Abstract
In order to study the functional structure of the transcription terminators and the mechanism of termination, a survey of the chromatin structure, including the location of DNase I hypersensitive sites and the nucleosome arrangement, of yeast ADHI and FLP terminators was made. The results show that there is no relationship between the function of the terminators and the existence of DNase I hypersensitive sites. However, it is found that there is always a nucleosome at the immediate upstream of the transcriptional termination sites. As a control, the chromatin structures of the pBR322 DNA fragments on the yeast shutter vectors are also investigated at the same time. The random nucleosome arrangement on lhe bacterial DNA in yeast agrees with the published reports. A new hypothesis, about the mechanism of transcriptional termination is put forward and the reason of different nucleosome arrangement on the DNAs which are originally from different species in yeast is discussed.
Similar content being viewed by others
Introduction
In-the field of eukaryotie gone expression and regulation, the mechanism of the yeasb transeription termination, as thai of higher organism, has not been clearly understood yet, although 3′ flanking regions of yeast genes have been considered as “terminators” in genetic engineering. The transoriptional termination with RNA polymerase II is considered generally to be coupled with the poly(A)proeessing of branseripts in yeast 1, 2, which is different from that in animals. It is also known that the 3′ flanking region of the years gene is always A/T-rich2, 3 by DNA sequencing. But the functional strueSure of yeast Sranseription terminabors is still open for further investigation.
A few hypotheses about the functional struelure of years terminalors have been put forward. By comparing DNA sequences using computer, Zaret and Sherman 2 found such a consensus sequences as: STOP 00DON ... 1∼l40bp ... (T-rich)...TAG...TA(T)GT...TTT...(Poly(A)site) in 3′ regions of 15 yeast mRNAs, and Shis was considered as a feature of yeast terminators. Bennetzen and Hall3 found thah the consensus sequence of TAAATAAA/G upsSream of 11. yeast transcription termination sites might be functionally similar to AATAAA polyadenylation signal in animal cells. From the results of deletion experimen∼ of fruit fly ADE–8 gene integrated into yeast chromosome, Henikoff 1, 4 concluded that TTTTTATA is the controlling signal of transeripbional termination, and moss of the termination sites were in the CAAT/GTTTG sequences. But all of the above results do not conform with each other, and none of them fits to all yeast terminators. The consensus sequence TAG–AT(T)GT–TTT in Zaret–Sherman's model, which conforms with mosb terminators and appears in more Shan 90% of the 3′ region of yeast transcripts, might also not reflect the true features of yeast terminalors because those with one–bp–mismatohed TATGT and thore witshout TTT triple in many cases were also considered as consensus sequences 2. Considering the feature of yeast terminators which usually contain 70—75% of A/T bases, it gets very high probability to randomly show Zaret–Sherman consensus sequence(calculation not shown).
More recently, it is found Shat there are two transcripSion termination sites in the A/T–rich region of She repeat sequences of yeast bransposon Ty–D15 5. The efficiency of termination on these two sites decrease by inserting a G/C–rich fragment downstream the termination sites5. It means that both transeripiion termination sises, and the downstream fragment play a role in the termination reaction. This is a new discovery and has not yet been noticed in any of the proposed models.
Therefore, we have every reason So believe that there is bulk of work waiting for further investigation on the functional s brucSure of terminators and the mechanism of yeast transcription termination.
The research of the interactions between the proteins and DNA has been developed very rapidly in these years. It is known that one DNA fragment can be recognized by two different proteins6, and the same protein may be able to bind two DNA fragments different in DNA sequences7. Is :it possible for yeast, terminators different in sequences be work in a resemble way? We suspect.
Years ARS region may be the first example in which 3–dimensional sructure instead of DNA sequence plays its role in function8. Moreover, it has boon found by Szent–Gyorgyi et al that there is specific chromatin structure at the transcription termination sites of yeast, HSP82 gene. They suspeeted that there might, be certain relationships between the chromatin structure and branseription termination9. So that the terminators for yeast RNA polymerase II transcription may stop the prolongation in a special 3–dimensional struebure, and this strueture may be reflected on its chromatin features.
