Host: Benjamin Thompson
Welcome back to the Nature Podcast. This week, we’ll be hearing about the researchers mapping mouse cells…
Host: Shamini Bundell
And learning about the potential benefits of cataloguing the world’s viruses. I’m Shamini Bundell.
Host: Benjamin Thompson
And I’m Benjamin Thompson.
[Jingle]
Host: Benjamin Thompson
What do you want to be when you grow up? The young me really wanted to be a professional footballer, but sadly complete lack of talent was against me. Shamini, what about you, what did you want to be when you grew up?
Host: Shamini Bundell
Oh, well I am still aiming to be a time-travelling space pirate actually.
Host: Benjamin Thompson
Nice, well, I guess there’s still time for that, particularly if time travel is involved, I suppose.
Host: Shamini Bundell
Yeah, I’ve got ages to work on it.
Host: Benjamin Thompson
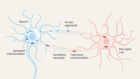
Well, today’s first story is about working out what happens to cells as they grow up – that’s something that developmental biologists often want to know when researching embryos. How do cells change over time? What kind of tissues will they end up being part of? And what are the processes that control how this fate gets decided? This week, Nature has published two papers from two different groups that each have produced extremely detailed molecular maps of mouse development, profiling the cells in the embryo as together they build a complete animal. Reporter Anand Jagatia has been speaking to the researchers involved.
Interviewee: Cole Trapnell
Over the span of around three weeks, a single cell develops into a mouse. In that time, millions and millions of cells are generated. Ultimately, they diversify into performing all kinds of different functions. It’s a tremendously complicated process and the programme that directs it is somehow encoded in the mouse genome.
Interviewer: Anand Jagatia
This is Cole Trapnell from the University of Washington, and for him, there’s no shortage of reasons for studying development.
Interviewee: Cole Trapnell
From a practical perspective, it’s important to understand the genetic architecture that controls development because so many developmentally regulated genes when misregulated cause disease, but I think that understanding the instructions for how to make all of the different types of cells that are found in our body is a fundamental goal in biology. Many of us who work on this are simply captivated by the complexity of the developmental programme. It’s a stunning example of how individual cells communicating with each other over short distances using proteins and nucleic acids can collaborate to make something as complicated as a whole animal.
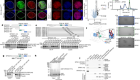
Interviewer: Anand JagatiaSo, how do you study something this complicated, involving millions of cells and tens of thousands of genes? One way is to produce an atlas, but instead of mapping continents and oceans, researchers map the identity and gene expression of all the cell types in the embryo as they start to form the organs of an adult mouse. This week, Nature features two maps from different groups, looking at key stages in this process. The first paper is from a group in the UK. Their atlas looks at a 48-hour window in development, starting at day 6.5. Bertie Göttgens from the University of Cambridge is one of the authors.
Interviewee: Bertie Göttgens
At about 6.5 days, the mouse embryo is composed of an outer layer of cells which surrounds an inner layer of cells, and those inner cells are unspecified, and what that means is they have the capacity to give rise to any cell in the adult body. There’s a few hundred of those cells at day 6.5, and then at day 8.5, the embryo would have grown to be more than 100,000 cells, so that we are generating the progenitors of all the major organs.
Interviewer: Anand Jagatia
Cole Trapnell, who we heard from earlier, is one of the authors that worked on the second atlas. This group looked at days 9.5 to 13.5.
Interviewee: Cole Trapnell
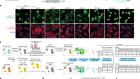
That interval in development is when many of the major organs really start to form. There’s so much that we don’t understand about the molecular regulation of almost every cell type in the body, in terms of how it develops. And our main goal in this project was to deploy a technology to watch as much of this particular window of development as we could.
Interviewer: Anand Jagatia
Across these two time windows, the number of cells in the embryo grows from the hundreds to the millions, and each one could be expressing thousands of genes, so producing an atlas of cell types is technically very challenging. Each of these papers used different techniques to do this, but the idea is the same. In a cell, the first stage of gene expression is to turn each stretch of DNA that you want to express into RNA, which is eventually read by the cell and turned into protein. This collection of RNA is called the cell’s transcriptome. Both these papers were able to sequence and label the entire transcriptome of individual cells in the embryo. This tells you exactly which genes are being expressed and by how much at different time points in development, and the labels mean you can trace each stretch of genetic sequence back to the cell it came from. Once you have that, you can analyse the fingerprint of gene expression from each cell and work out what kind of cell it is. For example, a heart muscle cell will express a specific set of hallmark genes that you can use to identify it. It’s a laborious process, but it allows you to reconstruct a picture of the whole embryo in incredible depth and at a resolution of single cells.
