Abstract
A member of tetraspanin CD151 is a scaffold protein of laminin-binding integrins and it plays an important role in stable interaction between cells and basement membrane. Although the upregulation of CD151 in tumor cells is thought to accelerate tumor invasion and metastasis, detailed pathological investigation on CD151 and its association with integrins has not been well documented, yet. In the present study, we showed that the expression levels of CD151 and its associated integrin subunits in epidermal carcinoma cell HSC5 were higher than those in immortalized epidermal cell HaCaT. By the stimulation of epidermal growth factor, CD151 was dissociated from cell surface and dispersed in the cytoplasm, and α3β1 integrin was concomitantly internalized. To understand the significance of CD151 in tumor cell dynamics, CD151 in HSC5 was knocked down (HSC5CD151−), and the expression of integrin subunits and matrix metalloproteinases (MMPs) were investigated. In HSC5CD151−, striking morphological alteration on Matrigel and laminin, and cytoskeletal rearrangements were demonstrated. α3β1 integrin was internalized in part, and α6β4 integrin was re-distributed from basal site to cell periphery. Quantitative RT-PCR, Western blot and zymography revealed that the expression levels of MMP2, MMP7 and MMP9 were markedly downregulated in HSC5CD151−. Immunoprecipitation assay demonstrated that MMP7 was co-immunoprecipitated with CD151. In double stainings, MMP7 was colocalized with CD151 at the leading edge of lamellipodia under migratory status. These results elucidated the importance of CD151 as one of the key molecules for integrin-dependent carcinoma–stroma interaction. It is indicated that CD151 might contribute not only to cell stabilization by associating with adhesion complexes but also to cell migration by inducing integrins re-localization and MMPs production.
Similar content being viewed by others
Main
The interaction between epithelial cells and basement membrane is supported by special adhesion complexes called hemidesmosome and focal adhesion. Hemidesmosome includes α6β4-integrin and provides the linkage between the intracellular keratin filament system to laminin of extracellular matrix (ECM).1, 2 Focal adhesion includes α3β1-integrin and controls actin-based cytoskeletal rearrangement.3 These integrin subsets are critical in various physiological and pathological events, that is wound healing and tumor invasion. In several tumor types, altered expression of integrins has been reported. For example, the expression level of α6β4-integrin increased during carcinogenesis of the skin,4 and its upregulation is correlated with poor prognosis in several types of carcinomas.5 α3β1-integrin was also shown to be upregulated in several advanced tumors.6, 7 In contrast, some other studies reported the downregulation of α6β4- and/or α3β1-integrins in certain advanced carcinomas.8, 9, 10 Collective pathological studies suggest that the expression of integrins in tumor cells can be upregulated or downregulated, or the lost polarity depending on invasive activities at local lesions and proinflammatory microenvironments of tumor nests.
CD151 is a member of tetraspanin, and is expressed in a wide variety of cell types such as hematopoietic cells, epithelial cells and tumor cell lines.11, 12, 13, 14 The interaction of CD151 with α3β1-integrin is known to be highly stable and stoichiometric,15, 16 and it is believed that CD151 associates directly with α3β1-integrin.17, 18 The association between CD151 and α6β4-integrin is also strong, which seems to be important in the pathophysiology of basal epithelial layer, because CD151-α6β4-integrin complex composes hemidesmosomes and links to the basement membrane.19, 20 In tumor progression, CD151-transfected carcinoma cells obtained the ability of invasion and metastasis.21 The expression level of CD151 was correlated with poor prognosis in colon cancer.13, 22 These reports suggest that CD151 may contribute to tumor progression. However, the importance of CD151 in association with upregulation or downregulation of laminin-binding integrins remains to be elusive. The recent studies on CD151 demonstrated the internalization of CD151 with α6β4-integrin in pancreatic adenocarcinoma cells in vitro,23 and the dissociation of CD151 from laminin-binding integrins in migratory basal keratinocytes in vivo.24 For better understanding of tumor migration and invasion, it is important to investigate the role of CD151 in integrin-dependent interaction between tumor cells and ECM.
In the present study, we investigated the expression of CD151 and laminin-binding integrins in epidermal-derived carcinoma cell HSC5. To understand the importance of CD151 in carcinoma cell migration, CD151 was knocked down and the effects on the distribution of laminin-binding integrin subunits, adhesion strength and the expression of matrix metalloproteinases (MMPs) were investigated. We demonstrated striking redistribution of α3β1- and α6β4-integrin subsets, attenuated adhesion strength and downregulation of MMPs.
Materials and methods
Reagents and Cells
Monoclonal antibodies against clones 14A2 and 11G5a that recognize different sites of CD15125 were purchased from BD Biosciences (San Diego, CA, USA) and Serotec (Oxford, UK), respectively. Monoclonal antibodies against CD9-, α3-, α6-, β1-integrins (DakoCytomation, Tokyo, Japan), CD63, CD81 (Novocastra, Newcastle upon Tyne, UK), CD82 (Santa Cruz Biotechnology, Santa Cruz, CA, USA), α2-, β4-integrins (BD Biosciences), MMP2, MMP7 and MMP9 (Daiich Fine Chemical, Toyama, Japan) were used with dilution of 1:100. Mouse immunoglobulin fraction was used as a negative control with dilution of 1:100. Human epidermal carcinoma cell line HSC5, established from the squamous cell carcinoma of the auricle skin, was obtained from Japanese Collection of Research Bioresources (Tokyo, Japan), and ovarian carcinoma cell line HTOA was obtained from RIKEN BRC Cell Bank (Tsukuba, Japan). Human immortalized keratinocyte HaCaT was kindly provided by Dr NE Fusenig.26 The cells were maintained in DMEM, supplemented with 10% fetal calf serum, 1% penicillin and streptomycin at 37°C in 5% CO2. Epidermal growth factor (EGF) was purchased from R&D Systems (Minneapolis, MN, USA).
RNA Isolation
Total RNAs were obtained using RNeasy Mini kit (Qiagen Hilden, Germany) and cDNA synthesis was performed using QuantiTect Reverse Transcription kit (Qiagen).