Dozens of articles on yeast chromatin structures of more than 20 DNA fragments have been published. A few rules and characteristics of yeast chromatins has be summarized10. In this paper, we get some data relevant to the chromatin strueture of the yeast terminators. We also checked Tthe loeation of DNase I hypersensitive sites and nucleosome arrangement on the chromatin strueture of the bacterial DNA fragments of the same plasmids. And the reasons for ordered arrangement of yeast nucceosomes are discussed.
Materials and Methods
1. Strains and media
Bacterium
E. coli HB 101 (recA 13 SupE44 (su2+) lacZ4 leuB6 proA2 thi-1 (B1) F-gal- r-m-)
Medium
LB(2% peptone, 1% Yeast Extract, 1% NaCl)
Yeast
S. cerevisiae Eal-103(a leu2 ura3 trpl)
Medium
YEPD(2% peptone, 1% Yeast Extract, 2% glucose)
SD(0.67% Yeast Nitrogen Base w/o amino acid, DIFCO, 2% glucose, with 40μg/ml appropriate amino acids and nucleotflles)
2. Enzymes and plasmids
All enzymes are from Biolabs, Sigma or Sine-American Biotechnology Co. All of the plasmids are constructed in our laboratory except pWR33-hCC, which is a gift of Li-he Guo of our Institute.
3. Recombinant D NA technique 11, transformation of bacterium 11 and yeast 12, nick translation and Southern hybridization 11 are done according to the references.
4. Extraction of yeast total RNA and Northern hybridization:
Total nucleic acid axe isolated as described by Elder et al. 13, electrophoresis is run in TA buffer (40 mM Tris-HCl/2 mM EDTA, pH8.0)on denaturing gel containing formalaehyde. Northern hybridization is done as Southern hybridization. For control, after RNAs are run on the den'Ituring gel, the gel is treated with 0.5 NNaOH/I.5 N NaCl for 1 hour, and neutralized in IM Tris-HCl, pH7.5, /1.5 N NaCl for 1 hour.
5. Digestion of yeast chromatin and naked DNA with micrococcal nuclease( MNase)
Cell pellet from 200 ml SD culture (1—1.50D 600)is washed in 10 ml 1M sorbitol/0.04 mg/ml Zymolyase 20T, and incubated at 37°C for 1 hour. Spheroplasts are washed with 1M sorbitol, and suspended in 3—4ml 5 mM KH2PO4, pH 7.5/0.1 mM CaC12/1 mM PMSF by Vigorous vortex. Suspension of nuclei is divided into 0.5 ml portions, and digested with 0—3.2mg gradient amount of MNase at 37°C for 10rain. The reactions are stopped by adding 50μl 0.5M EDTA, pH 8.0. The digested nuclei are broken with 50μl 10% SDS. DNA is purified and dissolved in 20μl TE(10mM Tris–HCl, pH 8.0/lmM EDTA). For naked DNA, (in an amount equivalent to one electrophoretic sample of chromatin DNA), 0.01–0.04 mg gradient amount of MNase is used in 25μl 5 mM KH2PO4, pH7.5/10 mM CaC12/1 mM PMSF at 37°C for 10rain, Then both chromatin DNA and naked DNA are treated with appropriate restriction endonuclease and electrophoresis is run on 1.2% agarose gel.
6. Digestion of yeast chromatin and naked DNA with DNase I
Spheroplasts in amount identical to that used in MNase digestion are suspended in 3–4 ml 10 mM Hepes, pH8.0/0.5mM MgCl2/0.1 mM GaCl2, divided into 0.5 ml portions, and digested with 0–0.6μg gradient amount of DNase I at 37∼C for :[0min, DNA was purified in 20pl TE. For naked DNA, (in an amount equal to one ele3trophoretic sample of chromatin DNA)0.1--0.4 ng gradient amount of DNase I in 10 m∼f Hepes, pH 8.0/10 mM MgCl2/10 mM CaC12 is used.
Result
1. Construction of plasmid models and its gene expression
As a model for investigation of chromatin structures of the yeast ADH1 and FLP terminators, a series of veetors are construeted to express the cDNA of the β–subunit of human chorionic gonadotropin (β–hCG) controlled by yeast ADH1 promoSer. The two Serminators are put in different plasmids in order to eliminate the effect of flanking strueture (boundaries) and chromatin folding (plasmid supercoilling and size)on the chromaSin structure of the Serminators.



