Abstract
New methods of elucidating the ligand-binding activity of receptors could improve our understanding of receptor function, key events they control, and their presence in normal and pathological states. We describe a method for visualizing receptor–ligand binding in cells and tissues that substitutes fluorescein for radioactive labels, and detects receptor bound, fluoresceinated ligand with an antifluorescein/horseradish peroxidase amplification system. Receptor-bound ligand is then visualized by light microscopy against a standard hemotoxylin-stained background of cell structure. Quantitative versions of the assay provide an apparent dissociation constant and number of receptors per cell at saturation in cell or tissue specimens. Receptors examined include the folate receptor, bombesin peptide-binding receptors, the epidermal growth factor receptor, the neuropeptide Y receptor, the asialoglycoprotein receptor, and RGD peptide-binding integrins. Using fluoresceinated versions of molecules, we show the method can visualize and quantitate receptor-bound ligands in cell culture monolayers and animal tissue specimens. Ligand binding to receptors present in tissues was visualized in normal and pathological samples of human tissue microarrays. The enzyme-amplified detection of receptor-bound fluoresceinated ligand is a simple and nonradioactive-based method that provides information on the receptor activity in tissue specimens.
Similar content being viewed by others
Main
Visualizing and quantifying the binding of ligands to receptors expressed in tissues is crucial to understanding the role receptors play and the events they control in normal and pathological tissues. Though antibody based immunohistochemistry provides valuable information about the distribution of immunoreactive structures (epitopes), it does not provide information on the presence or activity of ligand binding sites (Figure 1a). In a typical assay configuration, information about receptor distribution is obtained through antibody-mediated biotin–streptavidin interaction amplified by conjugated enzymes (eg horseradish peroxidase (HRP)).1, 2, 3 Receptor autoradiography requires the use of radiolabeled ligands, with the attendant issues of frequent radiochemical synthesis and handling of radioactivity (Figure 1b). In addition, the distribution of radioactivity is obtained on a blank background and must be superimposed on a second image, to visualize ligand binding against a background of cell structure.
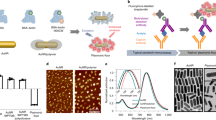
Methods of visualizing receptors. (a) Immunohistochemistry. A biotinylated antibody binds to a structural determinant on a 7TM receptor (or other type of structure). Visualization is with an avidin–enzyme conjugate. (b) Radioreceptor autoradiography. A radioactive peptide binds in the pocket of a 7TM receptor. Radioactivity is visualized with a photographic film. (c) Fluorescein hapten visualization (FHV) and fluorescein hapten assay (FHA). With FHV, fluorescein replaces the radioisotope and detection is with an antifluorescein HRP. With FHA, the fluoresceinated ligand binds the receptor, the specimen is digested, and fluorescein is quantified by an immunoassay for fluorescein. A slide chamber over the specimen contains the lysis buffer.
We hypothesized that it would be possible to substitute fluoresceinated ligands for radioactive ligands, and detect bound fluorescein using the HRP amplification system to achieve a high sensitivity. We further hypothesized that fluorescein hapten detection methodologies could be configured as a cell-based method for imaging receptors (fluorescein hapten visualization (FHV)) or as a cell-based assay for bound ligand (fluorescein hapten assay (FHA)) as shown in Figure 1c. In the visualization method (FHV), a cell monolayer or tissue specimen is incubated with the receptor-binding ligand in a slide chamber and bound ligand visualized by microscopy. In the assay method (FHA), the specimen is exposed to the fluoresceinated ligand as in FHV, but rather than proceeding with microscopy the specimen is solubilized and immunoreactive fluorescein quantified by a fluorescein immunoassay. When the number of cells is measured in the sample, the bound ligand per cell can be obtained. FHV and FHA provide information about ligand-binding receptors that can be used with different types of specimens and different types of receptor-binding ligands (small molecule inhibitors and active site binders, peptides and proteins).
