval mapping proposed by Knott et al. (1996). Briefly, there were 26 bulls and their 2260 daughters with phenotypes on five milk production traits (milk yield, fat yield, protein yield, fat percentage and protein percentage). The bulls and daughters were genotyped for 14 marker loci on chromosome 6 covering a total distance of 55.7 cM.
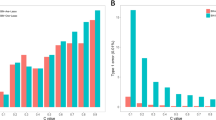
Separate MCMC analyses for the five traits were performed. In each MCMC process, Markov chain was run with 110 000 cycles, of which the first 10 000 cycles were discarded as burn-in. The estimates of the QTL positions and effects are given in Table 2. Figure 7 shows the t-statistic profile resulted from the analysis. There are two peaks for both milk yield and protein yield, one located at 55.7 cM between markers BM1329 and BMS2508 and the other at 9 cM close to BMS470. The peaks for fat yield and fat percentage are located at 18 cM between markers BMS1242 and BMS518 and 38 cM closed to marker RM028, respectively. In addition, there is another peak at 11 cM for fat percentage. The peaks for protein percentage are located at 7 cM close to marker BMS2508 and 50 cM close to BMS2460. All of these peaks exceed the significant level of P=0.01.
Discussion
The Bayesian shrinkage approach was originally developed for multiple QTL mapping for inbred line crosses (Xu, 2003; Wang et al., 2005) and has been proved effective and powerful. However, the inbred line cross is seldom used for QTL mapping in domestic animals, because it is extremely difficult to construct and maintain inbred lines for such populations. Furthermore, making a cross between outbred lines (breeds) is not feasible for some large animals such as dairy cattle because it is too time consuming and very expensive. We successfully extended the original Bayesian shrinkage method to map QTL in half-sib families. Both the simulation study and the real data analysis showed that the proposed method worked well and was powerful for detecting multiple QTL.
We compared the proposed Bayesian shrinkage method with the RJMCMC method through simulation with half-sib design. Besides the gain in precision of QTL position estimation, our method showed obvious advantage over RJMCMC in estimation of the number of QTL, the key parameter in multiple QTL mapping. Furthermore, we found that the RJMCMC method converged slower than our Bayesian shrinkage method and a longer Markov chain (200 000 iterations after burn-in period) was needed to the characteristics of low sampling efficiency in the RJMCMC. Wang et al. (2005) has also pointed out that RJMCMC for model selection was usually subject to poor mixing, that is, slow convergence, and demonstrated that the MCMC shrinkage analysis converged faster than RJMCMC via simulation with backcross design. Our findings herein were in accordance with those of Wang et al. (2005). Results from the simulation study clearly demonstrated that the Bayesian shrinkage approach had similar advantages in the half-sib design as in the backcross design over the RJMCMC approach.
Evidences for QTL on BTA6 affecting milk production traits have been reported by our earlier studies (Chen et al., 2006) and many other investigators (for example, Kuhn et al., 1999; Ron et al., 2001; Olsen et al., 2002; Freyer et al., 2003; Szyda et al., 2005). For further validating our method, we analyzed the same data set of Chen et al. (2006) using the proposed Bayesian shrinkage method. It is notable that the findings of Chen et al. (2006) were obtained based on a single-QTL model. On the contrary, our Bayesian shrinkage method was based on a multi-QTL model. In theory, the multi-QTL model should be superiors to the single-QTL model, since it is unreasonable and seldom true to assume that there exists only one QTL within the chromosomal region investigated. This aspect has been further confirmed via the real data analyses. Specifically, using the multi-QTL model-based Bayesian shrinkage method, we not only confirmed the QTL identified by Chen et al. (2006), but also identified some novel QTL. For example, for fat yield, in addition to the QTL found by Chen et al. (2006), two additional QTL at 18 cM and 38 cM, respectively, were identified. For these two additional QTL, one (at 8 cM) was previously reported by Kuhn et al. (1999), Szyda et al. (2005) and Freyer et al. (2003), the other (at 38 cM) was also detected by Szyda et al. (2005). Furthermore, a novel QTL affecting fat percentage beyond the findings of Chen et al. (2006) was identified at 18 cM, which was also reported by Ron et al. (2001) and Olsen et al. (2002). Comparisons among the findings in this study, by Chen et al. (2006), and by other investigators clearly demonstrate that our Bayesian multi-QTL model is more powerful and robust in QTL mapping than the single-QTL model. This also shows that our method proposed here has practical significance.
