- NEWS AND VIEWS
Transfer of ubiquitin protein caught in the act
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 578, 372-373 (2020)
doi: https://doi.org/10.1038/d41586-020-00325-9
References
Lydeard, J. R., Schulman, B. A. & Harper, J. W. EMBO Rep. 14, 1050–1061 (2013).
Saha, A. & Deshaies, R. J. Mol. Cell 32, 21–31 (2008).
Duda, D. M. et al. Cell 134, 995–1006 (2008)
Read, M. A. et al. Mol. Cell. Biol. 20, 2326–2333 (2000).
Podust, V. N. et al. Proc. Natl Acad. Sci. USA 97, 4579–4584 (2000).
Baek, K. et al. Nature 578, 461–466 (2020).
Zheng, N. et al. Nature 416, 703–709 (2002).
Cui, J. et al. Science 329, 1215–1218 (2010).
Sakata, E. et al. Nature Struct. Mol. Biol. 14, 167–168 (2007).
Scott, D. C. et al. Cell 166, 1198–1214 (2016).
Sievers, Q. L., Gasser, J. A., Cowley, G. S., Fischer, E. S. & Ebert, B. L. Blood 132, 1293–1303 (2018).
Verma, R., Mohl, D. & Deshaies, D. J. Mol. Cell https://doi.org/10.1016/j.molcel.2020.01.010 (2020).
Smith, B. E. et al. Nature Commun. 10, 131 (2019).
Competing Interests
Both R.J.D. and N.W.P. are employees of Amgen. R.J.D. is in addition an officer of the corporation.

 Read the paper: NEDD8 nucleates a multivalent cullin–RING–UBE2D ubiquitin ligation assembly
Read the paper: NEDD8 nucleates a multivalent cullin–RING–UBE2D ubiquitin ligation assembly
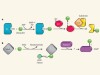 Ubiquitination without E1 and E2 enzymes
Ubiquitination without E1 and E2 enzymes
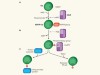 Deciphering the catalytic mechanism of bacterial ubiquitination
Deciphering the catalytic mechanism of bacterial ubiquitination