Materials and methods
Peptide Synthesis
Peptide synthesis using Fmoc chemistry was by the Tufts Core Facility (Boston, MA, USA). A fluoresceinated RGD peptide (GSSK(Fl)GGGCRGDC, ‘Fl-RGD’ and a scrambled version of the fluoresceinated RGD peptide (GSSK(Fl)GGGCDRGC, ‘Fl-scrRGD’) were obtained and cyclized by oxidation (room temperature, 24?h, less than 0.5?mg/ml peptide in 0.1?M ammonium bicarbonate). A bombesin peptide ((Fl)BRRRGQRL GNQWAVGHLM, ‘bombesin peptide (Fl-BN)’) and scrambled bombesin peptide ((Fl)BRRRGQMLGNHLAVGQWR, Fl-ScrBN) were also made. Fluoresceinated and nonfluroesceintated neuropeptide Y (NPY) was from Anaspec (San Jose, CA, USA) and are ‘fluoresceinated NPY (Fl-NPY)’ and ‘NPY, ‘respectively. The NPY sequence was YPSKPDNPGEDAPAEDMARYYSALRHYINLITRQRY with and without a N-terminal fluorescein. Fluoresceinated and nonfluoresceinated human epidermal growth factor (EGF) was from Invitrogen (Carlsbad, CA, USA) and are denoted Fl-EGF and EGF, respectively. The sequences were NSDSECPLSHDGYCLHDGVCMYIEALDKYACNCVVGYIGERCQYRDLQWWELR with and without an N-terminal fluorescein. Asialofetuin was from Sigma (Sigma-Aldrich, St Louis, MO, USA) and fluoresceinated by reaction with fluorescence isothiocyanate (FITC) to yield ‘fluoresceinated asialofetuin (Fl-ASF)’. A fluoresceinated folate was synthesized by reacting folate with 1,8-bis(methylamino)-3,6-dioxaoctane, to form an amino functionalized folate, followed by reaction with FITC, see Supplementary materials and is denoted ‘Fl-Fol’.
Cells
BT-20 and MDA-MB-468 were from ATCC (Manassas, VA, USA) and grown according to manufacturer's instructions. Cells were cultured on a four chambers slide system (Lab Tek Products, Naperville, IL, USA), which contained the fluid over the specimen and which was removed to process the sample for microscopy.
Tissue Specimens
Tissues were mouse liver and pancreas and rat colon. Tissues were snap frozen, cut into 7?μm sections and placed on glass slides. Tissue microarrays were from Tristar (Tristar Technology, Bethesda, MD, USA).
Fluorescein Hapten Visualization
A brief fixation procedure (acetone at 4°C, 5?min) was used to obtain high quality micrographs against which the ligand binding could be visualized. Microscopy of frozen sections gave poorer quality micrographs than those obtained with fixed specimens. Tissue microarrays (Figure 4) employed a fixation procedure of the vendor. Fluoresceinated ligands were dissolved in PBS (10% rabbit serum, 0.3% sodium azide and 0.5% H2O2 to block the endogenous peroxidases) and then placed in the sample chamber (1?h, room temperature, in a humidity chamber). The slide chamber was then removed so that specimens could be processed. Unbound ligand was removed by washing three times with the PBS. An antifluorescein-HRP conjugate (Invitrogen, Carlsbad, CA, USA), diluted to 5?μg/ml into PBS (2% rabbit serum), was applied to the sample (40?min, room temperature). Enzyme activity was developed for 10–20?min using AEC substrate (DAKO, Carpinteria, CA, USA). Slices were counterstained with hemotoxylin. Tissue microarrays were fixed and handled like cultured cells or tissue specimen.
FHV with tissue microarrays. (a) Fl-RGD and Fl-scrRGD binding to normal breast and ductal carcinomas. An adjacent slice was stained with the nonspecific peptide Fl-scrRGD. Note reddish areas of Fl-RGD binding in the ductal adenocarinoma. (b) Summary of tissue microarray data with Fl-RGD. Numbers of positive specimens are given followed by total numbers of specimens in that class. Thus, there were two positive specimens out of 10 total normal breast specimens. (c) Fl-BN and Fl-scrBN binding to normal pancreas and pancreatic ductal adenocarcinoma. (d) Summary of tissue microarray with Fl-BN.