Fig. 1 shows the construelion of the plasmids. ADH1 and FLP terminators are separated by a 350 bp HindIII–BamHI fragment from pBR322 in pY44. pY42 has similar structure as pY44 exeept that the β– hCG cDNA is insorted in opposite direotion, pYAI8 is a derivative of HindIII–parSially digested pY44. ADH1 terminator and HindIII–Bam HI fragment from pBR322 are deleted in pYA18. In pV3, ADHI promotor, 5′ portion of the β-hCG cDNA and BamHI-PvuII 170bp pBR322 fragment are doloted. The parts relevant to chromatin struoture assay of terminalors are shown in Fig, 2.
The terminators studied in this paper. (for legend, see Fig. 1)
The numerals under each diagram are the length of DNA fragments in Kb.
All of the four plasmids are transformed into yeast strain Eal–103. The transcription of She β–hCG cDNA in these plasmids is chocked. The result is shown in Fig. 3. Two RNA bands from the totol nucleic acids of She pY44 and pY42 Sransformants can be hybridized wish β–hCG cDNA, corresponding to the two transeripiion terminators in tandem. It shows that the upstream ADII1 terminator can not stop the elongation of the transerlption eomplotely, and some transcrlpts get over the 350 bp BamHI–HindIII pBR322 fragment and stop at the second terminator. But the dissineS difference betweon She inSonsiiy of She RNA bands in pY44 and pY42 transformants does not mean that the ADHI torminator in pY42 is more effieient then that in pY44.
As we can see from Fig. 4, the efficiency of the ADH1 torminators in pY44 transformed Eal-103 clones differs among themselves. The reason for this is not clear yet, but it is ovidont that the ADH1 terminator can work in bosh pY42 and pY44 plasmids. Nevertheless the efficiency of a termlnator in different plasmlds or different hosts, and even in different transformants with same plasmids in same host may be affected by many factors which have not been understood yet.
There is no yeast promoter in the same direction with ADH1 and FLP terminators in pV3, but the promoter of the Apr gone from pBR322 can control the gone transcription in yeast (unpublished data). Fig 3 shows that the Apr promoter does initiate the transcription of β–hCG cDNA in pV3. The transcripts from pYA18 also conform with its plasmid structure.
2. Chromatin structure of yeast transcription terminators
The main methods used to study the chromatin structure of the terminators in this paper are checking the arrangement of DNase I hypersensitive sites (HSs) and location of the nucleosomes on terminator region by indirect-end labeling. The chromatin structures of the terminator regions in all of the plasmids studied are shown in Fig. 5. There are at least three or four DNase I HSs on the region nearby the terminators in pY44(Fig. 5Ia), but only one of them is located on the 0.45Kb ADH1 terminator fragment. One of the others is on the repeated sequence of 2μ plasmid, close to the XbaI site. There is no DNase I HS on the FLP terminator region, pY42(Fig. 5IIa)has almost the same structure as pY44 except that the β–hCG cDNA insort is inverted, and the position of DNase I HSs on BamHI-HindIII pBR322 fragment changes while the one on 2μ–fragment near the XbaI site does not. This is caused by the changes of plasmid structure duo to inversion of β–hCH coding region. In pYA18(Fig. 5IIIa), ADH1 terminator and part of the pBR322 DNA are deleted, so that FLP terminator is closer to the probe, and its supereoil structure as well as plasmid size should be different from pY44 and pY42. No evident alternation in DNase I HSs position is found on FLP torminator of pYA18. In pV3(Fig. 5IVa), the DNase I HSs on ADPI1 terminator region as well as those on BamHI–piindIII pBR322 fragment are shifted for dozens of hase pairs, although the one near the XbaI site remains there.
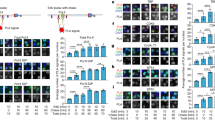
Chromatin struciure of ADH1 and FLP terminators. 32P–β–hCG cDNA SmaI-PstI fragment is used as probe. The length of each band indicates the distance between the DNase I/MNase cutting sits and the end of the probe, PstI site.
5–I. pY44/PstI 5–II. pY42/SmaI. 5–III pYAIS/PstI. 5–IV. pVS/PvuII.
a, DNA/DNaseI. b, DNA/MNase.
