Abstract
Genetic mapping techniques can be used to study the interaction between two different genomes after hybridization. This study investigated a Saccharum officinarum (Green German or GG, 2n≈11x≈110) × S. spontaneum (IND 81–146 or IND, 2n≈7x≈56) interspecific cross. Segregation of 193 microsatellite (SSR) loci was evaluated in the F1 progeny of 169 full-sibs of the cross. Following the two-way pseudo-testcross strategy and ‘cross pollination’ population type, linkage groups (LG) and phases were established for each parent map, using the criteria of LOD score ≥3.0 and a maximum recombination frequency of 0.35. Of the 193 markers analyzed, 61 were IND-specific, 106 were GG-specific, and 26 were heterozygous in both parents. About 78% of the markers segregated in a Mendelian fashion and 22% were distorted (as evaluated by χ2-tests, P≤0.01). The GG map included 91 marker loci arranged into 25 LG covering 1180 cM of the officinarum genome. The IND map consisted of 46 marker loci assembled into 10 LG, which spanned 614 cM of the spontaneum genome. A specific chromosome associated with segregation distortion was detected in the female (GG) genome only, probably as a result of double reduction. The segregation patterns of the marker loci indicated a centromere-driven distortion process with the shared allelic markers (as putative centromeres) regulating the placement and association of markers with opposite phase (coupling vs repulsion) and dosage on either side. Although incomplete, the framework maps were informative with respect to segregation distortion, chromosome fusion, rearrangements, and translocations, observed in both parental genomes as a result of their merger.
Similar content being viewed by others
Introduction
The world sugarcane (Saccharum sp.) breeding populations are founded on a very narrow genetic stock (particularly S. officinarum and S. spontaneum clones) and were developed by crossing within the progeny of interspecific crosses made in the early 1900 (Berding and Roach, 1987). Commercial sugarcane (2n=100–130) cultivars are complex alloploids or interspecific hybrids arising originally from the hybridization of multiple genomes (presumably S. officinarum, S. spontaneum, S. robustum, S. barberi, and/or S. sinense, with S. officinarum as the predominant cytoplasm), and as a consequence are highly heterozygous. There is still great interest among sugarcane breeders in broadening the genetic base of the crop and also in taping into the gene pool of the wild relatives to improve stress-resistance and sucrose content (Tai and Miller, 2001, 2002). Breeding gains in sugarcane, even when substantial (Edmé et al, 2005b), have been slow in recent years, possibly as a result of a founder and/or genetic bottleneck effect.
Classical genetics has been unreliable at ascertaining the introgression of beneficial alleles from the wild into the cultivated background and at eliminating linkage drag. Molecular approaches have improved the tracking of species-specific alleles in interspecific hybrid backgrounds and the investigation of colinearity and recombination of chromosomal segments between the parents. Recombination is crucial in the transfer of genes/alleles from wild species into the cultivated background and for this strategy to have an impact in plant breeding. Grivet et al (1996) confirmed that recombination is possible in interspecific sugarcane crosses.
Previous genetic maps for sugarcane have increased our knowledge of the organization of its genome with emphasis put on the number of cosegregating groups and pairing behavior of the Saccharum genomes via determination of linkage phases (da Silva et al, 1993, 1995; Ming et al, 1998; Aitken et al, 2005). However, these maps have made almost exclusive use of nondistorted SDM or DDM markers (da Silva et al, 1993; Ming et al, 1998). Grivet et al (1996) did take distortion into consideration in mapping using the progeny of a selfed cultivar. The possibility of meiotic irregularities (chromosome loss, translocations, rearrangements, double reduction, and distortion) was not considered, and it is well known that irregularities are common occurrences in intra- or interspecific crosses (Jenczewski et al, 1997) and can be related to specific generations (Paterson et al, 1991).
The present study used the Green German (GG) × IND 81–146 (IND) interspecific (GI) population (developed by Dr Peter Tai, USDA-ARS Canal Point, FL, USA) that combines two very different genomes (IND as a spontaneum and GG as an officinarum) in their DNA content (3.1 vs 6.7 pg), genome size (1474 vs 3239 Mbp), mean chromosome number (2n≈56 vs 2n≈110), and in their phenotype with GG possessing the larger values (Burner, 1997; Ming et al, 2002a, 2002b; Edmé et al, 2005a). The objectives of this study were to follow the segregation of SSR, alleles in the F1 interspecific GI population of full-sibs to investigate the structure of the Saccharum genome with respect to arrangement of loci and segregation distortion, and to construct framework genetic maps to begin identifying molecular markers linked to traits of interest for sugarcane.