The approach presented in this paper is focused on half-sib families. It is straightforward to extend it to complex pedigrees by using a general approach to infer marker–QTL linkage phases. Such a general approach has been developed by many investigators (Vogl and Xu, 2002). In addition, although our proposed method is focused on linkage analyses, it can be naturally extended to fine mapping by using both linkage and linkage disequilibrium (LD) information if highly dense markers are available. Generally, combining linkage and LD information can significantly improve the precision of QTL mapping. For this situation, the marker–QTL linkage phases in the founders can be inferred through the existing well-developed methods (Meuwissen and Goddard, 2000, 2001; Meuwissen et al., 2002; Meuwissen and Goddard, 2004; Lee and Van der Werf, 2006). However, extra effort is needed to remove the limitation of the original assumption that the QTL are in linkage equilibrium in the parental generation.
The approach presented here can be easily modified to handle categorical traits with discrete phenotypes. Specifically, the categorical traits can be modeled using a threshold model with a normal underline liability. In Bayesian estimation framework, the liability ‘phenotypic value’ can be treated as unknown and can be sampled from their conditional posterior distribution (Yi and Xu, 2000; Hoti and Sillanpaa, 2006; Huang et al., 2007). The efficiencies of our approach for categorical traits will be investigated in our future endeavors.
Based on the framework of a mixed model, the proposed method can be also extended for multi-trait QTL mapping. Multi-trait analysis has been studied substantially both in inbred line crosses (Jiang and Zeng, 1995; Knott and Haley, 2000; Xu et al., 2005; Fang et al., 2008) and in outbred populations (Korol et al., 2001; Mangin et al., 1998; Eaves et al., 1996; Liu et al., 2007). The advantage of multi-trait analysis is that it is more powerful to detect QTL. Yang and Xu (2007) recently developed an approach of Bayesian shrinkage mapping of QTL for dynamic trait, and they also discussed how to extend their method to multivariate analysis. Their method can be incorporated into the Bayesian shrinkage approach for multi-trait QTL mapping with some modification in sampling QTL genotypes.
References
Chen HY, Zhang Q, Yin CC, Wang CK, Gong WJ, Mei G (2006). Detection of quantitative trait loci affecting milk production traits on bovine chromosome 6 in a Chinese Holstein population by the daughter design. J Dairy Sci 89: 782–790.
de Koning DJ, Schulman NF, Elo K, Moisio S, Kinos R, Vilkki J et al. (2001). Mapping of multiple quantitative trait loci by simple regression in half-sib designs. J Anim Sci 9: 616–622.
Eaves LJ, Neale MC, Maes H (1996). Multivariate multipoint linkage analysis of quantitative trait loci. Behav Genet 26: 519–525.
Freyer G, Kühn C, Weikard R, Zhang Q, Mayer M, Hoeschele I (2002). Multiple QTL on chromosome six in dairy cattle affecting yield and content traits. J Anim Breed Genet 119: 69–82.
Fang M, Jiang D, Pu LJ, Gao HJ, Hong PJ, Wang Y et al. (2008). Multitrait analysis of quantitative trait loci using Bayesian composite space approach. BMC Genetics 9: 48.
Freyer G, Sorensen P, Kuhn C, Weikard R, Hoeschele I (2003). Search for pleiotropic QTL on chromosome BTA6 affecting yield traits of milk production. J Dairy Sci 86: 999–1008.
Fulker DW, Cardon LR (1994). A sib-pair approach tointerval mapping of quantitative trait loci. Am J Hum Genetics 54: 1092–1103.
Georges M, Nielsen D, Mackinnon M, Mishra A, Okimoto R (1995). Mapping quantitative trait loci controlling milk production in dairy cattle by exploiting progeny testing. Genetics 139: 907–920.
Grignola FE, Hoeschele I, Tier B (1996a). Mapping quantitative trait loci in outcross populations via residual maximum likelihood. I. methodology. Genet Sel Evol 28: 479–490.