Fluorescein Hapten Assay
Specimens were fixed as above, incubated with ligand, and washed free of unbound ligand as above for the fluorescein hapten receptor visualization method. The immunoassay for immunoreactive fluorescein was employed to quantitate bound fluorescein.4 To the slide chamber was added 500?μl of lysis buffer (PBS, 0.1% BSA, 0.1% Triton X-100, 1?mM 8-anilino-1-naphthalenesulfonic acid with 40?ng/ml of antifluorescein HRP. Incubation was for 2?h at room temperature. Duplicate 200?μl volumes were then transferred to a 96-well plate (MaxiSorp, Nunc) coated with 12.5?ng/ml FITC-labeled BSA (Sigma-Aldrich, St Louis, MO, USA) and incubated for 1?h at room temperature. Plates were then washed three times (PBS, 0.1% BSA, 0.1% Tween), to remove unreacted antifluorescein HRP, and bound peroxidase quantitated by absorbance at 650?nm using 200?μl of 3,3′,5,5′ tetramethylbenzidine-dihydrochloride after 30?min incubation at room temperature. The concentration of fluoresceinated ligand was determined from a standard curve of fluoresceinated ligand diluted in PBS, 0.1% BSA, 0.1% Tween.
For both FHV and FHA methods, receptor-specific ligand binding was determined from the binding of fluoresceinated nonreceptor-binding scrambled peptides (bombesin, RGD peptide) or by blocking binding with a nonfluoresceinated peptide or protein at 50-fold molar excess over the fluoresceinated ligand (folate, EGF, Asialofetuin, NPY).
To confirm the accuracy of FHA, cell-associated fluorescence was measured by fluorescein-activated cell sorter (FACS). Cells (BT-20 and MDA-MB-468) were trypsinized and collected by centrifugation (5?min at 1500?r.p.m.). They were fixed (suspended) in cold acetone for 5?min, collected by centrifugation and incubated with different concentrations of fluorescent ligands for 1?h at room temperature. Fluorescence was analyzed by FACS (FACScalibur, Becton Dickinson, Franklin Lakes, NJ, USA). The difference in mean fluorescence between the sample and the control was then plotted against the concentration of ligand. Data analysis yielded apparent dissociation constant (EC50), as described below, that could be compared with the apparent dissociation obtained by the FHA method.
To calculate the apparent dissociation constant (EC50) and number of binding sites per cell, the nonspecific binding was subtracted with either the scrambled peptide binding or blocking peptide method. The average of three determinations of bound ligand at each concentration were fit to a nonlinear mono-exponential binding isotherm equation using Prism4 software (GraphPad software Inc., San Diego, CA, USA); this yields an EC50 and the moles of binding sites per specimen at an infinite concentration of ligand. Cell number was determined as nuclei visualized after hemotoxylin stain using Image J software (NIH) on an adjacent slide. Dividing the moles of fluoresceinated ligand bound at saturation by the number of nuclei present gave the amount of ligand bound per cell. Uncertainties are the s.e. for each constant provided by GraphPad software. In all cases the coefficient of correlation was greater than 0.90, indicating a good fit of the data to single binding site model we employed.
