Abstract
It has repeatedly been suggested that the development of complex diseases can be elucidated by gene–gene interactions. Recently, we found that HTR2A, a member of the serotonin receptor family, is associated with rheumatoid arthritis (RA). This study was aimed to investigate the possibility of a gene–gene interaction between HTR2A and the major genetic risk factor for RA, HLA-DRB1 shared epitope (SE) alleles. We studied 4095 RA cases and 3223 controls from three different populations – from Sweden, the United States and the Netherlands – to test for interaction between the protective HTR2A haplotype and HLA-DRB1 SE alleles. Further, we analyzed mRNA and/or protein expression of HTR2A and HLA-DR in biopsy samples and in synovial fibroblasts from RA patients. The interaction was defined as departure from additivity of effects using attributable proportion due to interaction. First, we could demonstrate and further replicate an interaction between a protective haplotype in HTR2A and HLA-DRB1 SE alleles regarding risk of developing autoantibody-positive RA. Second, we could show that both genes are constitutively expressed in fibroblasts from synovial tissue of RA patients, and, by double immunofluorescence staining, we demonstrated that these two proteins are colocalized in these cells. In conclusion, our data demonstrate a statistical interaction between HTR2A and HLA-DRB1 SE alleles and colocalization of the product of these two genes in inflamed synovial tissue, which suggest a possible biological relationship between these two proteins. This finding may lead to the development of treatment based on enhancing the protective features of 5-HT2A in individuals with a certain HLA genotype.
Similar content being viewed by others
Introduction
Rheumatoid arthritis (RA (MIM 180300)) is a complex disease and much effort has been spent on finding risk genes for this disease. At present, >10 replicated RA susceptibility genes/loci, such as HLA-DRB1 shared epitope (SE) alleles,1 PTPN22,2 STAT4,3 TRAF1-C5,4, 5 CD406 and so on, have been discovered. However, except for MHC II genes, the odds ratios (ORs) are modest and estimations of the total genetic contributions to the disease indicate that additional genetic factors remain to be identified. We suggest that some of the genetic risk that remains to be identified resides in yet unidentified gene–gene7 and gene–environment interactions,8 and we have previously identified an example of gene–gene interaction between the two major susceptibility genes, HLA-DRB1 and PTPN22.7
Serotonin (5-hydroxytryptamine-5-HT) is a neurotransmitter that has an important role in brain homeostasis. It is surprising that only 1% of 5-HT in the human body is found in the CNS. The remaining 99% is found in other body compartments such as the gastrointestinal tract and immune tissues, wherein serotonin functions as a hormone regulating various physiological functions. Moreover, it is well documented that 5-HT has stimulatory and inhibitory effects on immune cells, including B, T, NK cells, monocytes and macrophages.9, 10 5-HT2A is one of the seven known serotonin receptors.11 HTR2A (MIM 182135) is localized on human chromosome 13q14–q21 and consists of three exons with five nonsynonymous and two synonymous variations and two introns with more than 200 known variations. In addition to neurons of the peripheral nervous system, 5-HT2A is highly expressed on platelets and fibroblasts, as well as in peripheral blood cells, and recently it was demonstrated that 5-HT2A is expressed on dendritic cells.12 Several biological and clinical facts serve as evidence for the connection between the function of 5-HT2A and immune response.11, 13 It has been demonstrated that the inhibition of production of TNF-α, an important proinflammatory cytokine, by 5-HT is mediated by 5-HT2A in PBMCs14 and that the activation of 5-HT2A suppresses TNF-α-induced inflammation in primary aortic smooth muscle cells.15
Previously we have demonstrated that genetic variations in HTR2A are in association with RA.16 However, as the strength of the association was moderate, we hypothesized that the strongest genetic risk factor for RA, HLA-DRB1 SE alleles, may modulate it and we can colocalize both products in rheumatoid tissue or in related cells.