Grignola FE, Hoeschele I, Zhang Q, Thaller G (1996b). Mapping quantitative trait loci in outcross populations via residual maximum likelihood. II. A simulation study. Genet Sel Evol 28: 491–504.
Haley CS, Knott SA (1992). A simple regression method for mapping quantitative trait loci in line crosses using flanking markers. Heredity 69: 315–324.
Heyen DW, Weller JI, Ron M, Band M, Beever JE, Feldmesser E et al. (1999). A genome scan for QTL influencing milk production and health traits in dairy cattle. Physiol Genomics 1: 165–175.
Hoti F, Sillanpaa MJ (2006). Bayesian mapping of genotype × expression interactions in quantitative and qualitative traits. Heredity 97: 4–18.
Huang H, Eversley CD, Threadgill DW, Zou F (2007). Bayesian multiple quantitative traits loci mapping for complex traits using markers of the entire genome. Genetics 176: 2529–2540.
Jansen RC (1993). Interval mapping of multiple quantitative trait loci. Genetics 135: 205–211.
Jiang C, Zeng ZB (1995). Multiple trait analysis of genetic mapping for quantitative trait loci. Genetics 140: 1111–1127.
Kao CH, Zeng ZB, Teasdale RD (1999). Multiple interval mapping for quantitative trait loci. Genetics 152: 1203–1216.
Knott SA, Haley CS (2000). Multitrait least squares for quantitative trait loci detection. Genetics 156: 899–911.
Knott SA, Elsen JM, Haley CS (1996). Methods for multiple-marker mapping of quantitative trait loci in half-sib populations. Theor Appl Genet 93: 71–80.
Korol AB, Ronin YT, Itskovich AM, Peng J, Nevo E (2001). Enhanced efficiency of quantitative trait loci mapping analysis based on multivariate complexs of quantitative traits. Genetics 157: 1789–1803.
Kuhn C, Freyer G, Weikard R, Goldammer T, Schwerin M (1999). Detection of QTL for milk production traits in cattle by application of a specifically developed markermap of BTA6. Anim Genet 30: 333–340.
Lee SH, Van der Werf JHJ (2006). Simultaneous fine mapping of multiple closely linked quantitative trait loci using combined linkage disequilibrium and linkage with a general pedigree. Genetics 173: 2329–2337.
Liu J, Liu Y, Liu X, Deng HW (2007). Bayesian mapping of quantitative trait loci for multiple complex traits with the use of variance components. Am J Hum Genet 81: 304–320.
Liu Y, Jansen GB, Lin CY (2004). Quantitative trait loci mapping for dairy cattle production traits using a maximum likelihood method. J Dairy Sci 87: 491–500.
Mangin B, Thoquet P, Grimslev N (1998). Pleiotropic QTL analysis. Biometrics 54: 88–99.
Meuwissen THE, Goddard ME (2000). Fine mapping of quantitativetrait loci using linkage disequilibria with closely linked marker loci. Genetics 155: 421–430.
Meuwissen THE, Goddard ME (2001). Prediction of identityby descent probabilities from marker-haplotypes. Genet Sel Evol 33: 605–634.
Meuwissen THE, Goddard ME (2004). Mapping multiple QTL using linkage disequilibrium and linkage analysis information and multitrait data. Genet Sel Evol 36: 261–279.
Meuwissen THE, Karlsen A, Lien S, Olsaker I, Goddard ME (2002). Fine mapping of a quantitative trait locus for twinning rate using combined linkage and linkage disequilibrium mapping. Genetics 161: 373–379.
Nadesalingam J, Plante Y, Gibson JP (2001). Detection of QTL for milk production on chromosomes 1 and 6 of Holstein cattle. Mamm Genome 12: 27–31.
Olsen HG, Gomez-Raya L, Olsaker I, Klungland H, Svendsen M., Sabry A (2002). A genome scan for quantitative trait loci affecting milk production in Norwegian dairy cattle. J Dairy Sci 85: 3124–3130.
Plante Y, Gibson JP, Nadesalingam J, Mehrabanni-Yeganeh H, Lefebvre S, Vandervoort G et al. (2001). Detection of quantitative trait loci affecting milk production traits on 10 chromosomes in Holstein cattle. J Dairy Sci 84: 1516–1524.
Rodriguez-Zas SL, Southey BR, Heyen DW, Lewin HA (2002). Interval and composite interval mapping of somatic cell score, yield, and components of milk in dairy cattle. J Dairy Sci 85: 2681–2691.
Ron M, Klinger D, Feldmesser E, Seroussi E, Ezra E, Weller JI (2001). Multiple quantitative trait locus analysis of bovine chromosome 6 in the Israeli Holstein population by a daughter design. Genetics 159: 727–735.
Satagopan JM, Yandell BS (1996). Estimating the number of quantitative trait loci via Bayesian model determination. Special contributed paper session on genetic analysis of quantitative traits and complex disease. Biometric Section, Joint Statistical Meeting, Chicago.