A, λDNA/PstI molecular weight standard. B. Naked DNA/DNaseI. C. Chromatin/DNaseI.
D. Naked DINA/MNase. E. Chromatin/MNase. F. Intact DNA as a control.
The spectra of MNase cutting sites in Fig. 5b shows that those cutting sites, 160 bp upstream of XbaI site on FLP terminator and 170 bp downstream of the β–hCG cDNA, are present in all of the four plasmids. In pY44 (Fig. 5Ib), there are three MNase cutting sites on 3′half of the ADH1 terminator fragment with short spaces of only 40–50 bps. This fact implies that there is no nucleosome on this region The cases in pY42 and pV3 are similar to that of pY44 but the spaces of the successive MNase sites seem a little longer. The rest of MNase sites are arranged with at least 140 bp space from each other, although their position may move depending on plasmid size and supercoiling. This phenomenon illustrates that in contrast to the random nucleosome arrangement in animals, the nucleosome arrangement on yeast DNA is sequence–dependent, and may be affected by DNA supercoiling.
The results of all of the chromatin structure assay of yeast ADH1 and FLP terminators are summarized in Fig. 6. There are two nueleosimes, one at 160 bp upstream of XbaI site of FLP terminator and the other at 170 bp downstream of HindIII–end of the ADHI terminator, locatod at the same position in all of the four plasmids studied. The rest of nucleosomes may shift with the plasmid structure, although their arrangements are also sequence–dependent.
Summary of chromatin structure assay of the transcription terminators.
▿, DNase I hypersensitive sites. ▾, MNase digestion sites. ○, Nucleosome.
For the rest of the legends, see Fig. 2.
3. Chromatin structure in yeast of pER322 DNA fragments on the plasmids we constructed
We studied the chromatin structures of Apr gene, replication origin and part of the Tor gone from pBR322. This is used as a control to check whother we are right in correlating funolional terminators to their chromatin struoiures. Fig. 7 shows that the background intensity of chromatin /MNase digestion pattern is heavier than that of naked DNA/MNase digestion pattern; the DNase I sensitivity of bacterial DNA chromatin is similar to that of yeast chromatins; the distances between the MNase bands are not regular and do not correspond to the length of yeast nueleosomal repoats, we come to the conclusion that these pBR322 DNA fragments are packaged into nucleosomes to form genuine ehromatin in yeast. However, the nueleosomes are randomly arranged. The eonoluslon agrees with those from obher reports.
Chromatin structure of pBR322 DNA fragment of plasmlds in yeast cell.
7—1, pY44/BamHI. 7—2, pY42/BamHI. 7—3, pYA18/BamH. 7—4, pV3/SmaI 32P–pBR322/BamHI–SalI 0.3Kb fragment as probe.
A, B, C, D, E and F, see the legend of Fig. 5.
Discussion
The ADH1 promoter-controlled B-hCG cDNA expression sys6em in yeast is chosen to study the chromatin structures of 6he ADH1 and FLP 6ermina6ors. The ADttl promoter is very strong 6o start transoription. Almost 1—2% of cellular poly(A)mRNA is the transcrfpts of ADH1 gene14 in wild type yeast cells, although its effioiency to express foreign genes varies. With this model, efficient transcription promoted from ADH1 promoter enables us to investigate the terminators in their functional state. Moreover, it had been noticed that the promoter is important for the RNA polymerase II to choose appropriate terminator structure to stop transcription. The transcripts from snRNA gene promoter can not be processed with poly(A)signal in animal cells. Those of pV3 promoted at 5′ region of bacterial Apr gene can be stopped at yeast terminators, as shown in Fig. 3. It means that the bacterial promoter region does not change the structure of RNA polymerase II which may be essential for recognizing the funetional strueture of the terminabors.
1. Common features of different yeast terminators without sequence homology
The DNA sequence analyses of both yeast ADH1 3 and FLP 15 terminators have been completed, and their RNA mapping has also been reported 3, 16. Sequence comparison shows that FLP terminator is of similar elements with Zaret-Sherman consensus sequences 16, however, ADHI terminator is of TAAATAAA/G homology sequence 3 instead of Zaret–Sherman Model 2. If there are two or more mechanisms for RNA polymerase II transcriptional termination in yeast, ADIII and FLP terminators may be their representatives. Pubting the twe terminators into different plasmid environments to survey their chromatin structure, we may get a more complete picture about the three–dimensional structure of yeast transcriptional tcrminators.