Interviewee: Bertie Göttgens
We are like explorers that are discovering new continents because there is so much to be discovered. It allows you to categorise all the different cells into different groups and also track over time how the status or the state of these cells changes and how an ancestral unspecified cell can take up multiple different outcomes.
Interviewer: Anand Jagatia
Previous work in this field has either focused on a handful of genes or just one time point, but what makes both of these atlases special is they look at all the genes in all cell types over time in a mammal. This kind of rich, in-depth data can be incredibly powerful. For example, in their paper, Bertie and his colleagues used their atlas to dig deeper into the early steps of blood development. In particular, they were able to shed light on a gene called TAL1, which regulates blood development but that can also cause leukaemia in humans and in mice models. Really, the atlases themselves are just a first step. Both datasets will be freely available online and provide a wealth of data that can be explored and interrogated by researchers in all sorts of ways. I asked Cole and Bertie about their hopes for the data.
Interviewee: Cole Trapnell
Right now, we have a measurement of all the genes that are being turned on and off as cells decide what they’re going to be, but I think what’s really important to do is be able to take those data and look at them and identify the genes that are actually controlling the fate decisions. So, try to use the data to infer the underlying genetic circuits that control a cell’s decision to become a heart muscle cell as opposed to something else.
Interviewee: Bertie Göttgens
We want to really use this technology to dig much deeper into the biological processes, some of those fundamental processes of how do these early cells make decisions. We are ready to embrace a phase of biological, hopefully even mechanistic, investigation of those early processes, taking full advantage, of course, of all the technologies that we now have.
Host: Benjamin Thompson
That was Bertie Göttgens, and before him, Cole Trapnell. You can read their papers over at nature.com/nature.
Host: Shamini Bundell
Coming up in the show, we’ll be hearing the latest lunar lander news – that’s in the News Chat. Now though, Anna Nagle is here with this week’s Research Highlights.
[Jingle]
Anna Nagle
Bat mums offer their offspring a helping wing when it comes to finding a place to roost, but draw a line when the young ‘uns are after food. To learn about bat parenting, researchers in Germany attached tiny trackers to 60 common noctule bats living in a Berlin forest. The team found that when the bats move to a new roosting site, mothers sometimes made repeated flights between the old and new sites to guide their progeny to the right location. However, when out foraging for food, young bats weren’t seen with their mothers any more often than they were with other bats, suggesting that parental guidance only goes so far. Fly over to that research in Biology Letters to read more.
[Jingle]
Anna Nagle
Most scientists will try and take research findings with a pinch of salt, but astronomers in the US have found a bit more than a pinch circling a star 1,300 light years away. A team of researchers have observed signatures of sodium chloride and potassium chloride out in the depths of space. Seasoned astronomers listening to this will of course know that these compounds have been seen before, and always around dying stars. However, in this new work, the compounds are around a young star, found in a hazy ring about the diameter of our solar system. It’s thought that the salt was formed when the dust that ultimately became the star was broken down into its component molecules. Mine that paper over at the Astrophysical Journal.
[Jingle]
Host: Shamini Bundell
There are a lot of viruses on the planet. It’s suggested that for every single organism on Earth, there is a specific virus that infects it. So, for example, there’s 400,000 species of beetle, and for every one of those, there’s probably at least one virus associated with them, if not more. So, when you start to think about all the animals, plants and bacteria out there, it’s possible that there are trillions of viral species on Earth. Now, trillions of viruses lurking out there in the world might seem a bit worrying, but many of these viruses could harbour useful resources. This week in Nature, virologist Jens Kuhn has co-written a Comment piece suggesting that we try to catalogue all the viruses in the world, in the hope that we can find something useful. Reporter Nick Howe wanted to learn more about creating an encyclopaedia of viruses, so he gave Jens a call and started by asking him how you might go about it.