Results
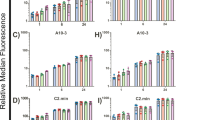
We initiated our experiments on the development of FHV and FHA methodologies with Fl-RGD, a cyclic peptide that binds to the αVβ3 integrin expressed on BT-20 tumor cells. Expression of αVβ3 on BT-20 cells was verified with an anti-αVβ3 antibody and FACS, data not shown. The cyclic RGD motif binds beta 3 integrins on a wide range of cell types including platelets, endothelial cells, tumor cells, smooth muscle cells and fibroblasts.5, 6, 7 The visualization of RGD-binding sites (FHV) is shown in Figure 2a, which can be compared to the staining obtained with the control peptide, Fl-scrRGD, the latter giving a distinctly lighter staining (Figure 2b). To confirm the visual impression of differential stain intensity, a quantitative assay for cell-associated peptide or FHA was performed over a range of peptide concentrations as shown in Figure 2c. Cells were exposed to peptide, washed, lysed, and cell-associated fluorescein determined using the fluorescein immunoassay.4 After subtracting the nonspecific binding component due to Fl-scrRGD, to obtain the specific binding of Fl-RGD, an apparent dissociation constant (EC50) and maximum binding was obtained for Fl-RGD. FHA values for the apparent dissociation constant (401?nM) and numbers of binding sites per cell (320?000) were obtained and are recorded with the FHA-generated values for other receptors (see below) in Table 1. In a second series of experiments, BT-20 cells were exposed to the Fl-RGD and Fl-scrRGD peptides, followed by determination of cell-associated fluorescence by FACS (Figure 2c and d). The apparent affinities based on cell-associated fluorescein obtained by immunoassay (401?nM) compared with that obtained by FACS (447?nM). Since FACS data were obtained as relative fluorescence, the amount of binding could not be determined.
Fluorescein hapten visualization (FHV) and fluorescent hapten assay (FHA) methods with cultured cells. Fluorescent hapten methodologies are shown using RGD peptides and integrin expressing BT-20 cells in (a–d). (a) The FHV with Fl-RGD. (b) The FHV with Fl-scrRGD. Fl-scrRGD gave a distinctly lighter stain than Fl-RGD. (c) The FHA of Fl-RGD and Fl-scrRGD peptides determined at different peptide concentrations. (d) Specific binding of Fl-RGD determined by FHA and FACS. Fluorescent hapten methodologies using Fl-EGF and growth factor expressing MDA-MB-468 cells are shown in (e–h). (e) The FHV with Fl-EGF. Arrows show high concentration of receptor at some faces of the membrane. (f) FHV with Fl-EGF and the EGF blocking peptide. EGF reduced binding of Fl-EGF. (g) The FHA of Fl-EGF at different peptide concentrations. (h) Specific binding determined of Fl-EGF determined by FHA and FACS.
The FHV and FHA methodologies were then applied to characterize the binding of fluoresceinated EGF (Fl-EGF) to MDA-MB-468 cells (Figure 2e–h). EGF binds to one of four members of the EGF receptor family, erbB1/Her1, which is the target of Erbitux (cetuximab). Unlike the Fl-RGD peptide, that bound uniformly over BT-20 cells, Fl-EGF concentrated at discrete points in the cell membrane (Figure 2e, arrows). Specific binding of Fl-EGF was obtained subtracting the binding blocked by EGF (compare Figure 2f and g). The FHA-generated apparent dissociation constant (EC50) for Fl-EGF and numbers of binding sites per cell are listed in Table 1. Again the apparent affinities for Fl-EGF binding obtained by FHA (51±29?nM) compared well with that obtained by FACS (31±3?nM).
Based on the ability of the FHA and FHV methods to provide information about the binding of peptides to receptors expressed on cultured cells, we applied these methods to four receptor/ligand combinations present in four different tissues. Receptors were chosen based on (i) the presence of substantial levels of receptors in the normal tissues of mice (or rats) and humans, (ii) the recognition of identical ligands by receptors in mice (or rats) and humans, and (iii) the ease of synthesizing fluoresceinated ligands with retention of binding activity. Four receptors meeting these criteria were the folate receptor in renal cells, the asialoglycoprotein receptor in hepatocytes, the NPY receptor in colonic epithelial cells, and the bombesin peptide-binding receptors in pancreatic acinar cells. (In mammals there are at least three pharmacological distinct receptors that bind bombesin-like peptides, which we refer to as ‘bombesin-binding receptors’ for simplicity.8, 9) Fluoresceinated ligands for these four receptors consisted of three types of molecules; a small organic molecule (Fl-folate), two peptides (Fl-NPY and Fl-BN), and a protein (Fl-ASF).
