Abstract
Multifocality and recurrence are clinically important features of urothelial carcinomas of the urinary bladder. Recent molecular genetic studies have suggested that multifocal urothelial carcinomas are monoclonally derived from an identical transformed progenitor cell. However, most of these studies investigated advanced and poorly differentiated tumors. The study presented focuses on early papillary tumors, including 52 superficial well-differentiated multifocal and recurrent bladder carcinomas from 10 patients. Microdissection separating urothelium from stromal cells was considered essential to obtain pure tumor cell populations. Genetic analysis was carried out by applying two different methods. Dual color fluorescence in situ hybridization (FISH) with centromeric probes for chromosomes 9 and 17 and gene-specific probes for chromosome loci 9q22, 9p21, and 17p13 was carried out in parallel to loss of heterozygosity (LOH) analyses applying 5 microsatellite markers on these chromosomes. Overall, deletions on chromosome 9p were found in 47 tumors (90%), at chromosome 9q in 36 tumors (69%) and at chromosome 17p in 3 tumors (6%). There was a very high correlation of the results between FISH and LOH analysis. Ten early superficial papillary tumors showed deletion of chromosome 9p without deletion of 9q, suggesting 9p deletions as a very early event in the development of papillary urothelial carcinoma. Although in four patients, all investigated tumors showed identical genetic alterations and one patient showed no genetic alterations at the loci investigated, in five patients, two or more clones with different deletions were found. In four of these patients, the results are compatible with clonal divergence and selection of different cell subpopulations derived from a common progenitor cell. However, in one patient different alleles in two markers at chromosome 9 were deleted, favoring an independent evolution of two recurring tumor cell clones. In summary, we could show that there is considerable genetic heterogeneity in early multifocal and recurring urothelial carcinoma and demonstrated the occurrence of two independent clones in at least one patient as an indicator of possible initial oligoclonality of bladder cancer.
Similar content being viewed by others
Introduction
One of the clinically most important features of urothelial cancer of the bladder is the very high frequency of synchronous and metachronous multifocal occurrence. Furthermore, it is common for urothelial carcinomas to be accompanied by surrounding abnormal urothelium that ranges from dysplasia to carcinoma in situ. An understanding of the underlying mechanisms of the multifocality of urothelial carcinoma is required to design appropriate strategies for early detection and treatment.
Two hypotheses have been proposed to account for the development of multiple tumors in bladder cancer patients. One hypothesis proposes a “field change” in which multiple cells become initiated or partially transformed as a result of carcinogenic events and acquire early genetic alterations. Multifocal tumors would subsequently arise synchronously or metachronously as result of further independent genetic alterations of the different clones (Harris and Neal, 1992; Heney et al, 1978). The alternative hypothesis postulates a clonal development of multifocal bladder cancer. According to this “intraluminal seeding and implantation hypothesis,” the progeny of one initial transformed cell is able to spread via intraluminal dispersion or intraluminal migration and gives rise to multifocal tumors (Soloway et al, 1989).
Recent molecular genetic studies of multifocal urothelial cancers have supported the seeding or intraepithelial spread hypothesis. Sidransky et al, (1992) studied X-chromosomal inactivation patterns in female patients with multiple bladder tumors. Their results provide evidence for the monoclonal origin of bladder cancer, because every tumor of a given patient in this study was shown to have the same pattern of X-chromosomal inactivation. A challenge to the widely accepted concept of monoclonality in bladder cancer has been presented by Tsai et al (1995), who reported that large patches of urothelium harbor the same inactivated X-chromosome in bladders from individuals without evidence of bladder cancer. These results suggest that large urothelial areas arise from a single precursor cell. Such a patch could be equally exposed to carcinogens and independent malignant clones could arise in this “field,” mimicking a monoclonal origin of multifocal tumors (Bender and Jones, 1998).
However, most of the molecular genetic studies (eg, Goto et al, 1997; Habuchi et al, 1993; Petersen et al, 1993; Spruck et al, 1994; Xu et al, 1996) have focused mainly on invasive and high-grade cancers. The determination of identical genetic alterations in later stages of tumor development could reflect monoclonal occurrence of multiple tumors as well as dominant overgrowth of the most malignant tumor cell clone, the latter mimicking monoclonality. Furthermore, in the majority of cases, only two or few tumors were investigated per patient.
Thus, in this study, we investigated the presence of deletions at chromosome 9 and 17 in multiple simultaneous and consecutive superficial papillary carcinomas in different locations within the urinary bladder at early tumor stages (52 non-muscle invasive tumors, grade 1 and 2), using both fluorescence in situ hybridization (FISH) and microsatellite analysis. This study provides the first evidence of different tumor cell clones in early bladder tumors by the finding of deletion of different alleles at chromosome 9, although the majority of the investigated cases showed evidence for monoclonality of multiple papillary superficial bladder tumors.