Materials and methods
Mapping population
An F1 sugarcane population was developed and later augmented via a biparental cross involving two highly heterozygous parents. Stored pollen of the S. spontaneum clone (IND, 2n≈7x≈56) was used to cross an S. officinarum clone (GG, 2n≈11x≈110) as the seed parent under controlled conditions. With a chromosome number greater than 80, the version of GG (2n≈97–117) used in this study and others (Burner, 1997; Edmé et al, 2005a; Ming et al, 1998, 2002a, 2002b) is not a true officinarum clone and therefore can be considered a hybrid of unknown origin. The progeny population was propagated in a nursery from seedlings in 2003 and immortalized in 2004 from vegetative cuttings, along with the parents, in a field experiment planted as an incomplete block design, with three replicates, at the Canal Point Sugarcane Field Station, Florida.
DNA isolation and microsatellite screening
A random subsample of 169 full-sibs was screened with a panel of sugarcane-based genomic SSR primers developed by the International Consortium of Sugarcane Biotechnology (ICSB) (Cordeiro et al, 2000). Approximately, 100 mg of the inner soft tissues of the leaf whorl of young plantlets were used for DNA extraction. DNA extraction was performed using the Qiagen DNeasy 96 Plant Kit (Qiagen, Valencia, CA, USA) according to the manufacturer's recommendations. DNA was eluted twice using 100 μl of elution buffer to give a final volume of 200 μl. Concentrations of DNA samples were determined by spectrophotometry at 260 nm and the purity was calculated by the ratio of optical density read at 260 and 280 nm.
DNA samples of the parents were scored for polymorphisms and the resulting polymorphic primer pairs were used to identify, in the progeny population, SSR markers that were present in the two parents. The two parents and seven progeny were replicated in all amplifications as quality controls of the reactions. PCR reactions were performed in 10-μl reaction volumes containing 1 μl (25 ng) DNA template, 0.8 U Taq DNA polymerase, 1.5 mM MgCl2, 1 × PCR reaction buffer, 250 μM each dNTP, 250 μM each forward and reverse primer. Forward primers were labeled with 6-FAM or HEX fluorescent dye. Amplifications were carried out in a thermal cycler with the following profiles: an initial denaturation at 94°C for 3 min, followed by 35 cycles of 94°C for 30 s, 54°C for 30 s, and 72°C for 45 s. The reactions were terminated with a 5-min extension at 72°C. Amplified fragments were resolved by capillary electrophoresis using an ABI 3730 Genetic Analyzer (Applied Biosystems, Foster City, CA, USA). DNA standards were used in the separation and scanning of the fragments. Raw data were analyzed by autobinning on the parental fragments using GeneMapper software v3.7 (Applied Biosystems) and automatic allele calls were confirmed visually.
Linkage analysis and map construction
The computer software Joinmap 3.0 (van Ooijen and Voorrips, 2001) was used for linkage analysis of the different segregation types and for map construction. SSR bands that were polymorphic in the parents and observed in the progeny were converted to a matrix of two-letter genotype codes based on the configuration of the parents and segregation of the bands as indicated by the software manual. Two maps were created, one for each parent, using a LOD score of ≥3.0, a maximum for recombination set at 0.35, and the ‘cp’ population type option as for a two-way pseudo-testcross. The pseudo-testcross strategy has been developed for linkage mapping in highly heterozygous diploid or polyploid outcrossers using marker-specific segregation types, based on the fact that segregation results from meiotic recombination between the parents (Grattapaglia and Sederoff, 1994). The segregation types considered were ‘nnxnp’, ‘lmxll’, and ‘hkxhk’ as heterozygous in the male, female, and both parents, respectively, with the phases automatically assigned by the software (van Ooijen and Voorrips, 2001).