RT-PCR and Real-Time RT-PCR
Reverse transcriptase-mediated (RT)-PCR was carried out with the following primers: (forward, F) 5′-GCTGGAGATCATCGCTGGTATC-3′ and (reverse, R) 5′-GGTGGTAG CGCCTGGTCAT-3′ for CD151, (F) 5′-TTCGGCCCAGGCTAAGTTAG-3′ and (R) 5′-CGGCAAGCCAGAAGATGAAG-3′ for CD9, (F) 5′-CTGGAAAGGAAACAGCTACATGATT-3′ and (R) 5′-CCTGCATCGTGTACCCAATATAGA-3′ for α3, (F) 5′-GTGACTGCTGCTGCCGAAA-3′ and (R) 5′-GGTACTCCCGAATACTGAGCTACAG-3′ for α6, (F) 5′-GTTTCGATGCCATCATGCAA-3′ and (R) 5′-AAAGTGAAACCCGGCATCTG-3′ for β1, (F) 5′-TCCAGGAAGATCCATTTCAACTG-3′ and (R) 5′-AGTCGCAATACGGGTACAGGTT-3′ for β4, (F) 5′-TCCACTGTTGGTGGGAACTCA-3′ and (R) 5′-TGGTCGCACACCACATCTTT-3′ for MMP2, (F) 5′-TTCCAAAGTGGTCACCTACAGGAT-3′ and (R) 5′-CTTTGCCCCACATGTTTAAAGC-3′ for MMP7, (F) 5′-GTGGCGGCGCATGAGT-3′ and (R) 5′-TGCCGGATGCCATTCAC-3′ for MMP9, (F) 5′-CCACCCATGGCAAATTCC-3′ and (R) 5′-TGAT-GGGATTTCCATTGATGAC-3′ for GAPDH. Conditions for PCR were as follows: at 95°C for 5 min, 30 cycles at 95°C for 15 s, 58°C for 15 s, 72°C for 1 min, with an extension step of 7 min at 72°C. For real-time RT-PCR analysis, QuantiTect SYBR Green PCR kit (Qiagen) was used. Conditions for PCR were at 50°C for 2 min, at 95°C for 15 min, 40 cycles at 95°C for 30 s and at 60°C for 30 s.
siRNA Transfection
The cells were transfected with siRNA using RNAiFect Reagent (Qiagen) according to the manufacturer's instructions. The siRNA directed against CD151 was 5′-UGGGUGAGUUCAACGAGAAGAAGAC-3′. The concentration of siRNA was 10 μg/5 ml per dish. Control siRNA (non-silencing) was purchased from Qiagen. In each examination, the efficiency of siRNA transfection was checked by fluorescence microscopy using FITC-labeled siRNA. Transfected cells were harvested at 48 h. The purity of CD151-negative cells was checked by FACScan (Becton Dickinson, Mountain View, CA, USA) each time before analysis.
Fluorescence Cytostainings
Chamber slides were washed with PBS, fixed with 4% paraformaldehyde for 10 min and stained according to the manufacturer's instruction. In addition to CD151 and integrins, MMP2, MMP7, MMP9 and phosphotyrosine (Upstate, Lake Placid, NY, USA) were used for fluorescence cytostainings. In the stainings for actin filaments, Alexa Fluor 488 phalloidin (Invitrogen) was used. Working dilution of phalloidin was 1:20 and of other antibodies was 1:100. The staining patterns were analyzed using fluorescence microscopy (Zeiss Axio Imager). For three-dimensional analysis of integrins, laser scan microscopy was used (Zeiss LSM510).
Flow Cytometry
Cell suspensions were stained with CD9, CD63, CD81, CD82, CD151 or integrins listed above, followed by FITC-conjugated goat anti-mouse immunoglobulins (ICN Pharmaceuticals, Aurora, OH, USA). The cells were cultured under the condition of 5 × 106 cells in 10 cm dish for 24 h, then harvested using EDTA (ICT Pharmaceuticals, Aurora, OH, USA) containing 5% trypsin. Cells were analyzed on flow cytometry (FACS), and relative mean fluorescence intensity (MFI) was quantified as absolute MFI/MFI of negative control cells using CELLQest software 2.1.1 (Becton Dickinson). The experiment was repeated three times and the expression levels were expressed as the mean±s.e.
In Vitro Morphogenesis Assay
Matrigel (BD Biosciences) was placed on 24-well plates (300 μl/well) and allowed to solidify at 37°C for 30 min. Other wells were coated with 2 μg/cm2 of laminin (Sigma). Laminin-5 was prepared according to the method described previously.27 Time-lapse morphological analysis was performed for 24 h using CCD camera VB-7000 (Keyence, Osaka, Japan). All the results were obtained from at least three independent experiments.
In Vitro Adhesion and Detachment Assays
Static adhesion assay and detachment assay were performed using the methods described previously.28, 29 Briefly, 105 cells were washed with 2% BSA and seeded onto laminin-coated (2 μg/cm2, Sigma) 24-well plates. After 0.5, 1 and 2 h, adherent cells were counted in three independent fields, respectively. The adherent cells were imaged using an inverted light microscope at × 20 objective, photographed by Olympus DP12 in three independent fields in triplicate wells, and the cell numbers were counted. For detachment assay, 105 cells were incubated for 24 h, and treated with trypsin/EDTA for 0, 1, 2, 3 and 5 min, respectively. For centrifugal force-based detachment assay, laminin adhesion was led to occur for 30 min. The wells of the plate were then completely filled with medium containing 0.35% BSA and sealed. The plates were inverted and centrifuged for 8 min at increasing forces (500, 750 and 1000 g). Detachment-resistant cells were counted using the same method as described above.
Immunoprecipitation Assay
A total of 200 μg of cell lysate in M2 buffer (20 mM Tris–HCl, pH 7.6, 350 mM NaCl, 3 mM EDTA, 2 mM DTT, 0.5 mM phenylmethylsulfonyl fluoride, 20 mM β-glycerophosphate and 0.5% NP-40) was mixed with 5 μg of antibody prebound to protein G plus/protein A agarose (Calbiochem) at 4°C overnight. Immune complexes were washed three times and then loaded on 12.5% sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). The blots were probed with specific antibody. Rabbit polyclonal antibodies against β1 (Santa Cruz), β4 (Chemicon), goat polyclonal antibody against α3 (Santa Cruz), and mouse monoclonal antibodies against α6 (Merck), MMP2, MMP7 and MMP9 (Daiich Fine Chemical) were used.