Results
Deletion Analysis by FISH and Microsatellite Analysis
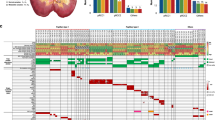
Eighteen synchronous papillary bladder tumors and 34 recurrent tumors from 10 patients were investigated. The clinical data, the localization of the tumor, and the timepoints of recurrence are given in Table 1.
Table 2 shows the summary of FISH and microsatellite analysis. There was good correlation between both methods, with discordant results in only 4 tumors (5P1, 5P2, 5RA1, 6RA1; see Table 2). There were deletions at chromosome 9p21 in 47 tumors (90%), at chromosome 9q in 36 tumors (69%), and at 17p13 in 3 tumors (6%). Only 1 of 10 patients revealed no genetic alterations at the two investigated chromosomes in all tumors. The majority of tumors (31 of 52, 60%) showed a monosomy of chromosome 9. A homozygous deletion (defined as more than 25% of cells with homozygous deletion) was detected by FISH solely at chromosome 9p21 (p16) in 16 tumors (31%).
Despite accurate microdissection of the tumor resulting in a tumor cell yield of at least 90%, a loss of both alleles in the microsatellite analysis was found in only 3 of the 16 cases with homozygous deletions. In 8 cases with a subpopulation of tumor cells with heterozygous deletion in addition to the homozygously deleted cells, LOH was detected, whereas both alleles were amplified in the remaining 5 cases. This was presumably due to amplification of both normal alleles from few contaminating normal cells in the case of a homozygous deletion in the tumor cells (Cairns et al, 1995). There was no significant difference in genetic alterations between low grade (G1) and intermediate grade (G2) tumors.
Distribution of Deletions Within the Patient Collective
Eleven early superficial papillary tumors showed deletion of chromosome 9p without deletion of 9q22, whereas none of the tumors had a deletion at chromosome 9q without involvement of 9p, suggesting 9p deletions as a very early event in the development of papillary urothelial carcinoma in this population. In 9 of these 11 patients the FISH results for chromosome 9q22 were confirmed using a different FISH probe on chromosome 9q34. There was no deletion of 9q34 in 8 tumors whereas the third recurrent tumor in patient 8 (tumor RC, see Table 2) showed a deletion in 48% of the cells when this probe was used.
In four patients all investigated tumors had identical genetic alterations (Patients 1 to 4), and one patient was found to have no genetic alterations at the investigated loci (Patient 10). However, in five patients two or more clones with different deletions were identified (patients 5 to 9, see Table 2). In four of these 5 cases (Patients 5, 6, 8,and 9) the results are compatible with clonal divergence and selection of different cell subpopulations derived from a common progenitor cell. Patient 5 had a primary bladder cancer with a p53 deletion and a recurrence after 5 months, whereas a second primary tumor in this patient showed no evidence for p53 deletion. Patient 6 had two primary tumors with deletion at 9p21 as the only genetic alteration, whereas the two recurrences 15 and 28 months after resection of the primary tumor accumulated additional genetic alteration with deletion at chromosome 9q in both and deletion of p53 in the first of the recurrent tumors (Fig. 4). Patient 8 had a primary bladder cancer without genetic alterations at the investigated loci and overall 13 recurrences with detection of two different clones that were found at several timepoints of recurrence. In Patient 9, both in the primary tumors and in one of two recurrent tumors (after 4 months), a deletion at 9p21 was found, whereas the second recurrent tumor had no deletion at this locus. On the other hand, this tumor showed the same microsatellite instability as the primary tumor. In contrast to these cases, we provide evidence that several tumor cell clones were coexistent in Patient 7. Whereas in all other cases, a deletion of identical alleles in the LOH analysis was found, in Patient 7 different alleles in two markers at chromosome 9q were deleted, favoring an independent evolution of two recurring tumor cell clones.
Silver-stained gels of microsatellite analysis of Patient 6 (see Tables 1 and 2). All five microsatellites for the normal DNA derived from the germline (N), two simultaneous primary tumors (P1, P2), and two recurrences 15 months (RA1) and 28 months (RB2) after the resection of the primary tumors are depicted. All tumors show retention of heterozygosity for 17p13.1 (p53Alu) and loss of the identical allele at both markers on chromosome 9p (D9S304 and D9S1751). On the other hand, there is retention of both alleles at the markers on chromosome 9q (D9S303 and D9S747) in both primary tumors, whereas both recurrent tumors show loss of the identical allele. These results are in complete concordance with the results obtained by FISH for these tumors. In addition, there is a reproducible microsatellite allele shift in tumor P1 at D9S747 that was not found in the other tumors of this patient. The results are compatible with a clonal origin with deletion of 9p and gain of an additional deletion of 9q in the recurrent tumors.