Folate Receptor
Figure 3a shows the FHV for Fl-Folate binding to a mouse kidney section from the cortical region. Displacement by folate (compare Figure 3a and b) indicated a receptor-specific binding component. The FHA data are shown in Figure 3c, with FHA-generated values for apparent dissociation constant and maximum binding given in Table 1. Folate receptors are found on many cell types but are particularly high in the proximal tubule of the kidney where they function to salvage folates.10
FHV and FHA methods with tissue specimens (a) FHV of Fl-folate binding to mouse kidney. (b) FHV with Fl-folate with binding blocked by folate. (c) FHA with Fl-folate at different peptide concentrations. (d) FHV of Fl-ASF binding to mouse liver. (e) FHV of Fl-ASF with binding blocked by asialofetuin. (f) FHV of Fl-ASF at different protein concentrations. (g) FHV of Fl-NPY binding to rat colon. (h) FHV of Fl-NPY with binding blocked by NPY. (i) FHA of Fl-NPY at different protein concentrations. (j) FHV of Fl-BN binding to mouse pancreas. (k) Binding of Fl-scrBN to pancreas. (l) FHA of Fl-BN binding.
Asialoglycoprotein Receptor
The FHV for Fl-ASF binding to a mouse liver section is shown in Figure 3d. Displacement by ASF (compare Figure 3d and e) provided the receptor-specific binding component. The FHA data is shown in Figure 3f, with FHA-generated apparent dissociation constant and maximum binding given in Table 1. The asialoglycoprotein receptor is found on hepatocytes11 and its expression used to measure hepatic function.12
NPY Receptor
The FHV for Fl-NPY binding to a tissue section from the descending colon of a rat is shown in Figure 3g. Displacement by NPY (compare Figure 3g and h) provided the receptor-specific binding component, with FHA data shown in Figure 3i and values recorded in Table 1. NPY receptors are found in many tissues of the gastrointestinal tract including the colon with the receptor expressed at particularly high levels on the mucosa, submucosa and myenteric plexus.13, 14, 15 NPY alters a variety of physiological functions with its effects on appetite control attracting recent attention.16
Bombesin Peptide-Binding Receptors
The FHV for Fl-BN binding to a tissue section from the tail of a mouse pancreas is shown in Figure 3j, with the binding of a scrambled peptide (Fl-scrBN) shown Figure 3k. FHA-generated apparent dissociation constants and maximum binding values are provided in Table 1. Bombesin-like peptides bind receptors upregulated in breast, prostate, and lung cancers.17 Bombesin-binding receptors are also present on acinar cells of the normal pancreas.17, 18, 19
Based on our results with cell monolayers and animal tissue sections, we applied the FHV method using the RGD peptides (Fl-RGD, Fl-scrRGD, Figure 2a–d) and the bombesin-like peptides (Fl-BN, Fl-scrBN, Figure 3j–l) to human tissues using commercially available microarrays. Tissue microarrays were handled as slides, with all tissue samples on the array simultaneously exposed to reagents of the FHV protocol. Figure 4a shows the binding of the RGD-like peptides to normal breast and ductal carcinoma specimens. The carcinoma bound the Fl-RGD peptide (pink color, arrows) but not the Fl-scrRGD peptide, while normal breast tissue failed to bind either peptide. The array consisted of 26 ductal carcinomas, 25 of which were evaluated, seven lobular carcinoma, six of which were evaluated, and 10 normal tissue specimens. Two samples were lost due to partial detachment. A cribriform carcinoma and a mucinous carcinoma were also not evaluated. The binding of Fl-RGD with the human breast cancer array had a sensitivity of 26/31 or 84% and specificity of 8/10 or 80%.
The FHV method using the bombesin-like peptides is shown in Figure 4c. Normal pancreatic tissue binds bombesin-like peptides while pancreatic cancers do not.17, 19, 20 The binding of Fl-BN, and lack of binding of Fl-scrBN, was apparent with normal pancreatic tissue while pancreatic ductal adenocarcinoma failed to bind either peptide. The pancreatic array consisted of 13 ductal adenocarcinomas and five normal samples, all of which were evaluated with results tabulated in Figure 4d. With this limited number of samples, the binding of Fl-BN to the pancreatic cancer array had a sensitivity and specificity of 100%. For tissue microarrays, the presence of receptor binding was deduced from the stain intensity of the FHV method because we lacked a device comparable to slide chamber to confine solutions over individual tissue specimens and perform the FHA assay (see Figure 1c).
