- NEWS FEATURE
The secret lives of cells — as never seen before
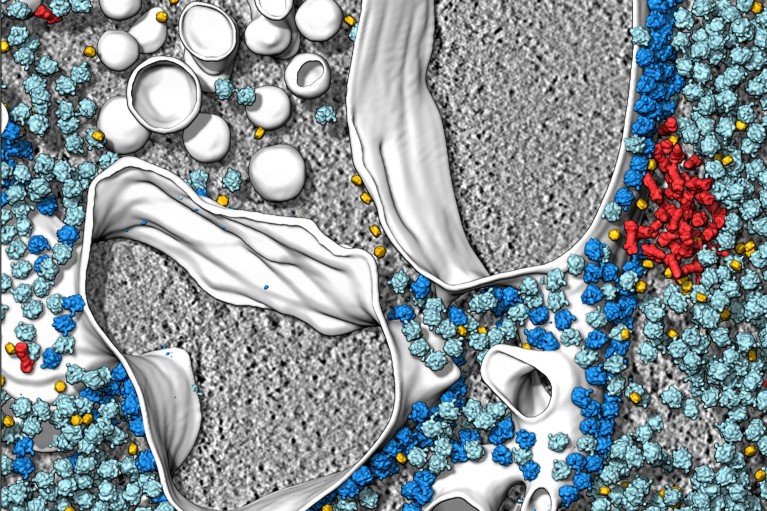
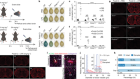
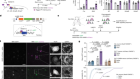
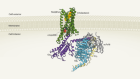
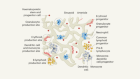
Cryo-electron tomography and related techniques can showcase the insides of cells in striking detail. Credit: S. Albert et al./PNAS (CC BY 4.0)
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 598, 558-560 (2021)
doi: https://doi.org/10.1038/d41586-021-02904-w
References
Medalia, O. et al. Science 298, 1209–1213 (2002).
Murphy, G. E., Leadbetter, J. R. & Jensen, G. J. Nature 442, 1062–1064 (2006).
Mahamid, J. et al. Science 351, 969–972 (2016).
Watanabe, R. et al. Cell 182, 1508–1518 (2020).
Deniston, C. K. et al. Nature 588, 344–349 (2020).
Bäuerlein, F. J. B. et al. Cell 171, 179–187 (2017).
Guo, Q. et al. Cell 172, 696–705 (2018).
Tegunov, D., Xue, L., Dienemann, C., Cramer, P. & Mahamid, J. Nature Methods 18, 186–193 (2021).
Ho, C.-M. et al. Nature Methods 17, 79–85 (2020).
Hoffman, D. P. et al. Science 367, eaaz5357 (2020).
Heinrich, L. et al. Nature https://doi.org/10.1038/s41586-021-03977-3 (2021).
Mendonça, L. et al. Nature Commun. 12, 4629 (2021).

 Revolutionary microscopy technique sees individual atoms for first time
Revolutionary microscopy technique sees individual atoms for first time
 The revolution will not be crystallized: a new method sweeps through structural biology
The revolution will not be crystallized: a new method sweeps through structural biology
 Calling cell biologists to try cryo-ET
Calling cell biologists to try cryo-ET