Sillanpää MJ, Arjas E (1998). Bayesian mapping of multiple quantitative trait loci from incomplete inbred line cross data. Genetics 148: 1373–1388.
Spelman RJ, Coppieters W, Karim L, van Arendonk JA, Bovenhuis H (1996). Quantitative trait loci analysis for five milk production traits on chromosome six in the Dutch Holstein-Friesian population. Genetics 144: 1799–1808.
Stephens DA, Fisch RD (1998). Bayesian analysis of quantitative trait locus data using reversible jump Markov Chain Monte Carlo. Biometrics 54: 1334–1347.
Szyda J, Liu Z, Reinhardt F, Reents R (2005). Estimation of quantitative trait loci parameters for milk production traits in German Holstein dairy cattle population. J Dairy Sci 88: 356–367.
Velmala RJ, Vikki HJ, Elo KT, de Koning DJ, Maki-Tanila AV (1999). A search for quantitative trait loci for milk production traits on chromosome 6 in Finnish Ayrshire cattle. Anim Genet 30: 136–143.
Viitala SM, Schulman NF, de Koning DJ, Elo K, Kinos R, Virta A et al. (2003). Quantitative trait loci affecting milk production traits in Finnish Ayrshire dairy cattle. J Dairy Sci 86: 1828–1836.
Vogl C, Xu S (2002). QTL analysis in arbitrary pedigrees with incomplete marker information. Heredity 89: 339–345.
Wang H, Zhang YM, Li X, Masinde GL, Mohan S, Baylink DJ et al. (2005). Bayesian shrinkage estimation of quantitative trait loci parameters. Genetics 170: 465–480.
Xu CW, Li ZK, Xu S (2005). Joint mapping of quantitative trait loci for multiple binary characters. Genetics 169: 1045–1059.
Xu S (2003). Estimating polygenic effects using markers of the entire genome. Genetics 163: 789–801.
Xu S (2007). Derivation of the shrinkage estimates of quantitative trait locus effects. Genetics 177: 1255–1258.
Xu S, Jia Z (2007). Genomewide analysis of epistatic effects for quantitative traits in barley. Genetics 175: 1955–1963.
Yang R, Tian Q, Xu S (2006). Mapping QTL for longitudinal traits in line crosses. Genetics 173: 2339–2356.
Yang R, Xu S (2007). Bayesian shrinkage analysis of quantitative trait loci for dynamic traits. Genetics 176: 1169–1185.
Yi N, Xu S (2000). Bayesian mapping of quantitative trait loci for complex binary traits. Genetics 155: 1391–1403.
Yi N, Xu S (2002). Mapping quantitative trait loci with epistatic effects. Genet Res 79: 185–198.
Yi N, Shriner D, Banerjee S, Mehta T, Pomp D, Yandell BS (2007). An efficient Bayesian model selection approach for interacting quantitative trait loci models with Many Effects. Genetics 176: 1865–1877.
Yi N (2004). A unified Markov chain Monte Carlo framework for mapping multiple quantitative trait loci. Genetics 167: 967–975.
Zeng ZB (1994). Precision mapping of quantitative trait loci. Genetics 136: 1457–1468.
Zhang Q, Boichard D, Hoeschele I, Ernst C, Eggen A, Murkve B et al. (1998). Mapping quantitative trait loci for milk production and health of dairy cattle in a large outbred pedigree. Genetics 149: 1959–1973.
Zhang YM, Xu S (2005). Advanced statistical methods for detecting multiple quantitative trait loci. Recent Res Devel Genet Breeding 2: 1–23.
Acknowledgements
We are grateful to the two anonymous reviewers whose constructive comments and suggestions led to a substantial improvement of the original article. This study was supported by the National Basic Research Program of China (Grant No. 2006CB102104), the Key project of the National Natural Science Foundation of China (Grant No. 30430500) and the National Natural Science Foundation (Grant No. 30700565).
Author information
Authors and Affiliations
Corresponding author
Appendix: Full conditional posterior distributions used in the MCMC sampling
Appendix: Full conditional posterior distributions used in the MCMC sampling
1. Overall mean: normal distribution

2. QTL effects: normal distribution

3. Residual polygene effects: normal distribution

where,

5. Residual polygenic variance: inverted chi-square distribution

where SSA = u′A−1u.
6. Residual error variance: inverted χ2 distribution

where

7. QTL variance: inverted χ2 distribution

Rights and permissions
About this article
Cite this article
Gao, H., Fang, M., Liu, J. et al. Bayesian shrinkage mapping for multiple QTL in half-sib families. Heredity 103, 368–376 (2009). https://doi.org/10.1038/hdy.2009.71
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/hdy.2009.71