Simplex (SDM, 1:1), duplex (DDM, 3:1, 11:3, and 7:2), triplex (TDM, 13:1, 15:1, and 11:1), and quadruplex (QDM, 64:1 and 69:1) markers, for either autoploidy or alloploidy, were tested for goodness-of-fit to the above-mentioned Mendelian segregation ratios by χ2-tests (P≤0.01 determined after Bonferroni correction) before inclusion in the construction of the linkage maps (da Silva and Sobral, 1996; Liu et al, 1998). Bands shared by both parents were used as anchor loci or allelic bridges to identify homologous and homeologous linkage groups (LG) and to create a composite map. Distorted markers were also mapped to identify regions of the genome associated with such distortions. Phases (coupling and repulsion) of the marker loci were automatically detected by the software by means of the underlying ‘cp’ option that allows loci of different phases to be linked on the same chromosome. The Kosambi mapping function (Kosambi, 1944) was used to calculate and convert recombination fractions into map distances, subsequently reported in centimorgans (cM). Original SSR marker names were used, followed by the amplified fragment size to identify corresponding loci. Linkage groups were named after the individual parents as GG and IND followed by an arbitrary number in descending length size. Locus names along with their respective distances were keyed in Mapchart software v2.1 (Voorrips, 2002) for map drawing.
As Joinmap does not include an option for segregation distortion, the software Map Manager QTXb20 (Manly et al, 2001) was used to assess linkage among the markers and to estimate the effect of distortion on the length of the maps. The Kosambi mapping function was used under a ‘Search linkage criterion’ of P=0.001 based on an ‘intercross’ model. The ‘Distribute’ option was used to allocate some of the unlinked markers to the most probable linkage groups at the same P-level.
Results
Segregation analysis
Segregation data on 193 polymorphic SSR markers between the two parents were retained for linkage analysis. Of the 193 markers used for mapping, 151 showed Mendelian segregation (1:1, 3:1, 7:2, 11:1, or 15:1) based on χ2-tests (P≤0.05; Table 1). In total, 74 of these markers were heterozygous in the female parent, whereas 51 were heterozygous in the male parent and 26 were heterozygous in both parents. On average, 78% of the markers segregated in a Mendelian fashion and 22% deviated (ie were distorted) from the expected ratios considered. Overall, distortion was three times as high with the ‘lmxll’ (16.6% female) than with the ‘nnxnp’ (5.2% male) markers. The parental alleles were not equally disturbed (30 vs 16%) within their respective class and no distortion was observed with the ‘hkxhk’ marker types.
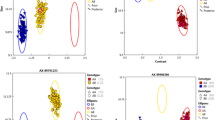
The distributions of the marker alleles were skewed toward the lowest ratios with the ratio class of 1 being the most typical for both the maternal and paternal markers (Figure 1). This implies that the majority of the markers could be considered to be single-dose markers (SDM). Higher dosages were predominant within the ‘hkxhk’ (shared-type) markers and also involved some of the specific parental alleles. However, a significant number of the <lmxll> (maternal) and ‘nnxnp’ (paternal) markers fell within the 0–0.25 ratio class, an indication that these markers were not being transmitted consistently and uniformly to the progeny.
Distribution of segregation ratios for 193 SSR markers used for mapping the F1 sugarcane population of 169 full-sibs and classified as being heterozygous in the female-GG (106 maternal), male-IND (61 paternal), or both (26 common) parents. Alleles with a segregation ratio >2.25 were considered to be of a dosage higher than that for single-dose markers (SDM).
Linkage analysis, segregation distortion, and map construction
Segregation data on 193 markers were used for linkage analysis of the F1 population, of which 106 and 61 markers were heterozygous in the female (GG) and male (IND) parents, respectively. The remaining 26 markers were heterozygous in both parents. In total, 137 marker loci were mapped on both genome maps, with 91 and 46 loci belonging to the female and male map, respectively. The loci on the GG map were distributed over 25 LG covering 1180 cM, for an average distance of 13 cM between markers or an average length of 47.2 cM per group (Figure 2). However, the marker loci were not well distributed across LG and gaps still remain. Linkage groups of the female map possessed 2–16 loci with GG1-1 and GG2-2 being the densest. These two LG were composed, each, almost exclusively of markers from a particular microsatellite, mSSCIR17 and mSSCIR32, respectively. Both these primers are repeats of the GA sequence, with mSSCIR32 having GG interspersed within. A total of 43 loci pertained to the 11 LG that covered 614 cM of the spontaneum male parent genome and that included from two to 11 loci, with an average distance of 13.3 cM between markers or an average length of 55.8 cM per group. IND1-1 also showed clustering of the same marker mSSCIR17 as did GG1-1 of the officinarum female map.
