- NEWS
The mutation that helps Delta spread like wildfire
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 596, 472-473 (2021)
doi: https://doi.org/10.1038/d41586-021-02275-2
References
Liu, Y. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.08.12.456173 (2021).
Peacock, T. P. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.05.28.446163 (2021).
Saito, A. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.06.17.448820 (2021).
Zhang, J. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.08.17.456689 (2021).
Lubinski, B. et al. Preprint at bioRxiv https://doi.org/10.1101/2021.06.30.450632 (2021).

 How do vaccinated people spread Delta? What the science says
How do vaccinated people spread Delta? What the science says
 How the Delta variant achieves its ultrafast spread
How the Delta variant achieves its ultrafast spread
 COVID vaccines slash viral spread – but Delta is an unknown
COVID vaccines slash viral spread – but Delta is an unknown
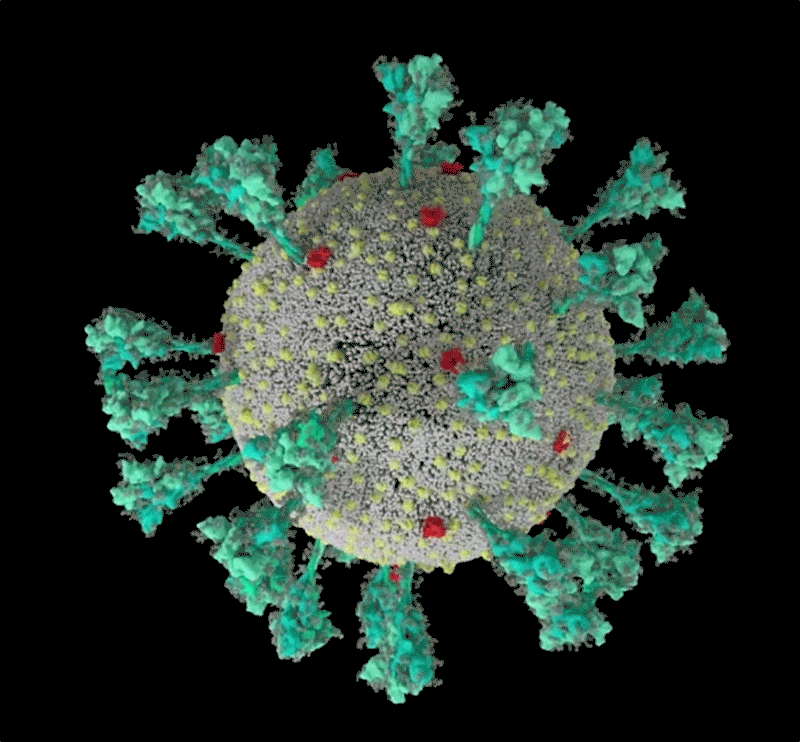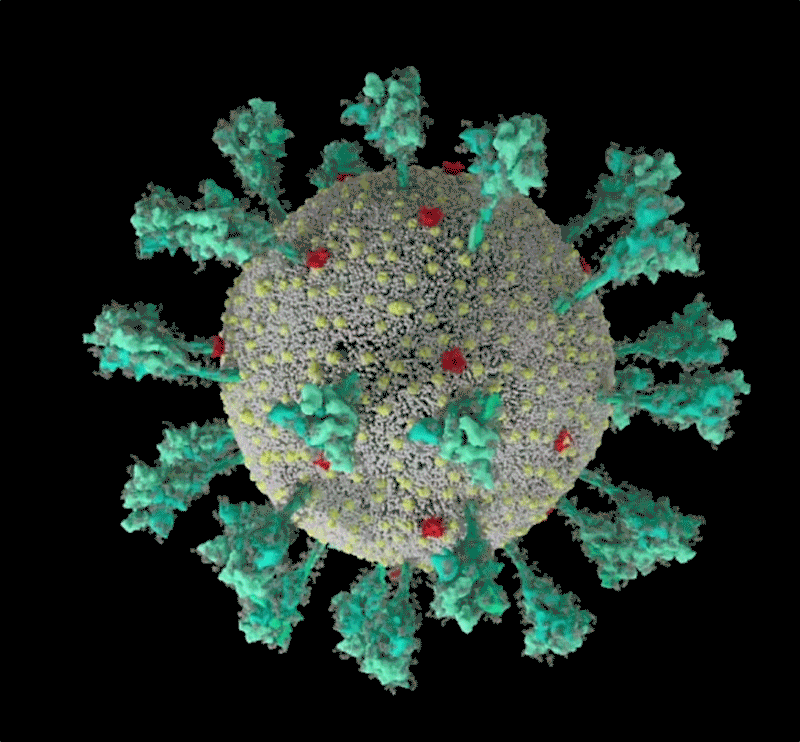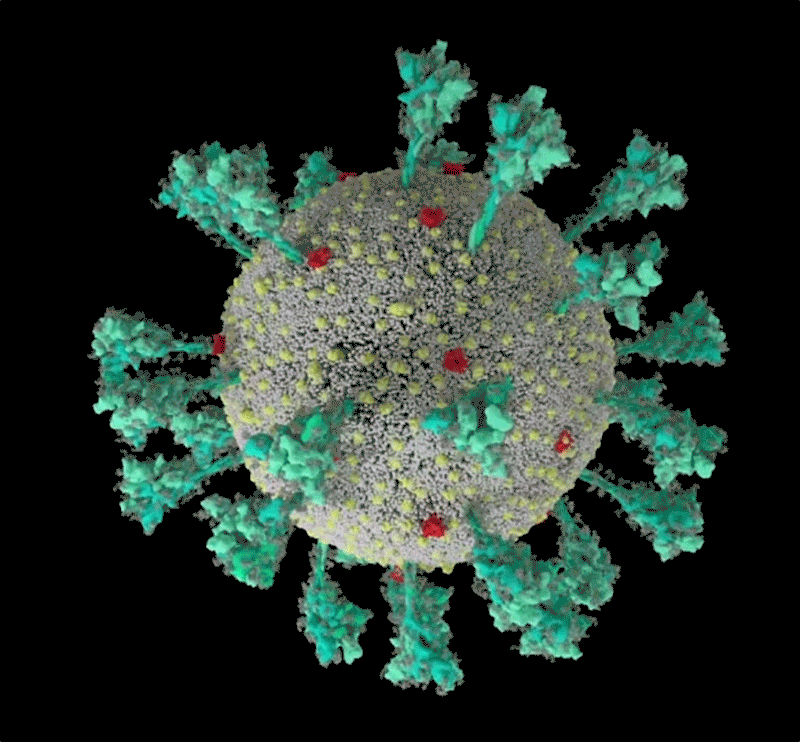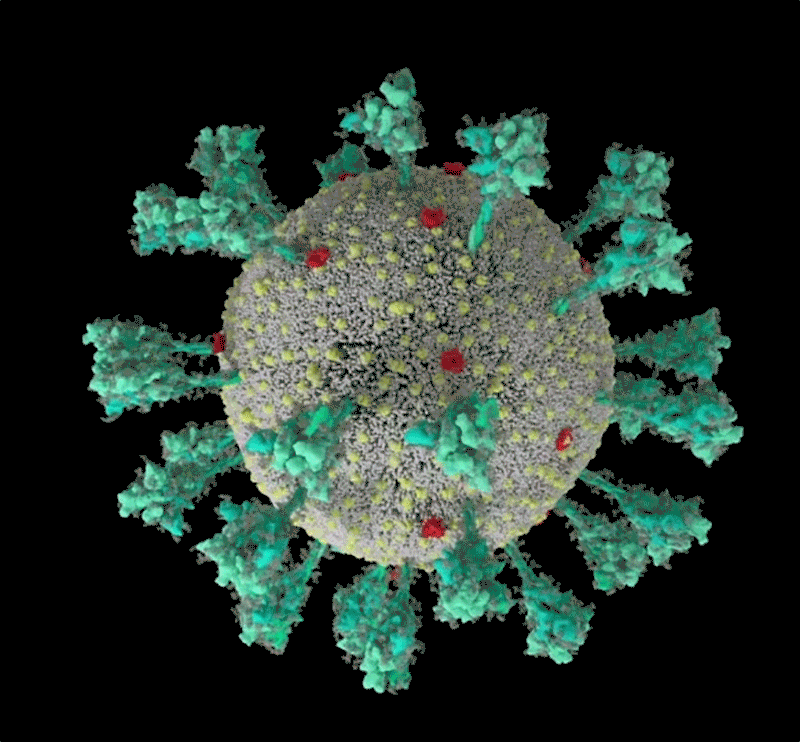 How the coronavirus infects cells — and why Delta is so dangerous
How the coronavirus infects cells — and why Delta is so dangerous