- OUTLOOK
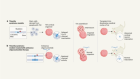
Could tracking RNA in body fluids reveal disease?

Illustration: David Parkins
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 582, S2-S4 (2020)
doi: https://doi.org/10.1038/d41586-020-01763-1
This article is part of Nature Outlook: Extracellular RNA, an editorially independent supplement produced with the financial support of third parties. About this content.
References
Van Deun, J. et al. Nature Meth. 14, 228–232 (2017).
McKiernan J. et al. JAMA Oncol. 2, 882–889 (2016).
McKiernan J. et al. Eur. Urol. 74, 731–738 (2018).
Geeurickx, E. et al. Nature Commun. 10, 3288 (2019).

 Artificial nanoparticles are not as good as the real thing
Artificial nanoparticles are not as good as the real thing
 The biologist on the hunt for extracellular ribosomes
The biologist on the hunt for extracellular ribosomes
 Dietary RNA is ripe for investigation
Dietary RNA is ripe for investigation
 Do the microRNAs we eat affect gene expression?
Do the microRNAs we eat affect gene expression?
 Unravelling the mysteries of microRNA in breast milk
Unravelling the mysteries of microRNA in breast milk
 How extracellular vesicles can enhance drug delivery
How extracellular vesicles can enhance drug delivery
 Inside the stem-cell pharmaceutical factory
Inside the stem-cell pharmaceutical factory
 Plant vesicles inspire methods to protect crops
Plant vesicles inspire methods to protect crops
 Research round-up: extracellular RNA
Research round-up: extracellular RNA