Partial microsatellite (SSR) genetic linkage maps of S. officinarum clone Green German (GG) and S. spontaneum clone IND 81-146 (IND) based on an F1 progeny population of 169 full-sibs. Linkage groups were established with LOD ≥3.0 and a recombination rate of 0.35. Genetic distances (cM) between loci, shown on the left of the chromosomes (open for GG and hatched for IND), were calculated via the Kosambi mapping function. SSR marker loci are shown on the right with their original names followed by the amplified fragment size. Shared marker loci are underlined and shown in bold typeface. Loci displaying distorted segregation ratios are indicated with asterisks (**), based on significant χ2-tests at P≤0.01. Repulsion-phase marker loci are indicated by the suffix ‘r’ attached to the loci. Multiple dosage markers are shown in italics. The first number after the name of the parent assigns homology groups within a parent, whereas it designates homoelogy groups between parents.
Linkage groups were arranged into putative cosegregating or homology groups (HG) for each parental map, based on microsatellite primers that have markers mapped on different regions of the genome or chromosomes. For GG, LG were assembled to give 11 putative HG that included 1–8 chromosomes. For IND, LG were ordered into six putative HG that included 1–3 chromosomes. The LG and HG were incomplete for both framework maps, given the expectations of 10 LG with 11 HG each for GG and eight LG with seven HG each for IND by assuming 2n≈11x≈110 for the former and 2n≈7x≈56 for the latter and normal meioses.
In all, 15 of the female LG showed preferential pairing of the marker loci. Various degrees of repulsion-coupling phase markers were mapped on these chromosomes, indicating that preferential pairing was predominant in GG. The other 12 chromosomes of the GG map included markers linked exclusively in the coupling phase. For IND, random pairing was more common with evidence of preferential pairing observed in only three of the 11 LG. The segregation patterns, chromosome pairing, and ratio of simplex to multiplex markers showed that GG behaved as an alloploid and IND as an autoploid (Tables 1 and 2).
Most of the distorted markers were unlinked and unmapped. Twice as many distorted marker loci and chromosomes were mapped on the female map (13 loci in six LG) than on the male map (six loci is three LG). No particular clustering was detected on IND chromosomes. Seven of these loci were clustered on GG2-2, which also included higher dosage markers linked in the coupling phase only. The other mapped distorted loci appeared to be randomly distributed across the two genomes in minor clusters or in between normal loci.
Differences in map lengths were observed between the Joinmap v3.0 and Map Manager QTXb20 linkage mapping programs. Although there was no significant difference in the length of the paternal map (614 vs 678 cM) between the two programs, there was a larger disagreement in the length of the maternal map (1180 vs 732 cM) when segregation distortion was accounted for in linkage analysis using Map Manager. The female map was reduced by 38%. To assess, without bias, the effect of distortion on linkage, Map Manager was used (on its own) with the ‘segregation distortion’ option turned on and off. Map Manager estimated a 111% increase in map length (1431 cM) for IND and a 156% increase in length (1880 cM) for GG (Figure 3).
Inflated microsatellite (SSR) genetic linkage maps of S. officinarum clone GG and S. spontaneum clone IND generated by Map Manager QTXb20 with the segregation distortion option turned off. Linkage groups (LG) were established at P=0.001 for an intercross model via the Kosambi mapping function and named as for the Joinmap map of Figure 2. LG that differ from the ones obtained with Map Manager by way of segregation distortion are reported only. Shared marker loci by the two parents are underlined and shown in bold typeface. Loci displaying distorted segregation ratios (χ2-tests at P≤0.01) are indicated with asterisks (**). The first number after the name of the parent assigns homology groups within a parent, whereas it designates homoelogy groups between parents.
Discussion
Two parental framework maps were built using microsatellite markers in conjunction with the pseudo-testcross strategy and the intercross ‘cp’ mapping population option in Joinmap. This allowed us to determine the phases of the markers and, at the same time, to map both coupling and repulsion-phase linked markers on the same linkage groups. The lengths of the maps were 1180 cM for GG and 614 cM for IND in this study, which, relative to the RFLP maps of Ming et al (1998), correspond to a 22.5 and 13% coverage, respectively. The number of SSR markers used for mapping the Saccharum genome was insufficient to cover the whole genome of the two species. Mapping in complex polyploids like sugarcane requires substantially more markers and progeny than it would for diploid genomes.