Discussion
A requirement of FHA and FHV methodologies is the design of fluoresceinated ligands recognized by receptors. With Fl-BN, C-terminal amino acids are essential for the binding of bombesin-like peptide to its receptor and amino acids toward the N terminus tolerate modification.21 With Fl-Folate, fluorescein was attached to carboxylic groups known to be nonessential for receptor binding.22, 23 With Fl-ASF, amino acids were modified by fluorescein since the asialoglycoprotein receptor binds to galactose residues. The activity of fluorescein Fl-EGF has been established.24, 25 To confirm a fluoresceinated ligand binds to the appropriate receptor, the binding of the ligand to cultured cells known to express the receptor can be employed as shown in Figure 2d and h.
A key feature of fluorescent hapten methodologies is the detection of bound fluorescein with a single reagent, an antifluorescein HRP conjugate. This reagent provides an amplification step using an antibody–enzyme conjugate which is used in immunoassay and immunohistochemistry and which is commercially available.
The use of fluorescein as a hapten for detection by an antifluorescein-HRP conjugate has several advantages over the biotin/avidin or the digitalis/antidigitalis systems. There are no reports of which we are aware of naturally occurring fluorescein or fluorescein-like molecules present in living systems, and reacting with antifluorescein antibodies. In contrast, biotin is involved in cell metabolism and present in the ng/ml range in human sera.26, 27 Substances reacting with antidigoxin antibodies, so-called endogenous digitalis-like factors, have long been noted.28 In addition, fluorescein has long been known to yield high affinity antifluorescein antibodies,29, 30, 31 presumably because it is so dissimilar from naturally occurring structures.
FHV and FHA methods provide a new approach for obtaining information about the binding of ligands to receptors present in cultured cell or tissue specimens. Several lines of evidence support the validity of these methods. First, similar apparent affinities were obtained by the FHA and FACS methods (Table 1), data that supplements the original comparison of the fluorescein immunoassay methodology with radioligand-based receptor methods.4 Second, when both FHV and FHA methods were performed (Figures 2 and 3), the visual distinctions seen by FHV were verified by the quantitative measurements using FHA. Finally, the pattern of bombesin peptide binding obtained with the FHV in normal and pancreatic cancer (Figure 4c) was that described with human specimens and radioreceptor methodology.20
In some cases it is impossible to compare our values with literature values because of differences in preparations and binding methodologies. However, three examples where the literature values are similar to FHV values for these numbers of sites per cell are provided. We obtained 300±24 bombesin-binding sites per cell (all numbers are in thousands per cell) for the normal mouse pancreas (Table 1). There are 44 bombesin receptor sites/cell on PC-3 cells32 and the density of bombesin peptide-binding sites in the normal mouse pancreas is approximately 10 times that of PC-3 cells, based on the accumulation of radioactive bombesin-like peptides after injection.33 Thus, the normal mouse pancreas has an eight- to 10-fold higher density of receptors than the PC-3 cells, which gives a value of between 320 and 440 sites per cell, compared with our value of 300±24 with the FHV method. The numbers of EGF receptors has been reported to be 1900 receptors per MDA-MB-468 cell,34 compared to our value of 1370±320 (Table 1). In addition to the values shown in Table 1, we determined number of EGF receptors per BT-20 cell to be 1340±400, compared values of 1000±40035 or 1500±500.36
Fluorescein-hapten visualization and assay methods hopefully will permit receptor-binding studies to be performed in pathology, cell biology, or pharmacology laboratories that lack radioligand receptor methodologies. In addition, the fluorescein hapten chemistry developed here will permit the development of microarray, tissue-based screening for receptor-binding ligands. With our tissue microarrays, we were limited to the FHV method because of an inability to confine reagents to individual specimens. However, overcoming this limitation of labware, and applying the chemistry shown in Figure 3, will yield tissue-based microarray receptor assays. For example, the affinity of drug candidates for the bombesin peptide-binding receptors could be measured through their inhibition of Fl-BN binding to a microarray of pancreatic tissue. FHA-based tissue microarrays might prove cheaper and be more reflective of receptors in tissues than receptor assays using cultured cells or purified protein.