According to Bennetzen and Hall 3, the terminus of the ADH1 gene transcript is located on three clusters of sites scattered from 230 to 290 bp upstream of the BamHI site, or in another words, the fronb of the region is at about 160 bp downsbream of the HindIII site. Our results show that there is a nucleosome positioned at just 160 bp downstream of the HindIII site on ADH1 terminators on all three plasmids studied(pY44, pY42, PV3), suggesting that the edge of the nucleesome is just at the front of the termination region, and the termination sites are located at the linker D NA between nueleosomes. The case in the FLP 4erminator is similar. The transcripiion terrninus of the FLP gene is mapped al about ]50 bp ups4ream of the XbaI site16, where a MNase cutting site is present. This indicates that a nucleosome is positioned at the immediately upstream of this point. The relationship between bhe nucloosome position and transeriptional termination site is diagramed in Fig. 8.
In all of the published reports about the nueleosome position of the 3′ region of yeast genes, the MNase sites spectrum of TDH3 gone 17 is not clear enough to be compared, but in PHO318, SUC219, and HSP829, there is a nuoleosome positioned at the immediately upstream of each transcription termination stbe (or sites cluster), followed by a short region free of nucleosome. In SUC2 20, the fact that the tight binding of nuoleosomes in coding region becomes relaxed in the downstream region of the transcriptional termination site, implies that there is a special structure at that site. In HSP82 geno10, the semi–nucleosome in coding region changes inio complete nucleosome al transcription termination site. Only the URA3 gene is an exception. Thoma found that both TATA box and transcriptional termination site of the URA3 are in the nucleosomal DNA 21. TRP1 gone 22 is a special example. There is no nucleosome positioned at immediately upstream of the terminalion site. However, the DNaseI hypersensitive region of ARS1 adjacent to the end of TRP1 gone may have boundary effect 23 on the nucleosome to be shifted further upstream.
On the basis of these reports and our results, the arrangement of nucleosomes on yeast transcription terminators may be importani for terminator function. Termination may occur on the linker DNA immediately downstream of a nucleosomo. The A/T–rich region, TAG, TA(T)GT elements and TAAAAAA/G 2 may have similar funolion in nucleosomo positioning. Insertion of G/C rich fragment into the region downstream of transcripiion termination site 5 may affect the affinity of adjacent DNA fragment to nucleosome probeins, and may change the nucleosome position and formation of the funcbional structure of terminators, and as a result, may decrease the efficiency of termination.
We fail to relate the DNaso I HSs to the function of terminators. There is no DNase I HSs in the FLP terminator, and that one near the Xba I site may be. caused 24 by Pu–Py altornative region of the inverted repeats 15. in ADIIL terminator, the closest DNase I HSs to the termination sites is at 90—160 bp downstream of the termination sties.
2. Nucleosome position and the sequence specificity in MNase digestion
After the MNase digestion, there are many bands of yeast chromatin corresponding to that of naked DNA, although some bands of naked DNA do not appear on chromatin due to the protection by nuoleosomo. The correspondence of MNase cutting sites between yeast chromatins and naked DNAs can be found in many other reports(for example, 25). It may be relevant to the sequence specificity of yeast nucleosome position. For a long time .it has been known thai the nucleo–some packaging requires some sequence features. Those structuros like Z–DNA 26, DNA cruciform 27and long stretch of poly (dA)–po]y (dT) 28 can not be packaged into nucleosome. On the other hand, yeast nucleosome arrangemont is ordered and sequence–specific. So that those DNA elements easily attacked by MNase may not be suitable to be packaged into nuoleosome. Instead, they may prefer to work as linker DNA.
3. Random arrangement of nucleosomes on bacterial DNA in yeast
It has been proved by nucleosome reconstruction it vitro that the nucleosomes formed by pBR322 from E. coli and the histones from higher organisms are randomly arranged 29, 30. The arrangement of nueleosomes in higher organisms is also basically random 32. We pay high attention to that the orderly arranged yeast nucleosomes losa bhoir positioning rule on pBR322 DNA fragments. Similar results have been reported by others33, 34. There may be a kind of special information, existing on DNA sequence, involved in nucloosome positioning and gone regulation in yeast. Lack of bhose informations in bacterial DNAs make thom unable to position nueleosome orderly in yeast cells. Consoquently, we believe the reason for the random nucleosome arrangement on pBR322 DNA in yeast is due io its original source of DNA from different species.