The HLA-DRB1 SE alleles have consistently been shown to be associated with subtypes of RA characterized by the presence of autoantibody production (eg, rheumatoid factor (RF)-positive RA and/or anti-citrulline protein antibody (ACPA)-positive RA).17, 18 The ACPA-positive subtype of RA represents a major clinically defined phenotype for the disease and accounts for ∼60% of all RA patients. Several facts show that both genetic and environmental risk factors for ACPA-positive and ACPA-negative RA are different,17, 19 as well as the clinical course, histology and response to therapy.20, 21
The aim of this study was to use three different relatively large case–control studies to investigate the potential interactions between HTR2A and HLA-DRB1 SE alleles in proving risk for RA, and to estimate where biological interaction may take place.
Materials and methods
Study population
In this report, the following three study populations have been included: Swedish Epidemiological Investigation of Rheumatoid Arthritis (EIRA, 2158 cases and 1068 controls), North American Rheumatoid Arthritis Consortium (NARAC, 908 ACPA-positive cases and 1260 controls) and the Leiden Early Arthritis Clinic (EAC) (1029 cases and 895 controls) (Table 1). The EIRA study population is a population-based case–control study of incident cases of RA in which all patients fulfilled the American College of Rheumatology (ACR) 1987 criteria.22 Controls were randomly selected from the Swedish national population registry, taking into consideration the patient's age, sex and residential area. More details about the EIRA study population have been described elsewhere.19 Dutch Caucasian individuals with RA, all of whom fulfilled the ACR classification criteria for RA were studied and described elsewhere.23 Controls were unrelated Dutch Caucasians with no history of RA. The cases in the NARAC study population consisted of RA patients of self-reported white ancestry who were randomly drawn from four different sample groups of patients and controls were recruited from the New York Cancer Project.24, 25 All these studies were conducted after obtaining approval from the Regional Ethics Committees and in accordance with the Declaration of Helsinki.
ACPA status and genotyping
The levels of ACPA in samples from the EIRA and Leiden EAC study were determined using Immunoscan-RA Mark2 ELISA (Euro-diagnostica, Arnhem, the Netherlands) as described elsewhere.26 Cases with antibody levels >25 U/ml were regarded as ACPA positive. In the NARAC study, ACPA levels were determined using the second-generation commercial anti-CCP ELISA (INOVA Diagnostics Inc., San Diego, CA, USA). Cases with serum antibody levels >20 U/ml were regarded as ACPA positive. The HLA-DRB1 type of samples of the Swedish cohort was determined by sequence-specific primer–PCR analysis using low-resolution HLA-DR and DRB1*04 kits from Olerup SSP (Saltsjöbaden, Sweden). HLA-DRB1 alleles DRB1*01, DRB1*04 and DRB1*10 were defined as SE. Subjects with the SE allele were classified as having a single or double SE allele. The method used for genotyping in the Dutch cohort and the NARAC cohort has been reported elsewhere.27 The method used for SNP genotyping of EIRA samples, which was previously performed,16 and of Leiden EAC samples was TaqMan allelic discrimination (Applied Biosystems, Foster City, CA, USA). The call rates for rs6314 and rs1328674 in all three cohorts were >97 and >98%, respectively. For the NARAC study, genotyping data for rs6314 and rs1328674 were extracted from the genome-wide association study, which was performed using the Illumina HumanHap550 array (Illumina, Inc., San Diego, CA, USA).4
Real-time PCR
For mRNA expression analysis, synovial fibroblasts from four individuals with RA were used (Dominion Pharmakine, SL, Bizkaia, Spain). Cells were cultured without/with 400 ng/ml LPS (Sigma Chemicals, St Louis, MO, USA) and 200 U/ml IFN-γ (Nordic BioSite, Taby, Sweden) for 24 h and harvested for RNA isolation. Cells were lysed and total RNA was extracted using RNeasy Mini Kit (QIAGEN, Valencia, CA, USA) according to the manufacturer's protocol. Samples were treated with DNAse (Qiagen RNase free DNase set) for 20 min at room temperature to avoid contamination with genomic DNA. A volume of 1 μg total RNA for each sample was converted into cDNA using SuperScript reverse transcriptase (Invitrogen, Carlsbad, CA, USA) and random hexamer primers. Real-time PCR was performed using the ABI PRISM 7900HT Sequence Detection System (Applied Biosystems) and TaqMan Gene Expression Assays for HTR2A, HLA-DRB1 and GAPDH (Hs00167241, Hs99999917 and 4310884E, respectively) (Applied Biosystems) with a two-step protocol (50°C for 2 min, 95°C for 10 min, followed by 40 cycles of 95°C for 15 s and 60°C for 1 min). The reactions were performed in duplicate in optical 384-well plates (Applied Biosystems) in a total volume of 20 μl. Relative gene expression was calculated using the calibration curve method and normalized to the GAPDH expression.