References
Stumpf WE . Drug localization and targeting with receptor microscopic autoradiography. J Pharmacol Toxicol Methods 2005;51:25–40.
Dumont Y, Chabot JG, Quirion R . Receptor autoradiography as mean to explore the possible functional relevance of neuropeptides: focus on new agonists and antagonists to study natriuretic peptides, neuropeptide Y and calcitonin gene-related peptides. Peptides 2004;25:365–391.
Reubi JC, Waser B, Horisberger U, et al. In vitro autoradiographic and in vivo scintigraphic localization of somatostatin receptors in human lymphatic tissue. Blood 1993;82:2143–2151.
Kelly KA, Reynolds F, Weissleder R, et al. Fluorescein isothiocyanate-hapten immunoassay for determination of peptide–cell interactions. Anal Biochem 2004;330:181–185.
Stupack DG, Cheresh DA . Integrins and angiogenesis. Curr Top Dev Biol 2004;64:207–238.
Westlin WF . Integrins as targets of angiogenesis inhibition. Cancer J 2001;7 (Suppl 3):S139–S143.
Byzova TV, Plow EF . Activation of alphaVbeta3 on vascular cells controls recognition of prothrombin. J Cell Biol 1998;143:2081–2092.
Smith CJ, Volkert WA, Hoffman TJ . Gastrin releasing peptide (GRP) receptor targeted radiopharmaceuticals: a concise update. Nucl Med Biol 2003;30:861–868.
Sano H, Feighner SD, Hreniuk DL, et al. Characterization of the bombesin-like peptide receptor family in primates. Genomics 2004;84:139–146.
Birn H, Nielsen S, Christensen EI . Internalization and apical-to-basolateral transport of folate in rat kidney proximal tubule. Am J Physiol 1997;272:F70–F78.
Schwartz AL . The hepatic asialoglycoprotein receptor. CRC Crit Rev Biochem 1984;16:207–233.
Kokudo N, Vera DR, Makuuchi M . Clinical application of TcGSA. Nucl Med Biol 2003;30:845–849.
Rettenbacher M, Reubi JC . Localization and characterization of neuropeptide receptors in human colon. Naunyn Schmiedebergs Arch Pharmacol 2001;364:291–304.
Ferrier L, Segain JP, Bonnet C, et al. Functional mapping of NPY/PYY receptors in rat and human gastro-intestinal tract. Peptides 2002;23:1765–1771.
Cox HM, Tough IR . Neuropeptide Y, Y1, Y2 and Y4 receptors mediate Y agonist responses in isolated human colon mucosa. Br J Pharmacol 2002;135:1505–1512.
Pedrazzini T, Pralong F, Grouzmann E . Neuropeptide Y: the universal soldier. Cell Mol Life Sci 2003;60:350–377.
Zhou J, Chen J, Mokotoff M, et al. Targeting gastrin-releasing peptide receptors for cancer treatment. Anticancer Drugs 2004;15:921–927.
Jensen RT, Coy DH, Saeed ZA, et al. Interaction of bombesin and related peptides with receptors on pancreatic acinar cells. Ann NY Acad Sci 1988;547:138–149.
Markwalder R, Reubi JC . Gastrin-releasing peptide receptors in the human prostate: relation to neoplastic transformation. Cancer Res 1999;59:1152–1159.
Fleischmann A, Laderach U, Friess H, et al. Bombesin receptors in distinct tissue compartments of human pancreatic diseases. Lab Invest 2000;80:1807–1817.
Nock BA, Nikolopoulou A, Galanis A, et al. Potent bombesin-like peptides for GRP-receptor targeting of tumors with 99mTc: a preclinical study. J Med Chem 2005;48:100–110.