Discussion
Understanding whether multifocal and recurring bladder cancer is a monoclonal disease arising from one transformed cell or if multiple tumor cell clones exist simultaneously in the urinary bladder of a patient with urothelial carcinoma affects the development of early tumor detection, prevention strategies, and also tumor recurrency screening (Hruban et al, 1994; Mao et al, 1996; Steiner et al, 1997; Xu et al, 1996). The aim of the latter studies is to characterize the genetic alteration of a tumor cell clone and use this genetic fingerprint for an early and efficient diagnosis of recurrent tumors. However, if there is genetic heterogeneity in patients with multifocal urothelial carcinoma, it would be unlikely that every developing new tumor shares the same genetic alterations with the originally evaluated primary tumor.
Most studies investigating the clonality of urothelial carcinomas focused on high-grade invasive cancer and investigated only two or three tumors in one patient (Goto et al, 1997; Habuchi et al, 1992; Lunec et al, 1992; Miyao et al, 1993; Rinaldi et al, 1995; Sidransky et al, 1992). Little is known about the genetic instability, heterogeneity, and clonality of low-grade superficial urothelial carcinoma, although these tumors represent the vast majority of urothelial carcinomas (Messing and Catalona, 1997).
The aims of our study were (i) to investigate patients with multiple (3 to 14) multifocal and recurrent superficial low-grade papillary urothelial carcinomas, (ii) to use accurate microdissection to obtain a population of pure tumor cells, (iii) to investigate deletions on chromosome 9p and 9q, regions known to be deleted very early in the development of superficial papillary bladder cancer (Cairns et al, 1995; Miyao et al, 1993; Ruppert et al, 1993; Sauter et al, 1995; Simoneau et al, 1996) in addition to deletions at 17p13(p53), and (iv) to compare the deletion analysis using FISH with LOH analysis after random preamplification using I-PEP-PCR in the same tumor.
Identical genetic alterations were detected in 4 of 10 investigated patients. In contrast, genetic heterogeneity with existence of tumor cell clones with different genetic alterations was found in 5 patients (5 to 9, see shaded areas in Table 2). A number of interesting points emerge from the analysis of these patients.
First, our analysis of early superficial tumors provides further evidence that in our investigated population, deletions on chromosome 9p21/p16 seem to precede the deletion on chromosome 9q (n = 10, see also Hartmann et al, 1999). The comparison of our findings with other studies is summarized in Table 3. A possible source for the differences described could be the different markers and methods used in these studies. Recently it was demonstrated that inactivation of multiple tumor suppressors on chromosome 9 may occur during bladder cancer development (Habuchi et al, 1997; Simoneau et al, 1996, 1999). To exclude the omission of small subchromosomal deletions in this region, in 9 of these 11 patients FISH analyses was confirmed using a gene-specific probe on chromosome 9q34 (clone 168A7, Habuchi et al, 1997; the investigated tumors are marked * in Table 2). Only in one of these 9 tumors (RC, Patient 8) with normal 9q22 was a deletion of 9q34 found. Interestingly, this was the only one of the 11 tumors where discrepancies between FISH and microsatellite analysis with loss of one allele in both investigated microsatellite markers at chromosome 9q occurred. In all other cases, the FISH results were validated by the identical results in the microsatellite analyses with retention of both alleles. In summary, we do not assume to have underestimated the frequency of deletions at 9q in our study. Larger sample numbers including early neoplastic flat lesions, like dysplasias and carcinomata in situ, need to be analyzed to clarify which of the deletions on chromosome 9 occurs first during tumorigenesis. The occurrence of several tumors with intact chromosome 9 in our study using different methods of detection of genetic alterations, including the primary tumor of Patient 8 (see Table 2), favors the hypothesis that deletions on different regions of chromosome 9 are a frequent but not necessary event in the development of papillary urothelial carcinoma. The conflicting results in the order of deletion at chromosome 9 could further be explained by the fact that there is no strict order of accumulation of genetic alterations in the multistep development of urothelial carcinoma as described for colorectal cancer (Fearon and Vogelstein, 1990).
Second, the results are compatible with clonality in the majority of patients. In 8 patients, there are either identical genetic alterations in all investigated tumors, or the heterogeneity of the tumors can be explained by clonal divergence and selection of different cell subpopulations derived from a common progenitor cell. These results are in agreement with the only other study investigating multifocal and recurring superficial papillary urothelial carcinomas, by Takahashi et al (1998), which reported that multifocal tumors were considered to be derived from a single progenitor cell in 20 of 25 patients.