In spite of the low coverage, the configuration of the markers on both maps was revealing of the distribution and linkage of loci on all linkage groups (LG). Segregation of marker loci in the two genomes appeared to be driven by the DDM or ‘hkxhk’ markers playing the role of allelic bridges for linkages between higher (TDM and QDM) and lower dosage (SDM) markers and between markers of opposite linkage phases (coupling vs repulsion). Markers tended to cluster around the DDM-allelic bridge markers which may point to the locations of the centromeres. In this case, segregation and linkage of the marker loci would be centromere-driven, a characteristic feature that usually controls the segregation of gametes at meiosis (Simchen and Hugerat, 1993; Dawe, 1998). This is further corroborated by the fact that seven of these DDM markers remain unlinked and unmapped. This observation is new in sugarcane mapping as Ming et al (2002a) reported that most of the published sugarcane maps are deficient in the regions associated with the centromeres.
Considering these allelic bridges as centromeres, meiotic irregularities were evidenced by the presence of putative multicentromeric chromosomes or compound (six of these loci were mapped on GG1-1 and IND1-1, on IND3-2, and two on GG1-12, GG3-15, and GG-22, IND1-11, and IND3-3) and acrocentric chromosomes, which could have resulted from chromosome breakages and subsequent fusion (GG1-1 and IND1-1, particularly) that led to rearrangements of the chromosomes and possible chromosome elimination. This interpretation was further supported by the observations that chromosome counts ranged from 73 to 129 in this set of full-sib hybrids (Burner, 1997; Edmé et al, 2005a) and by the presence of univalents at meiosis (Burner, 1997). Ming et al (1998), by aligning the genome structure of Saccharum sp. with that of Sorghum bicolor, detected two possible chromosome fusions in spontaneum and chromosomal rearrangements in both S. officinarum (GG) and S. spontaneum (IND). This explanation would fit with findings from other species: centromere sites in both rice (Oryza sativa) and maize (Zea mays) are frequently involved in chromosome breakages, fusions, and assortment (Moore, 2000). Two apparent translocations, associated with marker mSSCIR38, were noticed on chromosomes GG1-1 and with the mSSCIR14 locus on IND3-3. Loci detected by these particular primers were duplicated and predominant in other regions of the two genomes. More markers are being added to the maps to better comprehend the organization and pairing behavior of these two genomes when hybridized.
Duplication in the two genomes was evidenced by markers mapped to different regions or to specific chromosomes on the GG and IND maps. The SSR markers showed a high level of similarity between the two genomes (S. officinarum and S. spontaneum) based on four homeologous groups (GG1-IND1, GG3-IND3, GG4-IND4, and GG5-IND5). It appeared that sufficient homeology exists between the two parental genomes for recombination to take place between the chromosomes. Markers in repulsion were predominant on most chromosomes of the GG map. This pattern is indicative of the behavior of an autoalloploid with most of the genome showing disomic segregation (preferential pairing with repulsion) and other regions showing polysomic inheritance (random pairing by coupling) (Wu et al, 1992). These trends were taken also as further evidence that this GG clone was a recent hybrid of unknown origin. Conversely, the genome of IND underwent mostly random pairing with a low percentage of preferential segregation, indicative of the behavior of a true autopolyploid.
Previous linkage studies conducted in sugarcane (Al-Janabi et al, 1993; da Silva et al, 1993; Ming et al, 1998) have not mapped the genomic regions associated with distortion. Unraveling regions of the genome that display a propensity for segregation distortion is of practical importance for introgression breeding. The 22% distortion level obtained in this study was high enough to justify it being considered in mapping of sugarcane interspecific crosses. This level was comparable and within the range of distortion values observed in both intraspecific (from 2 to 35%) and interspecific crosses (from 10 to 48%) (Jenczewski et al, 1997). Distortion was less frequent in IND than in GG, which had twice as many loci and chromosomes with distorted segregation. However, the reverse was reported in Ming et al (1998), who reported 11% distortion for the GG loci and 26% for the IND, based on an RFLP study. In a S. officinarum×S. spontaneum hybrid, it would be expected that the female officinarum parent would transmit its ‘2n’ sporophytic component to some of the progeny (Edmé et al, 2005a) and the male spontaneum its ‘n’ gametophytic component. Distortion in this GI population may have resulted from the high degree of divergence of the two parental species, namely the difference in ploidy level, in number of chromosomes, and in genome size. Faure et al (1993) reported a similar finding in bananas (Musa acuminata). Competition among gametes for preferential pairing and/or selection at the gamete level (female in this study) are also reported causes for distortion in segregating populations (King et al, 1991; Lyttle, 1991).