Immunohistochemical analysis
To perform detection of proteins in synovial tissue, we used tissue biopsy samples from four RA patients. Staining of cryostat sections with mouse monoclonal anti-HLA-DR (BD Biosciences, San José, CA, USA) and rabbit polyclonal anti-5-HT2A (Sigma) antibodies was performed using a standard protocol.28 Isotype-matched irrelevant antibodies were used as negative controls. Tissue sections of mouse brain stem were used as positive tissue control for 5-HT2A. Stained sections were examined using a Polyvar II microscope (Reichert-Jung, Vienna, Austria) and photographed with a digital Leica camera 300F (Leica, Cambridge, UK). Positive staining was indicated as brown deposits. Double immunofluorescence staining was performed using mouse monoclonal anti-HLA-DR (BD Biosciences) and rabbit polyclonal anti-5-HT2A antibodies (Sigma Chemicals) as published previously.29
Statistical analysis
The χ2-test was used to compare the genotype and allele frequencies between patients and controls in all three studies. Haplotype analysis was performed using the HaploView 4.0 software (http://www.broad.mit.edu/mpg/haploview). When a DNA sample genotyping failed for >50% of the SNPs, it was excluded from statistical evaluation. The distribution of genotypes for SNPs rs6314 and rs1328674 in all three studies was in agreement with Hardy–Weinberg equilibrium (P>0.05). The haplotypes for each individual were assigned using the PHASE software.30 The interaction effects between the HTR2A haplotype and HLA-DRB1 SE alleles from all three studies were evaluated using the departure from the additive effects as the interaction criteria,31 and the attributable proportion (AP) due to interaction with 95% CI was calculated. A more detailed description has been described elsewhere.32 For meta-analysis, we used a random effects model (excluding estimates that deviated from the homogeneity of ORs) to calculate pooled ORs.33
Results
Interaction between HTR2A and HLA-DRB1 SE alleles
We performed an association study in the Swedish cohort as a follow-up of our previous finding of an association between HTR2A and risk of developing RA.16 Our previously found haplotype consisted of four tagging SNPs. However, because not all four SNPs were genotyped in the NARAC study, we decreased the number of SNPs to two. As could be seen from Table 2, the TC haplotype based on rs6314 and rs1328674, which is part of what was previously reported to be in association with RA,16 demonstrates a protective effect in the EIRA study (P=0.005 in χ2-test). Further, we addressed the hypothesis for association in two other study populations that was replicated in the NARAC study (P=0.006 in χ2-test) but not in the Leiden EAC study (P=0.8206 in χ2-test), although the control group had a 0.2% higher frequency compared with the patients group (Table 2).
The interaction between HTR2A and HLA-DRB1 SE alleles was quantified by means of calculating the AP (Table 3). For the EIRA and Leiden EAC studies, interaction was investigated in relation to the development of both ACPA-positive and ACPA-negative RA, respectively, but only a trend for an antagonistic interaction between these two genes regarding risk of developing ACPA-positive RA was observed in EIRA (AP 0.19 (95% CI –0.06 to 0.45)) and Leiden EAC studies (AP 0.2 (95% CI −0.15 to 0.55)) (Table 3). No interaction was observed with regard to risk of ACPA-negative RA in either of these two studies.
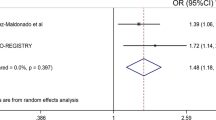
As EIRA and Leiden EAC studies demonstrated a trend toward interaction between HTR2A and HLA-DRB1 SE alleles in the development of ACPA-positive RA, we performed a meta-analysis for these two groups on the basis of ORs. This analysis demonstrated a statistically significant interaction (AP 0.19 (95% CI 0.04–0.34)) (Table 4).