Western Blot
Fifty micrograms of cell lysate proteins were electrophoresed on 12.5% SDS-PAGE and transferred to a PVDF membrane (Millipore, Bedford, MA, USA). Horseradish peroxidase-conjugated IgG (1:2500) was used as the secondary antibody. Bands were detected using an enhanced chemiluminescence system, according to the Hybond ECL protocol (GE Healthcare, Buckinghamshire, UK).
Zymography
Culture medium (CM) was prepared from either HSC5 or HTOA cells. A total of 105 cells were cultured in 1 ml of serum-free DMEM for 48 h, then 10 μl of CM was electrophoresed at 4°C in 10% crosslinked SDS-PAGE, containing either 0.1% gelatin (Difco, Detroit, MI, USA) or casein (Wako, Osaka, Japan). Following electrophoresis, the gel was washed with 2.5% Triton X-100 followed by incubation in Tris–HCl, 0.5 mM CaCl2, 10−6 M ZnCl2, pH 8.0, at 37°C for 16 h. Coomassie brilliant blue staining was then carried out.
Statistical Analysis
Values were expressed as the mean±s.e. Student's t-test was used for statistical evaluation in FACS and real-time RT-PCR data. One-way ANOVA was used in adhesion assay and detachment assay. Statistical significance was assumed when P<0.05 was obtained.
Results
Expression of Tetraspanins and Integrins, and Direct Interaction of CD151 with Laminin-Binding Integrins in Epidermal-Derived Carcinoma Cells and Immortalized Keratinocytes
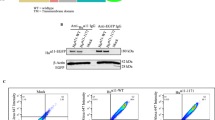
We compared the expression levels of CD151 and associated integrins between epidermal-derived carcinoma cell line HSC5 and immortalized keratinocyte HaCaT (Figure 1a–c). It was reported that the monoclonal CD151 antibodies 11G5a and 14A2 showed different staining patterns in cardiac muscles and intestinal mucosa, indicating that these clones recognize different epitopes of CD151.25 In our preliminary experiments, epidermal keratinocytes and hair follicles were positive for 11G5a and 14A2, and the former showed diffuse stainings in both basal and squamous layers whereas the latter showed restricted stainings in basal layer (data not shown). In this study, we used both clones to investigate CD151. Four other tetraspanins (CD9, CD63, CD81 and CD82) were also investigated. By FACS, CD9 was highly expressed, and 11G5a-recognized CD151 was also significantly expressed both in HSC5 and HaCaT. HSC5 showed higher expression of these molecules than HaCaT did (Figure 1a). 14A2-recognized CD151 is expressed at high level in HSC5 but at much lower level in HaCaT. MFI of CD63, CD81 and CD82 were either at very low or negative levels in both cell types. The expression levels of α2, α6-, β1- and β4-integrin subunits were higher in HSC5 than in HaCaT (Figure 1b). We also examined the expression of these molecules in another epidermal carcinoma cell line HSC1, and obtained similar results, although the MFI of CD151 was lower than that in HSC5 (data not shown). FACS results suggested that these tetraspanins and integrins were upregulated in invasive epidermal carcinoma cells. However, it should be noted that HaCaT is an immortalized cell and that the results may not represent the differences between epidermal carcinomas and normal keratinocytes.
The expression patterns of tetraspanins and integrins in HSC5 and HaCaT. (a and b) FACS result shows that CD9 and CD151 are expressed higher in HSC5 than in HaCaT. Other three tetraspanins are almost undetectable in both cell lines. The expression levels of α2-, α6-, β1- and β4-integrins are higher in HSC5 than in HaCaT. Values are expressed as relative MFI±s.e. (*P<0.05, **P<0.01). (c) CD151 was immunoprecipitated from HSC5 and HaCaT cells, respectively. Aliquots of the immunoprecipitate were electrophoresed and transferred to PVDF membrane, and probed with respective integrin antibodies (α3, β1, α6 and β4).
It is known that CD151 associates strongly with α3- and α6-integrin subsets in several cell types including epithelial cells and hematopoietic cells. To investigate whether CD151 in HaCaT and HSC5 interacted with these integrins, immunoprecipitation assay was performed. The bands of α3-, α6-, β1- and β4-integrin subunits were detected respectively (Figure 1c), demonstrating that CD151 in these cells formed complexes with the laminin-binding integrin subsets.
Dissociation of CD151 and Re-localization of Laminin-Binding Integrins Under Migratory Stimulation
To understand the roles of CD151 in tumor migration, HSC5 was treated with EGF at the concentration of 10 ng/ml for 24 h under serum-starved condition (Figure 2a–d). HSC5 expressed EGF-receptor at high level (data not shown), and we confirmed that HSC5 was actually stimulated through EGF by detecting the internalization of phosphotyrosine (Figure 2a). Under non-stimulatory condition, 11G5a-recognized CD151 was localized at cell–cell boundaries and basal contacts, and also in the cytoplasm (Figure 2b, left). 14A2-recognized CD151 was localized at membranous regions and basal contacts, but less significant in the cytoplasm (Figure 2b, right). It is known that up to 66% of total cellular CD151 is localized at cytoplasmic compartments in HUVEC cells.30 Therefore, CD151 in HSC5 is thought to be localized both at cell–cell boundaries and in the cytoplasm, and cytoplasmic CD151 can be detected by 11G5a more clearly. By EGF stimulation, both 11G5a-recognized and 14A2-recognized CD151 became dispersed markedly in the cytoplasm, and the re-distribution patterns of 11G5a and 14A2 were almost identical (Figure 2c). In Western blot analysis, neither the band intensity nor the molecular size of CD151 in EGF-stimulated HSC5 was changed (data not shown), excluding the possibility that diffuse cytoplasmic stainings reflected cleaved or newly produced CD151 protein within 24 h by EGF stimulation.