Recently, it is found that there are special chromatin struetures on the terminator region of the mouse k immunoglobulin gone different from the active chromatin structure pattern on the rest of the gene locus 35, implying that there may be some relationship between terminator and its chromatin structure even in animal cells. According to Resnokov et al. 36, transcription terminators in eukaryotic cells may be classified into three models. The terminators of yeast cells can be grouped wish one of the models in which it is supposed that terminators is a specific sequence with no apparent secondary structure but with speaJfic read through or terminalion factors. In She ]aek of definite sequence mark of terminators and knowledge of termination factors, it is worth studying the chromatin structure of the terminators. Our results show that the nucleosome immediately upstream of the termination site may be involved in the formation of the functional structure of terminalor. Future investigation on the isolation of possible termination factors on the key nucleosomes may be of high interest.
References
Henikoff S, killy JD, Cohen EH . Transcription terminates in yeast distal to a control sequence. Cell 1983; 33: 607–614.
Zaret KS, Sherman F . DNA sequence required for efficient transcription termination in yeast. Cell 1982; 28: 563–573.
Bennetzen JL, Hall BD . The primary structure of the Saccharomyces cerevisiae gene for alcohol dehydrogenase I. J Biol Chem 1982; 257: 563–573.
Henikoff S, Cohen E H . Sequences responsible for transcription tereimation on a gene segment in Saccharomyces ceaeviciae. Mol Cell Biol 1984; 4: 1515–1520.
Yu K, Elder RT . Some of the signals for 3′–end formation in transcription of the Saccharomyces cerevisiae Ty-D15 element are immediately downstream of the initiation site. Mol Cell Biol 1989; 9: 2413–2444.
Pfeifer K, Arcangioli B, Guareate L . Yeast HAP1 activator competes with the factor R02 for binding to the upstream activation site UAS1 of the CYC1 gene. Cell 1987; 49: 9–18.
Pfeifer K, Prezant T, Guazcnta L . Yeast HAP1 activator bind to two upstream activation sites of different sequence. Cell 1987; 49: 18–27.
Eoaton AH, Smith MM, Fine structure analysis of the DNA sequence requirements for autonomous replication of Sbccharomyves cerevisiae plasmids. Mol Cell Biol 1986; 6: 2354–2363.
Szent–Gyorgyi C, Finkellstein DB, Garrard WT . Sharp boundaries demarcate the chromatin structure of a yeast heat-shock gene. J Mol Biol 1987; 193: 71–80.
Gross DS, Szent-Gyorgyi C, Garrard WT . Yeast as a model system to dissect the relationship between chromatin structure and gene expression. UCLA Sump MIol Cell Biol 1986; 33" 345–366.
Relevant chapters, IN Mantiatis T., Fritsch EF, Sambrook J. eds. Molecular Cloning. Celd Spring Harbor Laboratory. New York 1982.
Hinnea A, Hicks JB, Fink GR . Transformation of yeast. Proc Natl Acad Sci USA 1987; 75: 1922–1933.
Elder RT, Loh EY, Davis RW . RNA from the yeast transposable element Tyl has both ends in the dilect repeats, a structure similar to retrovirus RNA. Proc Natl Acad USA 1983; 80: 2432–2436.
Danis CL, Ferhuson J, Young ET . mRNA levels for the fermentative alcohol dehydrogenase of Sacchaaomyces cerevisiae. Decrease upon growth on a nonfermentable carbon source. J Biol Chem 1983; 253: 1165–1171.
Hartley JL, Donelson JE . Nucleotide sequence of the yeast plasmid. Nature 1980; 386: 860–865.
Sutton A, Broach JR . Signals for transcription initiation and termination in the Sacoharmyecs cerexsiae plasmid 2μm circle. Mol Cell Biol 1985; 5: 2770–2780.