For the NARAC study, which was restricted to ACPA-positive patients only, the interaction between HTR2A and HLA-DRB1 SE alleles was statistically significant (AP 0.26 (95% CI 0.05–0.46)) (Table 3), which is a replication of our finding for two other studies (EIRA and EAC). We noticed that meta-analysis in all three groups demonstrated significant heterogeneity (not shown) and must be taken with caution.
5-HT2A and HLA-DR mRNA and protein expression
Using quantitative real-time PCR, we observed that HTR2A and HLA-DRB1 mRNA are expressed in untreated synovial fibroblasts from RA patients. Induction with LPS and IFN-γ induced the expression of HLA-DRB1 mRNA, whereas it did not markedly change the expression of HTR2A mRNA in RA synovial fibroblasts (Figure 1). Next, we investigated the expression of HTR2A and HLA-DR proteins in synovial tissue from patients with RA and we could detect the cytoplasmic and membranous staining for 5-HT2A in synovial lining and sublining cells (Figures 2a–c). As expected, HLA-DR was abundantly expressed in RA synovial tissue (data not shown).
HTR2A and HLA-DRB1 mRNA are expressed in untreated fibroblasts from rheumatoid arthritis (RA) patients. Expression of HTR2A and HLA-DRB1 mRNA in RA synovial fibroblasts untreated (open bars) or stimulated (filled bars) with LPS and IFN-γ for 24 h. The data represent mean (±SE) of values obtained from three separate experiments. The results are expressed as fold changes, considering 1 as the value of untreated cells.
5-HT2A is expressed in synovial tissues of patients with rheumatoid arthritis (RA) and colocalizes with HLA-DR in synovial fibroblasts. Photographs illustrating (a, b) brown immunoperoxidase staining of 5-HT2A-positive cells in representative synovial tissue sections from patients with RA counterstained with haematoxylin; (c) negative control (original magnification × 250–500). Double fluorescence staining illustrating (d) 5-HT2A staining, (e) HLA-DR staining and (f) merged images of rheumatoid arthritis synovial fibroblasts induced with IFN-γ and LPS (original magnification × 500).
To determine whether 5-HT2A and HLA-DR are colocalized in synovial fibroblasts, double staining of cells treated with IFN-γ and LPS was performed (Figures 2d–f). As can be observed from Figure 2, two proteins are colocalized in synovial fibroblasts, although there were cells that expressed only 5-HT2A or HLA-DR.
Discussion
The major finding in our study is a discovery of interactions between one of the serotonin receptors and MHC class II molecules, which is important for the development of RA. Such a relationship has never been shown or indicated previously and may demonstrate an unknown pathway in disease development, and it is also a potential tool to ameliorate the influence of the strongest genetic risk factor for RA, SE alleles. We found not only statistical evidence for such an interaction, but also identified cells that express both proteins, which is a totally new finding.
Rheumatoid arthritis is a complex disease in which genetic and environmental factors contribute to disease development. When investigating the association of single genes with the risk of a complex disease, ORs are almost always low to moderate. This reflects that a specific phenotype is a result of a combination of different low to moderately contributing genes and environmental factors. One example is the PTPN22 risk allele (R620W), which by itself is a moderate or no risk factor for the development of RA but, when interacting with HLA-DRB1 SE alleles, a gene strongly connected to different autoimmune diseases, the ORs for developing RA becomes very high.7 So taking in consideration the epistasis and interaction of genes with environmental factors is appropriate.