Re-distribution of CD151 and integrins by EGF stimulation. (a) HSC5 cells were cultured with or without 10 ng/ml EGF under serum-starved condition. At 24 h after EGF stimulation, the cells were fixed and stained with mAb against phosphotyrosine. Phosphotyrosine is predominantly localized at focal adhesions in the cells without EGF stimulation whereas it is internalized in stimulated cells. (b) Distribution of CD151 in the cells without EGF stimulation. 11G5a-recognized CD151 is detected both at membranous regions and in the cytoplasm (left), whereas 14A2-recognized CD151 is rather restricted to membranous regions (right). (c) Internalization of CD151 by EGF stimulation. 11G5a (green)-recognized CD151 and 14A2 (red)-recognized CD151 are concomitantly internalized (merged image). (d) Re-distribution of α3-, β1-, α6- and β4-subunits by EGF stimulation. In the cells without EGF stimulation, α3- and β1-subunits are localized at membranous regions, whereas α6- and β4-subunits are localized at basal areas (left panels). After EGF stimulation, α3- and β1-subunits are internalized whereas α6- and β4-subunits are re-localized to basolateral edges (right panels). Bars indicate 10 μm for fluorescent cytostainings.
Next, we examined the re-distribution of α3β1- and α6β4-integrins by EGF stimulation. In control non-stimulating HSC5, α3- and β1-subunits were localized mainly at cell–cell boundaries, and α6- and β4-subunits were detected mainly at basal regions (Figure 2d, left panels). When the cells were stimulated by EGF, the intensities of cytoplasmic stainings increased in α3- and β1-subunits, whereas α6- and β4-subunits were re-localized from basal to basolateral regions (Figure 2d, right panels). Collectively, it was suggested that CD151 was dissociated dynamically from adhesive sites in response to EGF stimulation, and that laminin-binding integrins were re-localized concomitantly with CD151 dissociation.
Preparation of HSC5CD151− Using siRNA
To investigate the roles of CD151 in integrin-dependent cell–ECM interaction, siRNA against CD151 was transfected into HSC5 (HSC5CD151−) (Figure 3a–c). Non-silencing siRNA-transfected HSC5 cells (control HSC5) were prepared for comparison. By FACS analysis, the expression level of CD151 in HSC5CD151− was almost completely negative, which was examined by 11G5a and 14A2 antibodies, respectively (Figure 3a). In contrast, the surface expression level of another tetraspanin CD9 in HSC5CD151− was as highly positive as that in control HSC5. By fluorescence cytostainings, neither intercellular nor cytoplasmic stainings were detected in HSC5CD151− cells. CD9 stainings at intercellular boundaries were detected clearly in both control HSC5 and in HSC5CD151− (Figure 3b). Quantitative RT-PCR revealed that the expression level of CD9 was not significantly changed in HSC5CD151− (Figure 3c). We also confirmed the knockdown of CD151 by Western blot, in which neither the band intensity nor the molecular size of CD9 was changed (data not shown). The results strongly suggested that the knockdown of CD151 did not change CD9 expression in this cell. However, further study is required to understand whether the function of CD9 is affected by CD151 knockdown through either direct or indirect mechanism.
Knockdown of CD151 in HSC5. (a) Expression of CD151 and CD9 is examined by FACS. In control HSC5, CD151 and CD9 are clearly expressed. In HSC5CD151−, the expression of CD151 is almost completely suppressed, but the expression level of CD9 is not significantly changed. (b) By fluorescent cytostainings, CD151 are clearly detected by 11G5a and 14A2 mAbs in control HSC5 (left panels), whereas neither membranous nor cytoplasmic CD151 are detected in HSC5CD151− (right panels). The membranous stainings for CD9 are observed both in control HSC5 and HSC5CD151−. Inset: merged image with DAPI. (c) Expression levels of CD151 and CD9 are examined by real-time RT-PCR. The band of CD151 is significantly weaker in HSC5CD151− than that in control HSC5. However, the bands of CD9 are clearly detected both in HSC5CD151− and control HSC5. The absolute level of each tetraspanin/GAPDH mRNA is shown in the graph. (d) Control HSC5 cells on Matrigel show close cell–cell contact and colony formation (upper left). HSC5CD151− cells become spindle-shaped and extend cellular cables (upper right). On laminin-coated plates, control-HSC5 cells are flat (lower left). HSC5CD151− cells show hemispherical feature with elongated cellular protrusions (lower right). (e) In control HSC5, actin filaments are stained at cell periphery (left). In HSC5CD151− cells, actin filaments form stress fibers (right). Bars indicate 10 μm for fluorescent cytostainings.
Morphological Alteration of HSC5CD151− on ECM
Control-HSC5 and -HSC5CD151− cells were cultured on Matrigel (from BD Biosciences) or laminin (from Sigma), and cell adhesion process was monitored using time-lapse video microscopy. Majority of control HSC5 attached firmly onto Matrigel in 8–9 h. The cells became polygonal in shape and contacted each other (Figure 3d, upper left). In contrast, HSC5CD151− cells adhered to Matrigel but intercellular contacts were not as tight as those in control cells. HSC5CD151− cells formed colonies in 17 h and connected each other by extending cellular cables (Figure 3d, upper right). On laminin-coated plates, morphological differences became significant in 8–9 h. Control-HSC5 cells became flat and adhered to laminin firmly (Figure 3d, lower left). HSC5CD151− cells did not become flat, instead, were hemispherical in shape and attached to laminin by extending protrusions (Figure 3d, lower right). Cytoskeletal stainings showed that, in HSC5CD151−, actin filaments formed stress fibers (Figure 3e). Significant morphological alteration indicated that tumor cell–ECM interaction was loosened by CD151 knockdown and that the presence of CD151 might be critical for stable adhesion of the cells to ECM.
Attenuated Surface Expression of Integrins in HSC5CD151− Cells
To clarify whether CD151 knockdown affects the expression levels of several integrin subunits on cell surface, FACS analysis was performed (Figure 4a and b). MFI of α3- and α6-subunits were shown to be markedly downregulated. MFI of β1- and β4-subunits were also downregulated, but high expression levels of these β-subunits were preserved, indicating that β-subunits might be indirectly affected by CD151 knockdown. The bands of these integrin subunits were clearly detected both in control HSC5 and HSC5CD151− by RT-PCR (Figure 4c). Real-time RT-PCR showed that the expression levels of these integrin subunits were not markedly changed although statistical significance was observed in α6-subunit (Figure 4d). It was reported that α3- and α6-integrins were clearly detected in the tissues of CD151-null mice.31 Taken together, the results suggested that these integrin subunits were not quantitatively downregulated but internalized in part by CD151 knockdown.