Third, “oligoclonality” of urothelial superficial papillary carcinoma is shown in at least one patient (Patient 7, see Table 2) with two different tumor cell clones with deletion of the opposite allele in both investigated microsatellite markers on chromosome 9q. Considering, that chromosome 9 deletions are thought to be the earliest event in the development of papillary bladder cancer, this result provides strong evidence for the development of two independent malignant clones in this patient. Both clones could be found in a recurrence 17 months after the resection of the primary tumor. In addition, one of these two clones accumulated a homozygous deletion at 9p21, whereas the other tumor showed only monosomy of chromosome 9 without evidence for homozygous deletion. This is the first report showing the deletion of different alleles at chromosome 9 in multifocal bladder carcinoma, providing evidence for the development of different tumor cell clones, ie, field cancerization.
In conclusion, our data of multifocal and recurrent early papillary tumors show considerable tumor heterogeneity with the occurrence of clones with different genetic alterations in more than 50% of the cases, and at least one patient with genetic indicators of “oligoclonal” urothelial carcinogenesis, compatible with the process of field cancerization. Investigation of the whole tumor-bearing urinary bladder, as proposed by Chaturverdi et al (1997) as “superimposed histologic and genetic mapping,” using FISH, microsatellite analyses, mutation analyses of specific genes like p53, determination of X-chromosomal inactivation, and gene expression analysis, should provide important clues on the importance of both field cancerization and intraepithelial/intraluminal spread in human urothelium.
Materials and Methods
Patient Material and Histopathological Diagnosis
Fifty-two topologically distinct urothelial tumors from 10 patients with at least 3 bladder cancers per patient were included in this study. Cystoscopy was performed after intravesical instillation of 5-ALA in patients participating in a clinical trial evaluating the photodynamic diagnosis of bladder cancer (Kriegmair et al, 1996). All patients gave written informed consent for the study. Biopsies were obtained from fluorescent lesions, immediately snap frozen in the operating room, and shipped on dry ice. Histologic diagnosis was established on serial frozen sections stained with hematoxylin-eosin (H&E). Staging was performed according to International Union Against Cancer (UICC) (Sobin and Wittekind, 1997) and grading according to World Health Organization (Murphy et al, 1994). Ten patients with 52 superficial urothelial carcinomas (28 pTaG1-, 23 pTaG2-, and 1 pT1G2-tumors) were selected for genetic analyses (Table 1). Recurrent tumors were only included in the study if there was a minimum of 4 months between cystoscopies.
Microdissection, Cell Dissociation, and DNA Isolation
From each frozen sample a 4-μm frozen section was stained with H&E and the presence of tumor was confirmed. Two consecutive 20-μm sections (for FISH analyses) and six 5-μm sections (for PCR analyses) were stained with methylene blue for approximately 15 seconds. The tumor was separated from stromal cells by microdissection with a needle (22G) under an inverted microscope (×40 magnification) or using laser microdissection (PALM; Wolfrathshausen, Germany; Schütze and Lahr, 1998). The microdissected probes contained at least 90% urothelial cells (Fig. 1, a and b). Cell dissociation was performed as described previously (Hartmann et al, 1999). The cells were pelleted on silanized glass slides by standard microcentrifugation, fixed in freshly prepared methanol/acetic acid (3:1), air dried, and stored at −20° C for up to 3 months. For PCR analysis, the microdissected tissue samples were digested in 10 to 30 μl Expand Lysis buffer (1× Expand HiFi buffer; Boehringer Mannheim, Mannheim, Germany) including 4 mg/ml Proteinase K and 0.5% Tween 20 (Merck, Darmstadt, Germany) at 55° C for 14 hours, followed by heat inactivation of Proteinase K for 15 minutes at 94° C. Normal DNA was isolated from 7.5 ml EDTA blood using the Qiagen DNA blood isolation kit (Quiagen, Hilden, Germany).