Polymorphisms in HTR2A have frequently been reported to be in association with different psychiatric diseases such as schizophrenia and bipolar anxiety disorder.34 However, evidence for the involvement of the serotonin system in inflammation is increasing.13, 14, 15 Notably, RA is negatively correlated with schizophrenia.35 Partition of RA into ACPA-positive and ACPA-negative subtypes is adequate, as more evidence points toward the fact that these two subtypes differ in etiology and pathophysiology. In this paper, we have shown that HTR2A, recently found to be in association with RA,16 interacts with HLA-DRB1 SE alleles in the ACPA-positive group of patients. HLA-DRB1 SE alleles have no effect on the development of ACPA-negative RA and that could be the reason why this interaction is not observed in this subgroup of the disease. The enhanced risk for development of RA due to interaction between HTR2A and HLA-DRB1 SE alleles was shown in the meta-analysis of EIRA and Leiden EAC studies. Our observation that the meta-analysis for the three cohorts did not pass the test for heterogeneity may have an obvious reason, which is that the estimates based on NARAC are significantly higher than the estimates from EIRA and Leiden EAC studies. However, we could demonstrate a statistically significant interaction between HTR2A and HLA-DRB1-SE both in meta-analysis (EIRA and EAC) and in the NARAC cohort. As previously noticed for the interaction between HLA-DRB1 SE alleles and PTPN22,7 it is illustrative that inspite of substantial differences in the design of these studies, data for all three populations consistently point to the gene–gene interaction. A possible biological background for the interaction between HTR2A and HLA-DRB1 SE alleles could be suggested at the cellular level by demonstrating that these two proteins are detected in inflamed synovial tissue simultaneously.
On the basis of the results from our study, we can speculate about a biological interaction between these two proteins. One possibility could be that there is a crosstalk between the signaling pathway related to HTR2A and MHC class II expression regulation. Furthermore, as we could detect these two proteins in the same cell in synovial fibroblasts in vitro, a physical interaction between them should not be excluded. The protective association of the TC haplotype in HTR2A with the development RA and the interaction between HTR2A and HLA-DRB1 SE alleles make it very tempting to speculate whether this could be one of the reasons for the inverse relation between schizophrenia and RA that has been reported frequently in literature.35 We can hypothesize that the protective haplotype in HTR2A with regard to RA is a susceptible haplotype with regard to schizophrenia. Moreover, it has been shown that HLA-DRB1 SE alleles are associated with schizophrenia,36, 37 hence, perhaps in contrast to RA, there is a synergistic interaction between this gene and the HTR2A TC haplotype among schizophrenia patients. More functional studies have to be carried out before further details regarding the interaction between 5-HT2A and HLA-DR proteins can be revealed, as our data are based on a limited number of experiments. In conclusion, our data show strong evidence for an interaction between 5-HT2A and MHC type II molecules in the development of RA.
References
Stastny P : Association of the B-cell alloantigen DRw4 with rheumatoid arthritis. N Engl J Med 1978; 298: 869–871.
Begovich AB, Carlton VE, Honigberg LA et al: A missense single-nucleotide polymorphism in a gene encoding a protein tyrosine phosphatase (PTPN22) is associated with rheumatoid arthritis. Am J Hum Genet 2004; 75: 330–337.
Remmers EF, Plenge RM, Lee AT et al: STAT4 and the risk of rheumatoid arthritis and systemic lupus erythematosus. N Engl J Med 2007; 357: 977–986.
Plenge RM, Seielstad M, Padyukov L et al: TRAF1-C5 as a risk locus for rheumatoid arthritis – a genomewide study. N Engl J Med 2007; 357: 1199–1209.
Kurreeman FA, Padyukov L, Marques RB et al: A candidate gene approach identifies the TRAF1/C5 region as a risk factor for rheumatoid arthritis. PLoS Med 2007; 4: e278.
Raychaudhuri S, Remmers EF, Lee AT et al: Common variants at CD40 and other loci confer risk of rheumatoid arthritis. Nat Genet 2008; 40: 1216–1223.
Kallberg H, Padyukov L, Plenge RM et al: Gene-gene and gene-environment interactions involving HLA-DRB1, PTPN22, and smoking in two subsets of rheumatoid arthritis. Am J Hum Genet 2007; 80: 867–875.
Padyukov L, Silva C, Stolt P, Alfredsson L, Klareskog L : A gene-environment interaction between smoking and shared epitope genes in HLA-DR provides a high risk of seropositive rheumatoid arthritis. Arthritis Rheum 2004; 50: 3085–3092.
Abdouh M, Storring JM, Riad M et al: Transcriptional mechanisms for induction of 5-HT1A receptor mRNA and protein in activated B and T lymphocytes. J Biol Chem 2001; 276: 4382–4388.