During hybridization, the marriage of the two genomes is not a harmonious union and is analogous to the intragenomic conflicts or competition alluded to by Lyttle (1991) and Hurst et al (1996), as the race for pairing of homologous (predominantly) chromosomes during meioses usually results in chromosome deletions, loss of DNA, and chromosome rearrangements, causing distortion (Eujayl et al, 1998). Moreover, studies in plants and animals have shown that meiotic drive may cause the abortion of certain classes of gametes, as those in the homozygous states are transmitted to fewer than half of the gametes in heterozygotes (King et al, 1991; Taylor and Ingvarsson, 2003). The observation that a large number of the GI hybrids in the field experiment show male and female sterility (shriveled anthers and stamens) is an indication that the meioses of these two genomes, when combined, are not without complications.
The only significant clustering of distorted loci observed was on GG2-2, which includes only higher dosage markers. This suggests either the existence of a gene on this particular chromosome linked to distortion, as is common on other crops, or the loss of a regulating gene after chromosome breakage. The presence of high dosage markers in the group of distorted marker loci on GG2-2 might be evidence of double reduction involving these gametes and resulting in distortion in this region. Double reduction occurs when two sister chromatids end up in the same gametes via random chromatid segregation after formation of quadrivalents and recombination between the locus and the centromere (Mather, 1935).
The order of marker loci was conserved on most of the homologous and homeologous chromosomes of these two genomes. Using shared marker loci between the two parental maps, comparison of the recombination rates between GG and IND was possible for GG1-1 and IND1-1 only. A similar number of meioses (nonsignificant at P≤0.05 using a χ2-test) was recorded for this region in the two genomes. Grivet et al (1996) and Ming et al (2002a) also reported a nonsignificant difference in recombination rates between S. officinarum and spontaneum in their mapping population. However, more SSR markers need to be mapped before a final assessment can be made across homeologous linkage groups. Probable rearrangements were observed for marker loci on GG3-9, GG3-11, and IND3-2. Chromosomal rearrangements in both S. officinarum (GG) and S. spontaneum (IND) were also detected in Ming et al's (2002a) study.
Segregation distortion had a considerable effect on the lengths of the maps, particularly that of the female parent. Being the recipient genome, GG was more disturbed, showing more loci with segregation distortion than IND. This resulted in Map Manager agreeing closely with Joinmap on the map length of IND and disagreeing significantly in that of GG when distortion was considered. The 2−2.5 × increase in the Map Manager map lengths resulted mostly from the addition of new marker loci (usually with loose linkage) at either end of the framework loci, or from joining multiple linkage groups together (Figure 3). Another reason for the discrepancy in the results obtained from the two programs is that Map Manager does not allow for a two-letter coding of the genotypes, which, in Joinmap, provides a better discrimination of the segregation patterns of the SSR markers and detection of their linkage phases. Based on the congruence of linkage groups and on the stringency of the significance level adopted, we argue that Joinmap was able to establish a sound framework map.
Unraveling the genomic constitution of both the officinarum and spontaneum species of Saccharum is essential to sugarcane breeding worldwide as they represent the main progenitors of sugarcane breeding populations, with officinarum conferring the high sucrose content and spontaneum the disease and stress tolerance. This study revealed that, although similarity exists in some portions of the two genomes for potential recombination, structural differences do exist in the form of substantial segregation distortion, translocations, and rearrangements which are likely to affect hybrid sterility and performance after hybridization. The recipient genome is more subject to segregation distortion. Only by identifying where the genomes resemble and differ from each other, with regard to the genes of interest, will sugarcane breeders be able to break the linkage blocks or linkats, reduce linkage drag, and exploit more heterosis in the crosses. The GI population is currently being scored with more SSR and AFLP markers with the objective of saturating the two parental framework maps, of creating a composite map where possible, and exploiting the structural differentiation of the two genomes for QTL mapping of the regions associated with sugarcane yield and quality.