Reduced-surface expression and re-distribution of α3-, α6-, β1- and β4-integrins in HSC5CD151−. (a) The expression levels of α3, α6, β1 and β4 are reduced significantly in HSC5CD151−. (b) Quantification of reduced MFI of integrins in HSC5CD151− in FACS. Values are expressed as relative MFI±s.e. (*P<0.05, **P<0.01). (c) The expression of α3, α6, β1 and β4 are examined by RT-PCR. The band intensities of these integrins are not significantly different between control HSC5 and HSC5CD151−. (d) The absolute level of each integrin/GAPDH mRNA is examined by real-time RT-PCR. The value of α6-subunit in control HSC5 is 0.67±0.01 and in HSC5CD151− is 0.48±0.01 (P<0.01), that of β1-subunit in control HSC5 is 0.73±0.01 and in HSC5CD151− is 0.99±0.1 (P=0.1). Other subunits demonstrate almost identical mRNA expression levels between control HSC5 and HSC5CD151−. (e) Re-distribution of integrin subunits in HSC5CD151−. In α3- and β1-subunits, membranous stainings become unclear, and strong perinuclear stainings are demonstrated. In α6- and β4-subunits, diffuse basal stainings are disappeared, and they are detected either in perinuclear regions or basolateral edges. Bars indicate 10 μm for fluorescent cytostainings.
Re-Distribution of Integrin Subunits in HSC5CD151− Cells
Previous studies have elucidated the importance of CD151 in the stabilization of laminin-binding integrins. However, the status of the integrins in the absence of CD151 has not been well documented, because mutant CD151 may not dissociate from integrin complex and site-specific CD151 antibodies may not inhibit the recruitment of integrin-free CD151 to adhesion complex. We compared the localization of α3-, α6-, β1- and β4-subunits between HSC5CD151− and control HSC5 on laminin. In control HSC5, the expression patterns of these integrin subunits were similar to those in original HSC5, as shown in Figure 2d. In contrast, in HSC5CD151−, staining patterns of the integrins were significantly changed. The accumulation of α3- and β1-subunits at cell–cell boundaries became unclear, and perinuclear cytoplasmic stainings were clearly demonstrated (Figure 4e, left). α6- and β4-subunits were re-distributed from basal sites to perinuclear regions and basolateral edges (Figure 4e, right). These findings supported the notion that CD151 may stabilize α3β1-integrin by strengthening high-affinity conformation of the heterodimer to bind firmly to laminin.32 The internalization of α3β1-integrin and basolateral accumulation of α6β4-integrin bore resemblance in part the feature of EGF-induced migratory form (Figure 2d) in which CD151 was dissociated from adhesion complexes.
Cellular Adhesion Strength was Attenuated by CD151 Knockdown
To understand whether integrin-dependent cell-binding ability onto laminin is affected by CD151 knockdown, we investigated the adhesion strength of HSC5CD151− cells (Figure 5a–c). In adhesion assay, it was shown that both control-HSC5 and -HSC5CD151− cells adhered onto laminin at almost similar levels (Figure 5a). In contrast, in detachment assay, it was shown that the adhesion strength in HSC5CD151− cells was significantly attenuated compared with that in control HSC5 (P<0.01; Figure 5b). Centrifugal force-based detachment assay also demonstrated the impaired adhesion strength of HSC5CD151− cells (P<0.01; Figure 5c). The results suggested that CD151 might not be indispensable for tumor cell adhesion to ECM, but that CD151 might be critical for supporting strong interaction between tumor cells and ECM by mediating the localization and conformation of its partner integrin subsets.
Adhesion strength onto laminin is attenuated by CD151 knockdown. (a) In static adhesion assay, no significant differences are demonstrated in a number of adherent cells between control HSC5 and HSC5CD151−. (b) In trypsin/EDTA-based detachment assay, adhesion strength of HSC5CD151− is shown to be significantly weaker than that in control HSC5 (**P<0.01). (c) In centrifugal force-based assay, 50% of HSC5CD151− cells are detached by 500 g centrifugation, whereas more than 80% of control-HSC5 cells keep adhesion to laminin by 500 and 750 g centrifugation (**P<0.01).
The Expression of MMPs was Suppressed by CD151 Knockdown
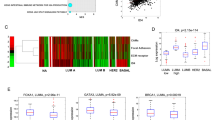
In tumor invasion, proteolytic activity is crucial for destruction and remodeling of surrounding stroma. MMPs are the key enzymes responsible for tumor invasion, controlled in part by integrin-mediated pathways.33 Recently, it was reported that CD151 was co-immunoprecipitated with MMP7 in rectal carcinoma cells.34 To understand whether CD151 mediates MMPs production in epidermal carcinoma cells at transcriptional level, we did quantitative RT-PCR. The expression levels of MMP2, MMP7 and MMP9 were downregulated in HSC5CD151−, and statistical significance was detected in MMP7 and MMP9 (Figure 6a and b). In Western blot analysis, the bands of MMP2, MMP9 and MMP7 were detectable in the lysate of control HSC5 and HSC5CD151−, and the band intensities of these MMPs were slightly stronger in the former than the latter (Figure 6c). We investigated further the secretion levels of MMPs by zymography. A total of 10 μl of CM from 1 × 105 cells for 48 h were added onto the gel. The bands of MMP2 and MMP9 were clearly detected both in the CM of control HSC5 and HSC5CD151−. However, the band intensities were weaker in the latter group (Figure 6d). The band of MMP7 was detected in the CM of control HSC5 concentrated 10 times but not in that of HSC5CD151− (Figure 6e, upper panels). Since the expression level of MMP7 in HSC5 was rather low, we examined whether MMP7 secretion was really suppressed by CD151 knockdown in a different cell line that produced endogenous MMP7 abundantly. An ovarian carcinoma cell HTOA was CD151 positive (data not shown) and produced significant amount of MMP7. Control HTOA was shown to secrete MMP7 clearly. In contrast, MMP7 band was almost invisible in HTOACD151− (Figure 6e, lower panels). Therefore, we carried out immunoprecipitation assay to understand whether CD151 interacts directly with MMPs. MMP7 was co-immunoprecipitated with CD151. MMP2 and MMP9 did not show significant bands (Figure 6f). Since it is known that MMP7 is often detected specifically in the tumor cells of invasive front,35, 36 we examined the localization of MMP7 in HSC5 cells on laminin. In the colonies that formed stable cell–cell adhesion, MMP7 was faintly distributed in peri-nuclear area in the cytoplasm (data not shown). In contrast, in migrating single cells, MMP7 was clearly expressed and was colocalized with CD151 in peri-nuclear cytoplasm and at the leading edges (Figure 6g, arrowheads). Collective data suggested that CD151 might contribute to effective tumor proteolytic activities by controlling MMPs production and secretion. Especially, the direct interaction between CD151 and MMP7 might be critical in integrin-mediated proteolysis.