Sternberg EM, Trial J, Parker CW : Effect of serotonin on murine macrophages: suppression of Ia expression by serotonin and its reversal by 5-HT2 serotonergic receptor antagonists. J Immunol 1986; 137: 276–282.
Hoyer D, Hannon JP, Martin GR : Molecular, pharmacological and functional diversity of 5-HT receptors. Pharmacol Biochem Behav 2002; 71: 533–554.
Idzko M, Panther E, Stratz C et al: The serotoninergic receptors of human dendritic cells: identification and coupling to cytokine release. J Immunol 2004; 172: 6011–6019.
Guillet-Deniau I, Burnol AF, Girard J : Identification and localization of a skeletal muscle secrotonin 5-HT2A receptor coupled to the Jak/STAT pathway. J Biol Chem 1997; 272: 14825–14829.
Cloez-Tayarani I, Petit-Bertron AF, Venters HD, Cavaillon JM : Differential effect of serotonin on cytokine production in lipopolysaccharide-stimulated human peripheral blood mononuclear cells: involvement of 5-hydroxytryptamine2A receptors. Int Immunol 2003; 15: 233–240.
Yu B, Becnel J, Zerfaoui M, Rohatgi R, Boulares AH, Nichols CD : Serotonin 5-hydroxytryptamine(2A) receptor activation suppresses tumor necrosis factor-alpha-induced inflammation with extraordinary potency. J Pharmacol Exp Ther 2008; 327: 316–323.
Kling A, Seddighzadeh M, Arlestig L, Alfredsson L, Rantapaa-Dahlqvist S, Padyukov L : Genetic variations in the serotonin 5-HT2A receptor gene (HTR2A) are associated with rheumatoid arthritis. Ann Rheum Dis 2008; 67: 1111–1115.
Plenge RM, Padyukov L, Remmers EF et al: Replication of putative candidate-gene associations with rheumatoid arthritis in >4,000 samples from North America and Sweden: association of susceptibility with PTPN22, CTLA4, and PADI4. Am J Hum Genet 2005; 77: 1044–1060.
Mattey DL, Dawes PT, Clarke S et al: Relationship among the HLA-DRB1 shared epitope, smoking, and rheumatoid factor production in rheumatoid arthritis. Arthritis Rheum 2002; 47: 403–407.
Klareskog L, Stolt P, Lundberg K et al: A new model for an etiology of rheumatoid arthritis: smoking may trigger HLA-DR (shared epitope)-restricted immune reactions to autoantigens modified by citrullination. Arthritis Rheum 2006; 54: 38–46.
van Oosterhout M, Bajema I, Levarht EW, Toes RE, Huizinga TW, van Laar JM : Differences in synovial tissue infiltrates between anti-cyclic citrullinated peptide-positive rheumatoid arthritis and anti-cyclic citrullinated peptide-negative rheumatoid arthritis. Arthritis Rheum 2008; 58: 53–60.
van Dongen H, van Aken J, Lard LR et al: Efficacy of methotrexate treatment in patients with probable rheumatoid arthritis: a double-blind, randomized, placebo-controlled trial. Arthritis Rheum 2007; 56: 1424–1432.
Arnett FC, Edworthy SM, Bloch DA et al: The American Rheumatism Association 1987 revised criteria for the classification of rheumatoid arthritis. Arthritis Rheum 1988; 31: 315–324.
van Aken J, van Bilsen JH, Allaart CF, Huizinga TW, Breedveld FC : The Leiden Early Arthritis Clinic. Clin Exp Rheumatol 2003; 21: S100–S105.
Jawaheer D, Seldin MF, Amos CI et al: A genomewide screen in multiplex rheumatoid arthritis families suggests genetic overlap with other autoimmune diseases. Am J Hum Genet 2001; 68: 927–936.
Jawaheer D, Lum RF, Amos CI, Gregersen PK, Criswell LA : Clustering of disease features within 512 multicase rheumatoid arthritis families. Arthritis Rheum 2004; 50: 736–741.