DNA Probes and Probe Labeling
For counts of chromosomes 9 and 17, biotin-labeled centromeric probes (D9Z1 and D17Z1; Oncor, Gaithersburg, Maryland) were used. These probes were combined with P1 probes (Shepherd et al, 1994), obtained from the LBNL/UCSF Resource for Molecular Cytogenetics (http://rmc-www.lbl.gov). These probes have a length of approximately 60 to 80 kB and are cloned in pAd10SacBII. The following probes were used: RMC09P007 for chromosome 9p21 (CDKI2/p16 locus), RMC09P008 for chromosome 9q22 (FACC locus), and RMC17P078 for chromosome 17p13 (p53 gene locus). Furthermore, a cosmid probe from chromosome 9q34.2, kindly provided by M. Knowles (Molecular Genetics Laboratory, Marie Curie Research Institute, Surrey, United Kingdom; clone 168A7, Habuchi et al, 1998), was used to confirm the FISH results obtained for 9q in a subset of samples with deletion of 9p21 and normal results for 9q22. DNA was isolated using alkaline lysis (Lee and Rasheed, 1990) and labeled with digoxigenin-11-dUTP using standard nick translation protocols (Boehringer Mannheim).
Fluorescence In Situ Hybridization
FISH was performed as described (Hartmann et al, 1999). Metaphase spreads were used as controls to assure specificity of the probes. Furthermore, for every hybridization, cytospins of cultured urothelial cells (Urotsa) without any alterations at the investigated gene loci were included to assure an estimation of hybridization efficiency (Fig. 2a).
Fluorescence in situ hybridization (FISH) using a centromeric probe for chromosome 9 and a gene-specific probe for 9p21 (p16) in urothelial cells from the Urotsa cell line (2a) and the papillary urothelial carcinoma shown above (2b; Patient 9; tumor P1). Whereas in control hybridizations with Urotsa cells two red centromere and two green gene-specific signals are visible in the majority of cells (2a), a deletion of 9p21 with two red centromeric signals and only one green gene-specific signal is seen in the tumor (2b). FISH with both probes for chromosome 9q showed normal results in this tumor.
Scoring of FISH Signals
Cells were selected for scoring with DAPI staining according to morphological criteria. Clearly distinguishable small lymphocytes were disregarded, all other cells were scored. Slides were analyzed if more than 75% of cells were interpretable. Copy numbers for centromeres and specific gene regions were counted in 200 cells if possible (minimum 80 cells). Only cells with non-overlapping and intact nuclei were counted. Cells without any signal were disregarded. Slides were counted by one of the investigators (UR) and every fifth slide was confirmed by a second observer. For these cases the mean of both counts was used. As a measure of deletion, the percentage of cells containing either one copy of centromere 9 or 17 or fewer gene-specific signals than centromeric signals was calculated for each hybridization (defined as percentage of deletion). The average percentage of deletion of 10 hybridizations of a normal urothelial cell line (Urotsa) and 6 hybridizations of dissociated normal urothelium from patients without bladder cancer was less then 15 ± 5% for every probe. Because there was no normal tissue available from patients treated in exactly the same manner as those investigated, a tumor was considered deleted for a specific chromosomal locus if the percentage of deletion was >40% (equivalent 2× mean ± sd) as a conservative evaluation of the results. Monosomy and homozygous deletion were defined as more then 50% of all deleted cells having either of these two alterations (eg, Fig. 3, a and b).
FISH using a centromeric probe for chromosome 9 and gene-specific probes for chromosome 9q22 (3a) and 9p21 (3b) in a papillary urothelial carcinoma (Patient 8, tumor RB2). FISH with the probe for chromosome 9q22 shows monosomy with one signal for both centromere and gene-specific probes (3a), whereas FISH with the probe for 9p21 reveals a homozygous deletion with one centomeric red signal and lack of gene-specific green signals in the majority of cells (3b). Note few cells with monosomy and retention of one copy of 9p21, indicating tumor heterogeneity.
Whole Genome Amplification by I-PEP-PCR and Microsatellite Analysis
Improved primer-extension-preamplification PCR (I-PEP) was performed using a MJR PTC200 thermocycler (Biozym, Oldenburg, Germany) as described previously (Dietmaier et al, 1999). In brief, after Proteinase K digestion 50 amplification cycles were performed, each consisting of a 1-minute step at 94° C, a 2-minute step at 37° C, a ramping step of 0.1° C/second to 55° C, a 4-minute step at 55° C, and a 30-second step at 68° C. I-PEP-PCR was set up by adding 50 μl I-PEP mix (final concentration 0.05 mg/ml gelatin, 16 μmol/L totally degenerate 15 nucleotide long primer; MWG Biotech GmbH, Ebersberg, Germany; 0.1 mmol/L dNTP, 3.6U Expand High fidelity polymerase, Boehringer, Mannheim, Germany; 2.5 mmol/L MgCl2, in 1× PCR buffer No 0.3, provided with the polymerase) to 10 μl lysed tissue.