Interviewee: Jens Kuhn
So now, it is possible to get to the genomes of viruses without necessarily growing them in culture. So, there are new methodologies out there now where you can simply take an environmental sample and sequence everything that is in there and then use computational methods to piece genomes together and then you can say so many different viruses are in this sample and their genomes look like x. So that has led to an explosion of virus data and so we had only like 3,000-4,000 viruses in our databases until about ten years ago, and now one paper after another is coming out describing thousands, and that poses problems of course. From an administrative point of view, somebody is needed to put these things into a database so that we at least know what’s out there. So, that’s ultimately the idea of the Comment – to raise awareness of the diversity of viruses out there, the changes that these new viruses bring to our understanding of the biosphere and the treasure trove that might be lying in this diversity, even in seemingly unimportant viruses.
Interviewer: Nick Howe
So, tapping into that treasure trove, as you put it, what things could we find it viruses that could be useful. What are the advantages of getting the taxonomy of viruses?
Interviewee: Jens Kuhn
I would say the easiest example is actually to go away from viruses a little bit and to look into bacteria. There’s a lot of bacteria out there that are seemingly completely unimportant for human life that live in deep-sea vents somewhere with really high temperatures. However, they encode a lot of enzymes and these enzymes have been useful in the past, for example, for dishwashing or laundry detergent manufacturers because if you want to break down protein stains, which of course is food on your clothes, you need to have ideally enzymes that break that up that work at higher temperatures so you can wash the clothes at higher temperatures. So now, if you look at viruses, viruses infect bacteria, they infect all other organisms, as far as we know, so that means that if you have a virus in a hot spring, deep-sea bacterium, that virus also needs to encode heat-stable proteins and so on.
Interviewer: Nick Howe
How could we go about cataloguing them all? Is it a feasible thing to undertake?
Interviewee: Jens Kuhn
Yeah, I think it is. I mean it’s daunting, of course, and sounds crazy at the moment. It’s a question of simple – well simple is in quotation marks – sequencing techniques to get as many genomes as possible and then very clever database programming that automatically sorts genomes in particular corners, maybe automatic alignments, methods, for these genomes to come up with rough phylogenetic trees without any human input, all the way to possible automatic classification.
Interviewer: Nick Howe
But how can we be certain if we’re not going about trying to culture these viruses in the lab, we’re not growing them in a host organism, that these are individual viruses and they’re not coming out as artefacts of the sequencing or they’re actually a different organism altogether?
Interviewee: Jens Kuhn
Yeah, that is definitely a problem. So, our methods are getting better in figuring out which pieces in a particular sample belong to which other piece, so the possibility that it’s a complete artefact, I would say, is relatively low. However, you need to assemble these individual pieces and therefore you do not know whether the actual gene that you have really works or whether you have individual, small mistakes in your assembly. So, if you are truly interested in a particular virus, then you will have to characterise it, you will have to try to grow it, you will have to try to see whether it’s pathogenic and so on.
Interviewer: Nick Howe
What would be the sort of cost and what resources would be required to do such a big taxonomical effort?
Interviewee: Jens Kuhn
For a very long time, there was no interest in ‘unimportant viruses’ and viruses of fungi and viruses of bacteria and so on, and that is a mindset that is still very much penetrating the field. I think the very first thing that needs to happen is that all virologists understand the immense treasure that is in virus diversity and to understand how many viruses are actually out there and why virus taxonomy is something that is now growing and is changing. The second thing is we need volunteers who actually want to help develop the databases and develop methodologies to classify viruses, and of course, actually become a part of the official process of taxonomy with the International Committee on Taxonomy of Viruses.
Interviewer: Nick Howe
Can we not just do a targeted approach and just look at a subset of viruses that are going to be affecting humans, for example?
Interviewee: Jens Kuhn
Well, you can, but I think the biggest problem is if you look at the big scourges that humans had in the last century, many of these viruses were simply not known to be human viruses before, right. So, SARS coronavirus or MERS coronavirus or even HIV, they were viruses of animals at the time, and then they made a species jump eventually and somehow ended up in the human population. So, I think it’s ultimately actually myopic to say we only look at things that are important to us because that means that you already know what is important, and I would say any good scientist would admit that we do not know what is important until it suddenly hits us.
Host: Shamini Bundell
That was Jens Kuhn. You can read his Comment over at nature.com/opinion.
Interviewer: Benjamin Thompson
Now, it’s time for the News Chat, and I’m joined by Lizzie Gibney, senior reporter here at Nature. Lizzie, thanks for stopping by.