References
Aitken KS, Jackson PA, McIntyre CL (2005). A combination of AFLP and SSR markers provides extensive map coverage and identification of hom(eo)logous linkage groups in a sugarcane cultivar. Theor Appl Genet 110: 789–801.
Al-Janabi AM, Honeycutt RJ, McClelland M, Sobral BWS (1993). A genetic linkage map of Saccharum spontaneum L. ‘SES 208’. Genetics 134: 1249–1260.
Berding N, Roach B (1987). Germplasm, collection, maintenance, and use. In: Heinz DJ (ed) Sugarcane Improvement through Breeding. Elsevier: NY. pp 143–210.
Burner D (1997). Chromosome transmission and meiotic behavior in various sugarcane crosses. J Am Soc Sugar Cane Technol 17: 38–50.
Cordeiro GM, Taylor GO, Henry RJ (2000). Characterisation of microsatellite markers from sugarcane (Saccharum sp. a highly polymorphic species. Plant Sci 155: 161–168.
da Silva JAG, Honeycutt RJ, Burnquist W, Al-Janabi SM, Sorrells ME, Tanksley SD et al (1995). Saccharum spontaneum L. ‘SES-208’ genetic linkage map combining RFLP- and PCR-based markers. Mol Breeding 1: 165–179.
da Silva JAG, Sobral BWS (1996). Genetics of polyploids. In: Sobral BWS (ed) The Impact of Plant Molecular Genetics. Birkhäuser: Boston, MA. pp 3–38.
da Silva JAG, Sorrells ME, Burnquist WL, Tanksley SD (1993). RFLP linkage map and genome analysis of Saccharum spontaneum. Genome 36: 782–791.
Dawe RK (1998). Meiotic chromosome organization and segregation in plants. Annu Rev Plant Physiol Plant Mol Biol 49: 371–395.
Edmé SJ, Comstock JC, Miller JD, Tai PYP (2005a). Determination of DNA content and genome size in field grown sugarcane interspecific hybrids and genotypes. J Am Soc Sugar Cane Technol 5: 1–7, Published online: www.assct.org/journal/journal.htm.
Edmé SJ, Miller JD, Glaz B, Comstock JC, Tai PYP (2005b). Genetic contribution to yield gains in the Florida sugarcane industry across 33 years. Crop Sci 45: 92–97.
Eujayl I, Baum M, Powell W, Erskine W, Pehu E (1998). A genetic linkage map of lentil (Lens sp.) based on RAPD and AFLP markers using recombinant inbred lines. Theor Appl Genet 97: 83–89.
Faure S, Noyer J, Horry J, Bakry F, Lanaud C, de Leon D (1993). A molecular-marker based linkage map of diploid bananas (Musa acuminata). Theor Appl Genet 87: 517–526.
Grattapaglia D, Sederoff R (1994). Genetic linkage maps of Eucalyptus grandis and Eucalyptus urophylla using a pseudo-testcross: mapping strategy and RAPD markers. Genetics 137: 1121–1137.
Grivet L, d'Hont A, Roques D, Feldmann P, Lanaud C, Glaszmann JC (1996). RFLP mapping in cultivated sugarcane (Saccharum sp.): genome organization in a highly polyploid and aneuploid interspecific hybrid. Genetics 142: 987–1000.
Hurst LD, Atlan A, Bengtsson BO (1996). Genetic conflicts. Quart Rev Biol 71: 317–364.
Jenczewski E, Gherardi M, Bonnin I, Prosperi JM, Oliveri I, Huguet T (1997). Insight on segregation distortions in two intraspecific crosses between annual species of Medicago (Leguminosae). Theor Appl Genet 94: 682–691.
King IP, Koebner RM, Reader SM, Miller TE (1991). Induction in the male fertility gene of the preferentially transmitted Aegilops sharonensis chromosome 4S and its application for hybrid wheat production. Euphytica 54: 33–39.
Kosambi DD (1944). The estimation of map distances from recombination values. Ann Eugen 12: 172–175.
Liu SC, Lin YR, Irvine J, Paterson A (1998). Mapping QTLs in autopolyploids. In: Paterson A (ed) Molecular Dissection of Complex Traits. CRC Press LLC: Boca Raton, FL. pp 95–102.