Leamon CP, Reddy JA . Folate-targeted chemotherapy. Adv Drug Deliv Rev 2004;56:1127–1141.
Sudimack J, Lee RJ . Targeted drug delivery via the folate receptor. Adv Drug Deliv Rev 2000;41:147–162.
Carraway III KL, Koland JG, Cerione RA . Visualization of epidermal growth factor (EGF) receptor aggregation in plasma membranes by fluorescence resonance energy transfer. Correlation of receptor activation with aggregation. J Biol Chem 1989;264:8699–8707.
Chatelier RC, Ashcroft RG, Lloyd CJ, et al. Binding of fluoresceinated epidermal growth factor to A431 cell sub-populations studied using a model-independent analysis of flow cytometric fluorescence data. EMBO J 1986;5:1181–1186.
Watanabe T, Yasumura S, Shibata H, et al. Biotin status and its correlation with other biochemical parameters in the elderly people of Japan. J Am Coll Nutr 1998;17:48–53.
Baker H . Assessment of biotin status: clinical implications. Ann NY Acad Sci 1985;447:129–132.
Hollenberg NK, Graves SW . Endogenous sodium pump inhibition: current status and therapeutic opportunities. Prog Drug Res 1996;46:9–42.
Muller JD, Nienhaus GU, Tetin SY, et al. Ligand binding to anti-fluorescyl antibodies: stability of the antigen binding site. Biochemistry 1994;33:6221–6227.
Bates RM, Ballard DW, Voss Jr EW . Comparative properties of monoclonal antibodies comprising a high-affinity anti-fluorescyl idiotype family. Mol Immunol 1985;22:871–877.
Voss Jr EW, Eschenfeldt W, Root RT . Fluorescein: a complete antigenic group. Immunochemistry 1976;13:447–453.
Reile H, Armatis PE, Schally AV . Characterization of high-affinity receptors for bombesin/gastrin releasing peptide on the human prostate cancer cell lines PC-3 and DU- 145: internalization of receptor bound 125I-(Tyr4) bombesin by tumor cells. Prostate 1994;25:29–38.
Maina T, Nock BA, Zhang H, et al. Species differences of bombesin analog interactions with GRP-R define the choice of animal models in the development of GRP-R-targeting drugs. J Nucl Med 2005;46:823–830.
Filmus J, Pollak MN, Cailleau R, et al. MDA-468, a human breast cancer cell line with a high number of epidermal growth factor (EGF) receptors, has an amplified EGF receptor gene and is growth inhibited by EGF. Biochem Biophys Res Commun 1985;128:898–905.
Falette N, Frappart L, Lefebvre MF, et al. Increased epidermal growth factor receptor level in breast cancer cells treated by 1,25-dihydroxyvitamin D3. Mol Cell Endocrinol 1989;63:189–198.
Imai Y, Leung CK, Friesen HG, et al. Epidermal growth factor receptors and effect of epidermal growth factor on growth of human breast cancer cells in long-term tissue culture. Cancer Res 1982;42:4394–4398.
Acknowledgements
We thank Dr Elena Aikawa, Dr Jan Grimm and Dr Rainer Kohler for helpful discussions. Xavier Montet was supported by a grant from the Swiss National Science Foundation. Work was supported by DOD Grant DAMD-170210089, RO1 EB00662 and P50 CA86355, and P01 CA69246.
Author information
Authors and Affiliations
Corresponding author
Additional information
Supplementary Information accompanies the paper on the Laboratory Investigation website (http://www.nature.com/labinvest).
Supplementary information
Rights and permissions
About this article
Cite this article
Montet, X., Yuan, H., Weissleder, R. et al. Enzyme-based visualization of receptor–ligand binding in tissues. Lab Invest 86, 517–525 (2006). https://doi.org/10.1038/labinvest.3700404
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700404
Keywords
This article is cited by
-
GRP Receptor Imaging of Prostate Cancer Using [99mTc]Demobesin 4: a First-in-Man Study
Molecular Imaging and Biology (2014)