Interviewee: Lizzie Gibney
Thank you, Ben.
Interviewer: Benjamin Thompson
Today we are going to do a little triple whammy of space stories, and one that just missed the cut for last week but I think is important that we do cover is, well, we need to tip our hats to Opportunity, the little Martian rover whose mission was declared over last week.
Interviewee: Lizzie Gibney
That’s right, and what an innings it had though. So, Opportunity was only supposed to have initially a 90-day mission, and it’s ended up lasting 15 years and travelling 45 kilometres across the Martian surface, so I think we can say it’s done pretty well.
Interviewer: Benjamin Thompson
In terms of bang for buck, I think it went above and beyond, right?
Interviewee: Lizzie Gibney
Absolutely, so one of the biggest things that it found was that there is this life-friendly water region on Mars, and that was quite nice actually because it was not really a serendipitous find, but it wasn’t an area they’d been planning to go to and then one of the project scientists spotted from orbit, using the Mars Reconnaissance Orbiter, that there was this region. They changed the trajectory of Opportunity and, yes, it found these lovely clay rocks that once hosted water.
Interviewer: Benjamin Thompson
And what ultimately happened to Opportunity?
Interviewee: Lizzie Gibney
So, unfortunately, it just basically stopped replying to signals coming from mission control, so there was a big dust storm, and its solar panels, which is what it relies on for energy and for electricity, were covered over, they think, by dust, and then mission control pinged it hundreds of times but alas did not get a response and eventually had to call it off.
Interviewer: Benjamin Thompson
Well, as you say, it has accomplished so much, and NASA has sent up another rover subsequently. What’s sort of the legacy of Opportunity and where do we go from here?
Interviewee: Lizzie Gibney
That’s right, so Opportunity had a twin called Spirit, but unfortunately that stopped working in 2009, but Curiosity, which landed in 2012, is still going – that’s nuclear-powered so we don’t have the same concerns about its solar panels – and there are three more rovers expected soon. So, NASA will have another one, the European Space Agency, and the Chinese National Space Administration, they are all planning to launch in 2020 and that’s when there is a particular alignment between the Earth and Mars which makes it easier to send a spacecraft there.
Interviewer: Benjamin Thompson
Lizzie, let’s move on to our next story today, and we’re going to move from Mars to the Moon, and a very different lander that could be ushering in a new era of lunar exploration.
Interviewee: Lizzie Gibney
Absolutely, so this is a lander called Beresheet, which in Hebrew means ‘in the beginning’, and that might be a clue as to who is launching it because it’s an Israeli craft. They are planning to launch this on Thursday evening US time, on 21st February. If successful, it would not only be the first Israeli craft on the Moon and that would be joining quite a prestigious bunch – the United States, China and Russia – but it would also be the first privately-backed lander. So, there’s a whole new bunch of commercial outfits who are trying to deliver payloads to the Moon. These are not commercial but they would be the first of those that would be private rather than totally run by some government space agency.
Interviewer: Benjamin Thompson
So, this is a very different way of funding space missions – is this the thin edge of the wedge, do you think?
Interviewee: Lizzie Gibney
I think so. So, there are about five or six commercial companies that I looked at I detail for this story, and there are no doubt more that are out there, and they are all really optimistic but obviously some of that is a bit of a sales pitch, but they really do think that they will have at least once launch by the end of 2021. So that’s not a long time away and you can imagine if we’ve got maybe six different private landers on the Moon by that point, that would be quite incredible, considering at the moment China is the only country to have any kind of lander or rover on the Moon. It would be a really different landscape, and think of what we could do – that’s exactly what the scientists are saying right now.
Interviewer: Benjamin Thompson
Well, let’s think about this mission which is launching this week, as you say. What’s that one geared up to do?
Interviewee: Lizzie Gibney
So, it’s mainly, as they say, about just trying to inspire a new generation. This is very much a symbolic mission and largely funded through philanthropy actually – they’ve had some massive donations from billionaires out there. It will also have a scientific mission as well, so it’s taking a magnetometer on board, so as it lands, it’s going to be studying the magnetic fields around the lunar rocks that it passes because there’s a bit of a mystery there. The Moon doesn’t currently have a magnetic field but it seems like it must have done at some point in order to magnetise these rocks, so by studying that they can try and figure out something out about the Moon’s history.