Ronnelid J, Wick MC, Lampa J et al: Longitudinal analysis of citrullinated protein/peptide antibodies (anti-CP) during 5 year follow up in early rheumatoid arthritis: anti-CP status predicts worse disease activity and greater radiological progression. Ann Rheum Dis 2005; 64: 1744–1749.
Huizinga TW, Amos CI, van der Helm-van Mil AH et al: Refining the complex rheumatoid arthritis phenotype based on specificity of the HLA-DRB1 shared epitope for antibodies to citrullinated proteins. Arthritis Rheum 2005; 52: 3433–3438.
Ulfgren AK, Lindblad S, Klareskog L, Andersson J, Andersson U : Detection of cytokine producing cells in the synovial membrane from patients with rheumatoid arthritis. Ann Rheum Dis 1995; 54: 654–661.
Westman M, Korotkova M, af Klint E et al: Expression of microsomal prostaglandin E synthase 1 in rheumatoid arthritis synovium. Arthritis Rheum 2004; 50: 1774–1780.
Stephens M, Donnelly P : A comparison of bayesian methods for haplotype reconstruction from population genotype data. Am J Hum Genet 2003; 73: 1162–1169.
Rothman K : Epidemiology. An Introduction. New York: Oxford University Press, 2002.
Andersson T, Alfredsson L, Kallberg H, Zdravkovic S, Ahlbom A : Calculating measures of biological interaction. Eur J Epidemiol 2005; 20: 575–579.
Fleiss JL : The statistical basis of meta-analysis. Stat Methods Med Res 1993; 2: 121–145.
Norton N, Owen MJ : HTR2A: association and expression studies in neuropsychiatric genetics. Ann Med 2005; 37: 121–129.
Eaton WW, Hayward C, Ram R : Schizophrenia and rheumatoid arthritis: a review. Schizophr Res 1992; 6: 181–192.
Wright P, Donaldson PT, Underhill JA, Choudhuri K, Doherty DG, Murray RM : Genetic association of the HLA DRB1 gene locus on chromosome 6p21.3 with schizophrenia. Am J Psychiatry 1996; 153: 1530–1533.
Sasaki T, Matsushita M, Nanko S, Fukuda R, Kennedy JL, Tokunaga K : Schizophrenia and the HLA-DRB1 gene in the Japanese population. Am J Psychiatry 1999; 156: 771–773.
Acknowledgements
The EIRA study was supported by grants from the Swedish Medical Research Council, Swedish Council for Working life and Social Research, King Gustaf V's 80-year foundation, the Swedish Rheumatism Foundation, Stockholm County Council, the insurance company AFA and Combine (Vinnova). We thank the RA patients and controls for participating in the study; Eva Jemseby, Marie-Louise Serra, Lena Nise and Camilla Bengtsson for invaluable contributions to the collection of data and maintenance of the EIRA database; as well as Johan Rönnelid (Unit of Clinical Immunology Uppsala University/Akademiska Hospital) and Hiba Mahdi (Karolinska University Hospital) for performing the analysis of anti-CCP. The Leiden EAC study was supported by grants from the Duthch Arthritis foundation, the European program grant FP6 AutoCure and FP1 Masterswitch, as well as a grant from the center for Medical System Biology within the framework of the Netherlands Genomic Initiative. We thank Peter Gregersen and the NARAC investigators for providing continuing access to genotyping data on the NARAC cohort; the NARAC study is supported by grants from the National Institutes of Health RO1-AR44422 and NO1-AR-2-2263.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Rights and permissions
About this article
Cite this article
Seddighzadeh, M., Korotkova, M., Källberg, H. et al. Evidence for interaction between 5-hydroxytryptamine (serotonin) receptor 2A and MHC type II molecules in the development of rheumatoid arthritis. Eur J Hum Genet 18, 821–826 (2010). https://doi.org/10.1038/ejhg.2010.12
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ejhg.2010.12
Keywords
This article is cited by
-
New polymorphisms associated with response to anti-TNF drugs in patients with moderate-to-severe plaque psoriasis
The Pharmacogenomics Journal (2018)
-
Genetic variation in the serotonin receptor gene affects immune responses in rheumatoid arthritis
Genes & Immunity (2013)