Lyttle TW (1991). Segregation distorters. Annu Rev Genet 25: 511–557.
Manly FF, Cudmore RH, Meer JM (2001). Map Manager QTX, cross-platform software for genetic mapping. Mamm Genome 12: 930–932.
Mather K (1935). Reductional and equational separation of the chromosomes in bivalents and multivalents. Genetics 30: 53–78.
Ming R, Liu SC, Lin YR, da Silva JAG, Wilson W, Braga D et al (1998). Detailed alignment of Saccharum and Sorghum chromosomes: comparative organization of closely related diploid and polyploid genomes. Genetics 150: 1663–1682.
Ming R, Liu SC, Moore PH, Irvine JE, Paterson AH (2002a). Construction of a Saccharum consensus genetic map from two interspecific crosses. Crop Sci 42: 570–583.
Ming R, Liu SC, Moore PH, Irvine JE, Paterson AH (2002b). QTL analysis in a complex autopolyploid: genetic control of sugar content in sugarcane. Genome Res 11: 2075–2084.
Moore G (2000). Cereal chromosome structure, evolution, and pairing. Annu Rev Plant Mol Biol 51: 195–222.
Paterson AH, Damon S, Hewitt JD, Zamir D, Rabinowitch HD, Lincoln SE et al (1991). Mendelian factors underlying quantitative traits in tomato: comparison across species, generations, and environments. Genetics 127: 181–197.
Simchen G, Hugerat Y (1993). What determines whether chromosomes segregate reductionally or equationally in meiosis? BioEssays 15: 1–8.
Tai PYP, Miller JD (2001). A core collection for Saccharum spontaneum L. from the world collection of sugarcane. Crop Sci 41: 879–885.
Tai PYP, Miller JD (2002). Germplasm diversity among four sugarcane species for sugar composition. Crop Sci 42: 958–964.
Taylor DR, Ingvarsson PK (2003). Common features of segregation distortion in plants and animals. Genetica 117: 27–35.
van Ooijen JW, Voorrips RE (2001). Joinmap v3.0, Software for the Calculation of Genetic Linkage Maps. Plant Research International: Wageningen, The Netherlands.
Voorrips RE (2002). MapChart: software for the graphical presentation of linkage maps and QTLs. J Hered 93: 77–78.
Wu KK, Burnquist W, Sorrells ME, Tew TL, Moore PH, Tanksley SD (1992). The detection and estimation of linkage in polyploids using single-dose restriction fragments. Theor Appl Genet 83: 294–300.
Acknowledgements
This study was partly supported by the USDA-ARS-SAA and the Florida Sugarcane League, Inc. We thank the Editor and two anonymous reviewers for invaluable suggestions that were incorporated in the final version of the manuscript. We are indebted to JAG da Silva, B Scheffler, and B Glaz for helpful comments and suggestions on the first draft. We acknowledge the help of B Wesley and M Gonzalez in field and lab work. Product names and trademarks are mentioned to report factually on available data; however, the USDA-ARS neither guarantees nor warrants the standard of the product, and the use of the name by USDA-ARS does not imply approval of the product to the exclusion of others that may also be suitable. The experiments reported comply with current US laws.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Edmé, S., Glynn, N. & Comstock, J. Genetic segregation of microsatellite markers in Saccharum officinarum and S. spontaneum. Heredity 97, 366–375 (2006). https://doi.org/10.1038/sj.hdy.6800888
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/sj.hdy.6800888
Keywords
This article is cited by
-
Sucrose-Specific Genic Microsatellites to Analyse the Genetic Structure Among the Commercial Hybrids and Clones of Interspecific and Intergeneric Origin in Saccharum Complex
Sugar Tech (2023)
-
QTL mapping and identification of candidate genes linked to red rot resistance in sugarcane
3 Biotech (2023)
-
Genome wide association studies in sugarcane host pathogen system for disease resistance: an update on the current status of research
Indian Phytopathology (2021)
-
Progress and Prospects of Association Mapping in Sugarcane (Saccharum Species Hybrid), a Complex Polyploid Crop
Sugar Tech (2020)
-
Development of an Axiom Sugarcane100K SNP array for genetic map construction and QTL identification
Theoretical and Applied Genetics (2019)