Pavlovic V, Horz W . The chromatin structure at the promoter of a glyceraldehyde phosphate dehydrogenase gene from Saccharomyccs cerevisiae reflects its functional state. Mol Cell Biol 1988; 8: 5513–5520.
Almer A, Horz W . Nuc]ease hypersensitive regions with adjacent positioned nucleosomes mark the gene boundaries of the PH05/PH03 locus in yeast. EMBO J 1986; 5: 2681–2687.
Perez-Ortin JE, Estruch F, Matallana E, ∼ranco L . Fine analysis of the chromatin structure of tke yeast SUC2 gene and of its changes upon depression. Comparison between the chromosomal and plasmid-inserted genes. Nucleic Acids l∼es 1987; 15: 6937–6954.
Perez-Ortin JE, Estruch F, Matallan∼ E, Franco T,. DNase I sensitivity of the yeast SUC2 gene for invertase. Mol Gen Genet 1986; 205: 422–∼27.
Thoma F . Protein-DNA interactions and nuclease-sensitive region determine nucleosome positions on yeast plasmid chromatin. J Mol Biol 1986; 190: 177–190.
Lohr D, Torchia T . Structure of the chromosomal copy of yeast ARS1. Biochem 1988; 27: 3961–3965.
Thoma F, Zatchei M . Chromatin folding modulates nucleesome positioning in yeast minichromosomes. Cell 1988; 55: 945–953.
Fagrelius T J, Strand AD, Linvingston DM . Changes in the DNAase I sensitivity of DNA sequenes within the yeast 2μm plasmid nucleoprotein complex effected by plasmid-encoded products. J Moi Biol 1987; 197: 415–423.
Klnezetic JA, Luse DS . The presence of nucleosomes on a DNA Lemplate presents initiation by RNA polymerase II in vitro. Cell 1986; 45: 95–1C4.
Garner MM, Felsenfeld G . Effects of Z-DINA on nucleosome placement. J Mol Biol 1987; 196: 581–590.
Nobile C, Nickol J, Martin, RG . Nucleosome phasing in a DNA fragment from the replication origin of silan virus 40 and rephrasing upon cruciform formation of the DNA. Mol Cell Biol 1936; 6: 2916–2922.
Kunkel GR, Martinson HG . Nucleosomes will not form on double-stranded RNA or over poly(dA)- poly(dT) tracts in recombinant DNA. Nucleic Acids Res 1981; 9: 6869–6888.
Caffarelli E, De Santis P, Leoni L, Palleschi A, Savino M . Preferential positioning of nucleosomes on pBR322 as evaluatel via fourier transform of data from electron microscopy. Eur J Biochem 1988; 171: 497–501.
Chao MV, Garlla J, Martinson HG . DNA sequence directs placement of histone cores on restriction fragments during nucleosome formation. Biochem 1979; 18: 1068–1074.
McNamara PT, Winecov I, Harrington RE . Preferential nucleosome placement of pBR322 restriction fragments. Biochem Biephys Res Comraun 1986; 138: 110–117.
Eissenberg JC . Selected topics in chromatin structure. Ann Rev Genet 1985; 19: 485–536.
Bloom KS, Carbon J . Yeast centromere DNA is in a unique and highly ordered structure in chromosomes and small circular minichromosomes. Cell 1982; 29: 305–317.
Estruch F, Perez-Crtin JE, Matallana E, Franco L . In vivo assembly of chromatin on pBR322 sequence cloned into yeast plasmids. Plasmid 1989; 21: 113–119.
Xu M, Barnard MB, Rose SM, Cockerill PIN, Huang SY, Garrard WT . Transcription termination and chromatin structure of the active immunoglobulkin k gene locus. J Biol Chem 1986; 261: 3838–3845.
Resnekov O, Ben-Asher E, Bengal E, Choder M, Hay N, Kessler M, de et. Transcription termiaation in animal viruses and cells. Gene 1988; 72: 91–104.
Author information
Authors and Affiliations
Additional information
*Project supported by National Science Foundation of China.
Rights and permissions
About this article
Cite this article
Hu, G., Yu, Y. & Kuang, D. Fine analysis of the chromatin structure of yeast RNA polymerase II transcription terminators. Cell Res 2, 67–81 (1992). https://doi.org/10.1038/cr.1992.7
Received:
Revised:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/cr.1992.7