Downregulation of MMPs by CD151 knockdown. (a) The expression of MMP2, MMP7 and MMP9 mRNAs are examined by RT-PCR. The band intensities of these MMPs are significantly weaker in HSC5CD151− cells than in control-HSC5 cells. (b) The absolute level of each MMP/GAPDH mRNA is compared between control HSC5 and HSC5CD151− by real-time RT-PCR. Downregulation of MMP7 and MMP9 mRNAs in HSC5CD151− is statistically significant (*P<0.05, **P<0.01). (c) Western blot analysis of MMP2, MMP7 and MMP9 is carried out using cell lysate samples. Significant bands are detected in control-HSC5 lysate, whereas the band intensities of MMPs in HSC5CD151− are rather weak. (d) Gelatin zymography is performed using 10 μl of CM from each sample electrophoresed onto the gel containing 0.1% gelatin. MMP2 and MMP9 are clearly detected both in control-HSC5 and -HSC5CD151− groups, but the band intensities in HSC5CD151− are weaker than those in control HSC5. (e) Casein zymography is carried out using 10 μl of 10 times concentrated CM from HSC5 samples electrophoresed onto the gel containing 0.1% casein. MMP7 in control HSC5 is detected as pro-enzymatic form (upper left). MMP7 in control HTOA is detected as both pro-enzymatic and active forms (lower left). In the CM of control HTOA on Matrigel, active form of MMP7 is dominant. In HSC5CD151− and HTOACD151− samples, neither the band of pro-enzymatic form nor active form of MMP7 is detected (upper and lower right). Plain, no matrix: Ln5, coated with laminin-527: MG, coated with Matrigel. (f) CD151 was immunoprecipitated and precipitates were electrophoresed, then probed with each MMP mAbs. The band of MMP7 (29 kDa) is detected. The bands of MMP2 (72 kDa) and MMP9 (92 kDa) are not detectable. *Cross reactive protein. (g) Migratory HSC5 cell stained with CD151 (clone 14A2, green) and MMP7 (red) is shown. CD151 and MMP7 are diffusely stained in the perinuclear cytoplasm and accumulated at the leading edges (upper panel, arrow heads). High magnification of lamellipodia (dotted rectangles of upper panel) is shown in lower panel. CD151 and MMP7 are colocalized at the leading edges (arrow heads).
Discussion
The principal role of CD151 in epithelial cells under physiological condition is thought to stabilize cell–ECM adhesion system by associating with α6β4-integrins at hemidesmosomes and tethering α3β1 at cell–cell boundaries.17, 18 In β4-integrin-deficient epidermolysis bullosa PA-JEB keratinocytes, CD151 and α3β1-integrin were clustered together at basal sites, and when transfected with β4-integrin, CD151 bound more strongly to α6β4 than to α3β1 in vitro,19 which indicates that CD151 contributes to the stabilization of hemidesmosomes for maintaining proper cell–ECM interaction. In addition to the studies on adhesive cells, several studies on hematopoietic cells and immune cells have elucidated the importance of CD151 in physiological activities of non-polarized cells.37 CD151 in these cells are co-immunoprecipitated with several types of integrin repertoire, and cellular motility is affected by the antibodies against CD151-integrin complexes. Therefore, CD151 in non-polarized cells is involved at least in cell–cell and cell–matrix events such as lymphocyte–endothelium interaction and platelet aggregation. These events suggest that CD151 participates not only in cell stabilization but also in migratory process. In tumor cells, the expression patterns and dynamics of α3β1- and α6β4-integrins can be heterogonous depending on tumor cell types and microenvironments.38 Currently, the importance of CD151 in tumor migration in association with cytological localization of laminin-binding integrins and proteolytic activities is not well understood.
In the present study, we demonstrated that (i) The expression levels of CD151 and laminin-binding integrins in an epidermal carcinoma cell HSC5 were higher than those in an immortalized keratinocyte HaCaT; (ii) Knockdown of CD151 resulted in cytoskeletal rearrangements, impaired adhesion strength and re-localization of laminin-binding integrin subunits and (iii) The expression levels of MMP2, MMP7 and MMP9 were downregulated by CD151 knockdown and CD151 was directly associated with MMP7. These results suggested that the presence of CD151 was critical for stabilizing the localization of laminin-binding integrins and that the upregulation of CD151 was favorable for tumor cells to exert strong proteolytic activity by producing and secreting MMPs.
Depletion of CD151 from tumor cells resulted in attenuated adhesion strength in this experiment, which was consistent with those reported in the platelets of CD151-null mice and of NIH 3T3 cells transfected with mutant CD151.14, 31 In tumor cells, it remains unclear whether the dissociation or depletion of CD151 actually impairs cell motility, or increases the ability of cell motility. Varied results have been reported; treatment of HaCaT with a CD151 monoclonal antibody induced cell migration,24 and the melanocytes transfected with CD151–siRNA enhanced cell motility,39 whereas CD151-silenced A431 cells showed impaired cell motility.40 In the present study, we demonstrated the downregulation and re-distribution of laminin-binding integrin subunits by CD151 knockdown. Although the cell types and experiment systems used in the studies were different, it is indicated that the dissociation of CD151 from adhesion complexes could induce the cytoskeletal rearrangement and morphological alteration, which could be interpreted as a feature of increased motility. However, attenuated cell–ECM interaction and proteolytic activities may represent a feature of reduced migratory potential. Since tumor cells can change contact sites and interaction strength sequentially, the role of CD151 in tumor cells should be considered in the context of cell dynamics and tumor microenvironments.