Specific microsatellite PCR (0.2 mmol/L dNTP, 0.3 μmol/L primers, 0.5 U Taq polymerase, Life Technologies, 1.5 mmol/L MgCl2) was performed using 2 μl of the preamplified PCR as template in a final volume of 20 μl or 30 μl, respectively, in a PTC100 thermocycler (MJ Research, Watertown, Maryland) for 35 cycles: 94° C for 1 minute, 50 to 60° C for 1 minute, 72° C for 1 minute, followed by a final extension at 72° C for 8 minutes. Amplified microsatellites were detected by polyacrylamid gel electrophoresis and silver staining as described (Schlegel et al, 1995). The silver-stained gels were assessed visually, and informative cases were scored as allelic loss when intensity of the signal for a tumor allele was decreased to 50% relative to the matched normal allele. Microsatellite instability was defined as the occurrence of additional alleles in the tumor tissue compared with the normal DNA. All cases of allelic loss (LOH) and microsatellite instability were confirmed at least once. Twenty microsatellites with evident or questionable allelic loss were selected and reinvestigated using capillary electrophoreses on an ABI301 automatic sequencer and Genescan software as described (Canzian et al, 1996). In all cases the results confirmed the results of the silver-stained gels (data not shown). Primers were obtained from MWG Biotech GmbH. Primer sequences and annealing temperatures were as follows: p53Alu (17p13.1.): 5′AGGAGGTTGCAGTAAGCGGA3′ and at 60° C 5′AACAGCTCCTTTAATGGCAG3′ (Futreal et al, 1991), D9S304 (9p21): 5′GTGCACCTCTACACCCAGAC3′ and at 60° C 5′TG- TGCCCACACACATCTATC3′, D9S1751 (Pky11, 9p21): 5′TTGTTGATTCTGCCTTCAAAGTCTTTTAAC3′ and at 55° C: 5′CGTTAAGTCCTCTATTACACAGAG3′ (Cairns et al, 1995), D9S303 (9q22) at 55° C 5′CAACAAAGCAAGATCCCTTC3′ 5′GGTACTTGGAAACTCTTGGC3′, D9S747 (9q32) at 56° C 5′GCCATTATTGACTCTGGAAAAGAC3′ 5′CAGGCTCTCAAAATATGAACAAAAT3′.
References
Baud E, Catilina P, Boiteux J-P, and Bignon Y-J (1998). Human bladder cancers and normal bladder mucosa present the same hot spot of heterozygous chromosome-9 deletion. Int J Cancer 77:821–824.
Bender CM, and Jones PA (1998). Molecular genetics in carcinoma of the bladder. In: Petrovich Z, Baert L, and Brady LW, editors. Carcinoma of the bladder: Innovations in management. Heidelberg, New York:Springer, pp. 37–51.
Cairns P, Polascik TJ, Eby Y, Tokino K, Califano J, Merlo A, Mao L, Herath J, Jenkins R, Westra W, Rutter JL, Buckler A, Gabrielson E, Tockman M, Cho KR, Hedrick L, Bova GS, Isaacs W, Gocj W, Schwab D, and Sidransky D (1995). Frequency of homozygous deletion at p16/CDKN2 in primary human tumours. Nature Genet 11:210–212.
Canzian F, Salovaara R, Hemminki A, Kristo P, Chadwick RB, Aaltonen LA, and de la Chapelle A (1996). Semiautomated assessment of loss of heterozygosity and replication error in tumors. Cancer Res 56:3331–3337.
Chaturverdi V, Li L, Hodges S, Johnston D, Ro JY, Logothetis C, von Eschenbach AC, Batsakis JG, and Czerniak B (1997). Superimposed histologic and genetic mapping of chromosome 17 alterations in human urinary bladder neoplasia. Oncogene 14:2059–2070.
Dietmaier W, Hartmann A, Wallinger S, Heinmöller E, Kerner T, Endl E, Jauch KW, Hofstädter F, and Rüschoff J (1999). Multiple mutation analyses in single tumor cells with improved whole genome amplification. Am J Pathol 154:83–95.
Fearon ER, and Vogelstein B (1990). A genetic model for colorectal tumorigenesis. Cell 61:759–763.
Futreal PA, Barrett JC, and Wiseman RW (1991). An Alu polymorphism intragenic to the p53 gene. Nucl Acids Res 19:6977.
Goto K, Konomoto T, Hayashi K, Kinukawa N, Naito S, Kumuzawa J, and Tsuneyoshi M (1997). p53 mutations in multiple urothelial carcinomas: A molecular analysis of the development of multiple carcinomas. Mod Pathol 10:428–437.
Habuchi T, Takahashi R, Yamada H, Kakehi Y, Sugiyama T, and Yoshida O (1993). Metachronous multifocal development of urothelial cancers by intraluminal seeding. Lancet 342:1087–1088.