Interviewer: Benjamin Thompson
I mentioned before that this might be a new era of lunar exploration – is that true do you think?
Interviewee: Lizzie Gibney
It kind of depends. So, at the moment there are this plethora of companies and probably not all of them will last. The question is how many people are willing to pay to get to the Moon. Obviously, NASA always want lunar data so they will, but are there going to be enough other customers who want to get there? One of the big draws of the Moon is that it seems to have water – there’s certainly ice – and the question is could we mine water on the Moon. Water – you split it up into hydrogen and oxygen and boom, you’ve got rocket fuel. So, a lot of people think it could be very profitable as almost like a gas station for the rest of the solar system, but then that part is quite far away, so in this interim, will the industry survive? That is definitely something that is still a big question.
Interviewer: Benjamin Thompson
Okay, well let’s move on to our third space story then, Lizzie, and you talked about water there, and so let’s go to Neptune who is of course the god of the sea – how’s that for a link – and well, a new discovery has happened around this planet. What’s going on there?
Interviewee: Lizzie Gibney
Yes, so Neptune has a new moon, and it’s called Hippocamp. So, many of the really small inner moons around Neptune were discovered by Voyager 2, which is the spacecraft that’s now actually left our solar system, and that was back in 1989. This is another moon that was just discovered or that had kind of been seen quite hazily over the past decade or so but has now really been pinned down, and it was seen by astronomers looking at data from the Hubble Space Telescope, so rather than being a spacecraft winging past this moon and capturing it then, it was actually seen from Earth.
Interviewer: Benjamin Thompson
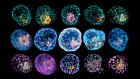
And well, you showed me a picture of what this moon looked like, and it looks like a little white dot in a sea of other white dots. How on Earth do you ascertain that that is a moon and not just sort of background noise?
Interviewee: Lizzie Gibney
Absolutely, well, it is pretty tiny. It’s 34 kilometres across, and this is the whole problem with trying to spot these moons, right, is they don’t produce any light of their own so they’re pretty dark and really, really far away, so it’s very, very hard to spot in the first place. What they did here was rather than just capturing one orbit of the moon, effectively they used a long exposure and they were able to build up images over the top of each other of subsequent orbits. So, when they were all piled on top of each other and you capture it at this one particular point in space and time, you were able to build up a much better image of this little tiny moon.
Interviewer: Benjamin Thompson
And what do we know about this little tiny moon?
Interviewee: Lizzie Gibney
It’s the smallest of Neptune’s moons and it sits just inside the orbit of Neptune’s second largest moon, Proteus. The researchers of this paper suggest that it might have formed through ejected fragments of some kind of bigger, previous moon perhaps, after a big comet impact. So, this all adds support to the idea that it was perhaps quite a turbulent environment around Neptune back in the early days.
Interviewer: Benjamin Thompson
Well, last question from me, Lizzie, and it’s about the name, Hippocamp. What can you tell me about this one?
Interviewee: Lizzie Gibney
Yes, so of course, Neptune was named after the Roman god of the sea, as you mentioned, and so the moons are all kind of lesser sea gods and nymphs in Greek mythology. So, Hippocamp was one of those. Now, people might be confused because they’ve probably heard of the hippocampus which – and this is something I hadn’t realised and I looked up – but the hippocampus is called that because that part of the brain resembles a seahorse, so they kind of tie together.
Interviewer: Benjamin Thompson
The things I learn doing the News Chat. Well, listeners, I hope you’ve learned something too, and for more on these stories head over to nature.com/news.
Host: Shamini Bundell
That’s it for this week’s show. In the meantime, if you need more science for your ears and eyes, head over to our YouTube channel for a mini documentary on some multi-coloured mutant ants. That’s at youtube.com/NatureVideoChannel. I’m Shamini Bundell.
Interviewer: Benjamin Thompson
And I’m Benjamin Thompson. Thanks for listening.

 Classify viruses — the gain is worth the pain
Classify viruses — the gain is worth the pain
 Opportunity lost: NASA says goodbye to pioneering Mars rover
Opportunity lost: NASA says goodbye to pioneering Mars rover
 Read the paper: The seventh inner moon of Neptune
Read the paper: The seventh inner moon of Neptune