The contribution of some tetraspanins including CD151 to MMPs expression has been demonstrated in previous works.34, 41 In this study, the expression of MMP2, MMP7 and MMP9 were downregulated by CD151 knockdown (Figure 6a–e). Immunoprecipitation assay and double stainings demonstrated that CD151 was colocalized with MMP7 (Figure 6f and g). The finding, together with the notion that MMP7 is expressed preferentially at the leading edge of tumor invasive sites,34, 35 indicates that CD151–MMP7 complex may play an important role in integrin-mediated proteolytic activity during tumor invasion. MMP2 and MMP9 were not co-immunoprecipitated with CD151, but we cannot exclude the possibility that these MMPs may associate with CD151 either directly or indirectly. A recent study using melanoma cells showed that transfected CD151 activated AP-1 sites in the MMP9 gene promoter by c-Jun binding to AP-1.41 It should be further investigated whether CD151 transports specific MMP(s) to integrin–ligand contacting sites for effective proteolysis.
Currently, it remains to be enigmatic whether the upregulation of integrins reported in several aggressive tumors actually strengthen the integrin–ligand interaction or accelerate hemidesmosome-independent migration.42, 43 Tumor cells may strengthen cell–ECM interaction to invade,43 but may also loosen the interaction to relocate from primary site. To cope with changeable microenvironments, the scaffold molecules like CD151 might accelerate and decelerate outside–in signalings swiftly between integrins and ligands. It is necessary to extend the studies on CD151, other tetraspanins and associated integrins in relation to cell dynamics such as migration and invasion in progressive tumors.
References
Stepp MA, Spurr-Michaud S, Tisdale A, et al. Alpha6 beta4 integrin heterodimer is a component of hemidesmosomes. Proc Natl Acad Sci USA 1990;87:8970–8974.
Koster J, van Wilpe S, Kuikman I, et al. Role of binding of plectin to the integrin beta4 subunit in the assembly of hemidesmosomes. Mol Biol Cell 2004;15:1211–1223.
DiPersio CM, Hodivala-Dilke KM, Jaenisch R, et al. Alpha3 beta1 integrin is required for normal development of the epidermal basement membrane. J Cell Biol 1997;137:729–742.
Tennenbaum T, Weiner AK, Belanger AJ, et al. The suprabasal expression of alpha6 beta4 integrin is associated with a high risk for malignant progression in mouse skin carcinogenesis. Cancer Res 1993;53:4803–4810.
Mercurio AM, Rabinovitz I . Towards a mechanistic understanding of tumor invasion – lessons from the alpha6beta 4 integrin. Semin Cancer Biol 2001;11:129–141.
Natali PG, Nicotra MR, Bartolazzi A, et al. Integrin expression in cutaneous malignant melanoma: association of the alpha 3/beta 1 heterodimer with tumor progression. Int J Cancer 1993;54:68–72.
Giannelli G, Bergamini C, Fransvea E, et al. Human hepatocellular carcinoma (HCC) cells require both alpha3beta1 integrin and matrix metalloproteinases activity for migration and invasion. Lab Invest 2001;81:613–627.
Liebert M, Washington R, Wedemeyer G, et al. Loss of co-localization of alpha 6 beta 4 integrin and collagen VII in bladder cancer. Am J Pathol 1994;144:787–795.
Pignatelli M, Cardillo MR, Hanby A, et al. Integrins and their accessory adhesion molecules in mammary carcinomas: loss of polarization in poorly differentiated tumors. Hum Pathol 1992;23:1159–1166.
Hashida H, Takabayashi A, Tokuraha T, et al. Integrin alpha3 expression as a prognostic factor in colon cancer: association with MRP-1/CD9 and KAI1/CD82. Int J Cancer 2002;97:518–525.
Fitter S, Tetaz TJ, Berndt MC, et al. Molecular cloning of cDNA encoding a novel platelet–endothelial cell tetraspan antigen, PETA-3. Blood 1995;86:1348–1355.
Zhang XA, Bontrager AL, Hemler ME . Transmembrane-4 superfamily proteins associate with activated protein kinase C (PKC) and link PKC to specific beta(1) integrins. J Biol Chem 2001;276:25005–25013.
Kohno M, Hasegawa H, Miyake M, et al. CD151 enhances cell motility and metastasis of cancer cells in the presence of focal adhesion kinase. Int J Cancer 2002;97:336–343.
Lammerding J, Kazarov AR, Huang H, et al. Tetraspanin CD151 regulates alpha6beta1 integrin adhesion strengthening. Proc Natl Acad Sci USA 2003;100:7616–7621.
Yauch RL, Berditchevski F, Harler MB, et al. Highly stoichiometric, stable, and specific association of integrin alpha3beta1 with CD151 provides a major link to phosphatidylinositol 4-kinase, and may regulate cell migration. Mol Biol Cell 1998;9:2751–2765.
Kazarov AR, Yang X, Stipp CS, et al. An extracellular site on tetraspanin CD151 determines alpha 3 and alpha 6 integrin-dependent cellular morphology. J Cell Biol 2002;158:1299–1309.
Berditchevski F . Complexes of tetraspanins with integrins: more than meets the eye. J Cell Sci 2001;114:4143–4151.
Stipp CS, Kolesnikova TV, Hemler ME . EWI-2 regulates alpha3beta1 integrin-dependent cell functions on laminin-5. J Cell Biol 2003;163:1167–1177.
Sterk LM, Geuijen CA, Oomen LC, et al. The tetraspan molecule CD151, a novel constituent of hemidesmosomes, associates with the integrin alpha6beta4 and may regulate the spatial organization of hemidesmosomes. J Cell Biol 2000;149:969–982.
Penas PF, Garcia-Diez A, Sanchez-Madrid F, et al. Tetraspanins are localized at motility-related structures and involved in normal human keratinocyte wound healing migration. J Invest Dermatol 2000;114:1126–1135.
Testa JE, Brooks PC, Lin JM, et al. Eukaryotic expression cloning with an antimetastatic monoclonal antibody identifies a tetraspanin (PETA-3/CD151) as an effector of human tumor cell migration and metastasis. Cancer Res 1999;59:3812–3820.