Habuchi T, Yoshida O, and Knowles MA (1997). A novel candidate tumour suppressor locus at 9q32–33 in bladder cancer: Localization of the candidate region within a single 840 kb YAC. Hum Mol Gen 6:913–919.
Habuchi T, Luscombe M, Elder PA, and Knowles MA (1998). Structure and methylation-based silencing of a gene (DBCCR1) within a candidate bladder cancer tumor suppressor region at 9q32–q33. Genomics 48:277–288.
Harris AL, and Neal DE (1992). Bladder cancer-field versus clonal origin. N Engl J Med 326:759–761.
Hartmann A, Moser K, Kriegmair M, Hofstetter A, Hofstaedter F, and Knuechel R (1999). Frequent genetic alterations in simple urothelial hyperplasias of the bladder in patients with papillary urothelial carcinoma. Am J Pathol 154:721–727.
Heney NM, Daly J, Prout GR Jr, Nieh PT, Heaney JA, and Trebeck NE (1978). Biopsy of apparently normal urothelium in patients with bladder carcinoma. J Urol 120:559–560.
Hruban RH, Van der Riet P, Erozan YS, and Sidransky D (1994). Brief report: Molecular biology and the early detection of carcinoma of the bladder: The case of Hubert H. Humphrey. N Engl J Med 330:1276–1278.
Keen AJ, and Knowles MA (1994). Definition of two regions of deletion on chromosome 9 in carcinoma of the bladder. Oncogene 9:2083–2088.
Kriegmair M, Baumgartner R, Knuchel R, Stepp H, Hofstadter F, and Hofstetter A (1996). Detection of early bladder cancer by 5-aminolevulinic acid induced porphyrin fluorescence [see comments]. J Urol 155:105–109.
Lee S, and Rasheed S (1990). A simple procedure for maximum yield of high-quality plasmid DNA. Biotechniques 9:676–679.
Linnenbach AJ, Pressler LB, Seng BA, Kimmel BS, Tomaszewski JE, and Malkowicz SB (1993). Characterization of chromosome 9 deletions in transitional cell carcinoma by microsatellite assay. Hum Mol Genet 2:1407–1411.
Lunec J, Challen C, Wright C, Mellon K, and Neal DE (1992). C-erbB-2 amplification and identical p53 mutations in cocomitant transitional carcinomas of renal pelvis and urinary bladder. Lancet 339:439–440.
Mao L, Schoenberg MP, Scicchitano M, Erozan YS, Merlo A, Schwab D, and Sidransky D (1996). Molecular detection of primary bladder cancer by microsatellite analysis. Science 271:659–662.
Messing EM, and Catalona W (1997). Urothelial tumors of the urinary tract. In: Campbell's Urology, ed. 5, vol. 3: 2327–2410. New York: Saunders.
Miyao N, Tsai YC, Lerner SP, Olumi AF, Spruck CH III, Gonzalez Zulueta M, Nichols PW, Skinner DG, and Jones PA (1993). Role of chromosome 9 in human bladder cancer. Cancer Res 53:4066–4070.
Murphy WM, Beckwith JB, and Farrow GE (1994). Atlas of tumor pathology. In: Rosai J, and Sobin LH, editors. Third Series, Fascicle 11, Tumors of the kidney, bladder, and related urinary structures. Washington, DC:Armed Forces Institute of Pathology.
Petersen I, Ohgaki H, Ludeke BI, and Kleihues P (1993). P53 mutations in phenacetin-associated human urothelial carcinomas. Carcinogenesis 14:2119–2122.
Rinaldi F, Mairs R, Wheldon T, Smith G, Yates A, and Symonds P (1995). Clonal analysis of phenacetin-implicated urothelial carcinoma. Int J Cancer 63:881–882.
Ruppert JM, Tokino K, and Sidransky D (1993). Evidence for two bladder cancer suppressor loci on human chromosome 9. Cancer Res 53:5093–5095.
Sauter G, Moch H, Carroll PC, Kerschmann R, Mihatsch MJ, and Waldman FM (1995). Chromosome-9 loss detected by fluorescence in situ hybridization in bladder cancer. Int J Cancer 64:99–103.
Schlegel J, Bocker T, Zirngibl H, Hofstädter F, and Rüschoff J (1995). Detection of microsatellite instability in human colorectal carcinomas using a non-radioactive PCR-based screening method. Virchows Arch 426:223–227.
Schütze K, and Lahr G (1998). Identification of expressed genes by laser-mediated manipulation of single cells. Nature Biotechnol 16:737–742.
Shepherd NS, Pfrogner BD, Coulby JN, Ackerman SL, Vaidyanathan G, Sauer RH, Balkenhol TC, and Sternberg N (1994). Preparation and screening of an arrayed human genomic library generated with the P1 cloning system. Proc Natl Acad Sci U S A 91:2629–2633.
