- NEWS AND VIEWS
Killer T cells show their kinder side
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 51 print issues and online access
$199.00 per year
only $3.90 per issue
Rent or buy this article
Prices vary by article type
from$1.95
to$39.95
Prices may be subject to local taxes which are calculated during checkout
Nature 555, 594-595 (2018)
doi: https://doi.org/10.1038/d41586-018-03510-z
References
Linehan, J. L. et al. Cell 172, 784–796 (2018).
Sabaté Brescó, M. et al. Front. Microbiol. 8, 1401 (2017).
Naik, S. et al. Nature 520, 104–108 (2015).
Wang, C.-R. et al. Cell 82, 655–664 (1995).
Legoux, F., Salou, M. & Lantz, O. Annu. Rev. Cell Dev. Biol. 33, 511–535 (2017).
Van Rhijn, I. et al. Nature Immunol. 14, 706–713 (2013).
Godfrey, D. I., Uldrich, A. P., McCluskey, J., Rossjohn, J. & Moody, D. B. Nature Immunol. 16, 1114–1123 (2015).
Rodgers, J. R. & Cook, R. G. Nature Rev. Immunol. 5, 459–471 (2005).
Goyos, A., Sowa, J., Ohta, Y. & Robert, J. J. Immunol. 186, 372–381 (2011).

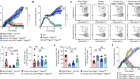
 The patterns of T-cell target recognition
The patterns of T-cell target recognition
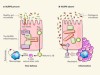 In command of commensals
In command of commensals