Hashida H, Takabayashi A, Tokuhara T, et al. Clinical significance of transmembrane 4 superfamily in colon cancer. Br J Cancer 2003;89:158–167.
Gesierich S, Paret C, Hildebrand D, et al. Colocalization of the tetraspanins, CO-029 and CD151, with integrins in human pancreatic adenocarcinoma: impact on cell motility. Clin Cancer Res 2005;11:2840–2852.
Chometon G, Zhang ZG, Rubinstein E, et al. Dissociation of the complex between CD151 and laminin-binding integrins permits migration of epithelial cells. Exp Cell Res 2006;312:983–995.
Geary SM, Cambareri AC, Sincock PM, et al. Differential tissue expression of epitopes of the tetraspanin CD151 recognised by monoclonal antibodies. Tissue Antigens 2001;58:141–153.
Boukamp P, Petrussevska RT, Breitkreutz D, et al. Normal keratinization in a spontaneously immortalized aneuploid human keratinocyte cell line. J Cell Biol 1988;106:761–771.
Goldfinger LE, Stack MS, Jones JC . Processing of laminin-5 and its functional consequences: role of plasmin and tissue-type plasminogen activator. J Cell Biol 1998;141:255–265.
Lau LM, Wee JL, Wright MD, et al. The tetraspanin superfamily member CD151 regulates outside–in integrin alphaIIbbeta3 signaling and platelet function. Blood 2004;104:2368–2375.
Raymond K, Kreft M, Janssen H, et al. Keratinocytes display normal proliferation, survival and differentiation in conditional beta4-integrin knockout mice. J Cell Sci 2005;118:1045–1060.
Sincock PM, Fitter S, Parton RG, et al. PETA-3/CD151, a member of the transmembrane 4 superfamily, is localised to the plasma membrane and endocytic system of endothelial cells, associates with multiple integrins and modulates cell function. J Cell Sci 1999;112:833–844.
Wright MD, Geary SM, Fitter S, et al. Characterization of mice lacking the tetraspanin superfamily member CD151. Mol Cell Biol 2004;24:5978–5988.
Nishiuchi R, Sanzen N, Nada S, et al. Potentiation of the ligand-binding activity of integrin alpha3beta1 via association with tetraspanin CD151. Proc Natl Acad Sci USA 2005;102:1939–1944.
DiPersio CM, Shao M, Di Costanzo L, et al. Mouse keratinocytes immortalized with large T antigen acquire alpha3beta1 integrin-dependent secretion of MMP-9/gelatinase B. J Cell Sci 2000;113:2909–2921.
Shiomi T, Inoki I, Kataoka F, et al. Pericellular activation of proMMP-7 (promatrilysin-1) through interaction with CD151. Lab Invest 2005;85:1489–1506.
Shiomi T, Okada Y . MT1-MMP and MMP-7 in invasion and metastasis of human cancers. Cancer Metastasis Rev 2003;22:145–152.
Kerkelä E, Saarialho-Kere U . Matrix metalloproteinases in tumor progression: focus on basal and squamous cell skin cancer. Exp Dermatol 2003;12:109–125.
Tarrant JM, Robb L, van Spriel AB, et al. Tetraspanins: molecular organisers of the leukocyte surface. Trends Immunol 2003;24:610–617.
Janes SM, Watt FM . New roles for integrins in squamous-cell carcinoma. Nat Rev Cancer 2006;6:175–183.
Garcia-Lopez MA, Barreiro O, Garcia-Diez A, et al. Role of tetraspanins CD9 and CD151 in primary melanocyte motility. J Invest Dermatol 2005;125:1001–1009.
Winterwood NE, Varzavand A, Meland MN, et al. A critical role for tetraspanin CD151 in alpha3beta1 and alpha6beta4 integrin-dependent tumor cell functions on laminin-5. Mol Biol Cell 2006;17:2707–2721.
Hong IK, Jin YJ, Byun HJ, et al. Homophilic interactions of tetraspanin CD151 up-regulate motility and matrix metalloproteinase-9 expression of human melanoma cells through adhesion-dependent c-Jun activation signaling pathways. J Biol Chem 2006;281:24279–24292.
Avizienyte E, Wyke AW, Jones RJ, et al. Src-induced de-regulation of E-cadherin in colon cancer cells requires integrin signalling. Nat Cell Biol 2002;4:632–638.
Dajee M, Lazarov M, Zhang JY, et al. NF-kappaB blockade and oncogenic Ras trigger invasive human epidermal neoplasia. Nature 2003;421:639–643.
Acknowledgements
We express our greatest sorrow over the death of Professor Dr H Ishikura, our beloved and respected director of Department of Molecular Pathology. We thank Dr T Takenouchi for providing materials, Professor Dr NE Fusenig for providing HaCaT cell and Dr N Morinaga and Dr Y Nakayama for expert help with real-time RT-PCR system and laser scan microscopy, respectively. This work was supported by grants in aid from the Ministry of Education, Culture, Sports, Science and Technology of Japan (18590320), Inohana Foundation of Chiba University and Sagawa Cancer Research Foundation (to MF).
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website (http://www.laboratoryinvestigation.org)
Rights and permissions
About this article
Cite this article
Hasegawa, M., Furuya, M., Kasuya, Y. et al. CD151 dynamics in carcinoma–stroma interaction: integrin expression, adhesion strength and proteolytic activity. Lab Invest 87, 882–892 (2007). https://doi.org/10.1038/labinvest.3700657
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700657
Keywords
This article is cited by
-
FAK-ERK activation in cell/matrix adhesion induced by the loss of apolipoprotein E stimulates the malignant progression of ovarian cancer
Journal of Experimental & Clinical Cancer Research (2018)
-
Characterization of the early molecular changes in the glomeruli of Cd151 −/− mice highlights induction of mindin and MMP-10
Scientific Reports (2017)
-
Remodeling of extracellular matrix by normal and tumor-associated fibroblasts promotes cervical cancer progression
BMC Cancer (2015)
-
CD151 in cancer progression and metastasis: a complex scenario
Laboratory Investigation (2014)
-
Application of RNA-Seq transcriptome analysis: CD151 is an Invasion/Migration target in all stages of epithelial ovarian cancer
Journal of Ovarian Research (2012)