Shigyo M, Sugano K, Fukayama N, Taniguchi T, Tobisu K, Fujimoto H, Doi N, Niwakawa M, Shimamura K, Tsukamoto T, Sekiya T, and Kakizoe T (1998). Allelic loss on chromosome 9 in bladder cancer tissues and urine samples detected by blunt-end single-strand DNA conformation polymorphism. Int J Cancer 78:425–429.
Sidransky D, Frost P, Von Eschenbach A, Oyashy R, Preisinger AC, and Vogelstein B (1992). Clonal origin of bladder cancer. N Engl J Med 326:737–740.
Simoneau AR, Spruck III CH, Gonzalez-Zulueta M, Gonzalgo ML, Chan MF, Tsai YS, Dean M, Steven K, Horn T, and Jones PA (1996). Evidence for two tumor suppressor loci associated with proximal chromosome 9p to q and distal chromosome 9q in bladder cancer and the initial screening for GAS1 and PTC mutations. Cancer Res 56:5039–5043.
Simoneau M, Aboulkassim TO, LaRue H, Rousseau F, and Fradet Y (1999). Four tumor suppressor loci on chromosome 9q in bladder cancer: evidence for two novel candidate regions at 9q22.3 and 9q31. Oncogene 18:157–163.
Sobin LH, and Wittekind C (1997). TNM classification of malignant tumours, 5th ed. New York, Weinheim:Wiley-Liss.
Soloway MS, Jordan AM, and Murphy WM (1989). Rationale for intravesical chemotherapy in the treatment and prophylaxis of superficial transitional cell carcinoma. 310:215–236.
Spruck III CH, Ohneseit PF, Gonzalez-Zulueta M, Esrig D, Miyao N, Tsai YC, Lerner SP, Schmütte C, Yang AS, Core R, Dubeau L, Nichols PW, Hermann GG, Steven K, Horn T, Skinner DG, and Jones PA (1994). Two molecular pathways to transitional cell carcinomas of the bladder. Cancer Res 54:784–788.
Steiner G, Schoenberg MP, Linn JF, Mao L, and Sidransky D (1997). Detection of bladder cancer recurrence by microsatellite analysis of urine. Nat Med 3:621–624.
Takahashi T, Habuchi T, Kakehi Y, Mitsumori K, Akao T, Terachi T, and Yoshida O (1998). Clonal and chronological genetic analysis of multifocal cancers of the bladder and upper urinary tract. Cancer Res 58:5835–5841.
Tsai YC, Simoneau AR, Spruck III CH, Nichols PW, Steven K, Buckley JD, and Jones PA (1995). Mosaicism in human epithelium: Macroscopic monoclonal patches cover the urothelium. J Urol 153:1693–1700.
Xu X, Stower MJ, Reid N, Garner RC, and Burns PA (1996). Molecular screening of multifocal transitional cell carcinoma of the bladder using p53 mutations as biomarkers. Clin Cancer Res 2:1795–1800.
Acknowledgements
We thank Helmut Kutz, Andrea Schneider, and Monika Kerscher for excellent technical support. Furthermore, we thank M.A. Knowles for providing the FISH probe for chromosome 9q34.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supported by DFG grant 263/7–1 (RK) and grant 10–1096-Ha I from the Dr. Mildred Scheel Foundation of Cancer Research (AH, RK).
Rights and permissions
About this article
Cite this article
Hartmann, A., Rösner, U., Schlake, G. et al. Clonality and Genetic Divergence in Multifocal Low-Grade Superficial Urothelial Carcinoma as Determined by Chromosome 9 and p53 Deletion Analysis. Lab Invest 80, 709–718 (2000). https://doi.org/10.1038/labinvest.3780074
Received:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3780074
This article is cited by
-
Non-invasive papillary urothelial carcinoma of the vagina: molecular analysis of a rare case identifies clonal relationship to non-invasive urothelial carcinoma of the bladder
Virchows Archiv (2017)
-
Determining the origin of synchronous multifocal bladder cancer by exome sequencing
BMC Cancer (2015)
-
Comparison of the clonality of urothelial carcinoma developing in the upper urinary tract and those developing in the bladder
SpringerPlus (2013)
-
Molecular heterogeneity in malignant peripheral nerve sheath tumors associated with neurofibromatosis type 1
Human Genomics (2012)
-
Analysis of molecular intra-patient variation and delineation of a prognostic 12-gene signature in non-muscle invasive bladder cancer; technology transfer from microarrays to PCR
British Journal of Cancer (2012)