Abstract
Early diagnosis of oral squamous cell carcinoma (OSCC) may have a major impact on survival and quality of life. Recent studies have shown that the majority of OSCC is preceded by precursor lesions characterized by genetic alterations. The aim of this study was to develop and evaluate a noninvasive screening test for oral preneoplastic lesions, based on genetic alterations as marker. Various methods to obtain a high yield of cells by brushing a small area of the oral mucosa were compared. A novel genetic assay, multiplex ligation-dependent probe amplification (MLPA), was applied that enables the measurement of gains and losses at 40 different chromosomal locations in one PCR reaction using 150 ng DNA. MLPA was performed on DNA of normal and dysplastic oral mucosa as well as of OSCC with the intention to select a specific probe set for accurate detection of precursor lesions in the oral cavity. The assay was correlated to loss of heterozygosity analysis using microsatellite markers, and evaluated on noncancer subjects and patients with oral leukoplakia. A noninvasive sampling method was developed with DNA yields ranging from 150 to 600 ng. Using 120 probes, we could detect large differences with MLPA in the number of alterations between normal vs dysplastic and dysplastic vs tumor tissue with P-values <0.001. A significant correlation was found between the number of alterations as detected by MLPA and the analysis for allelic loss. The available data enabled the selection of a set of 42 MLPA probes, which had the power to optimally discriminate between normal and dysplastic tissue. Our data show that MLPA is a sensitive, reliable, high-throughput and easy-to-perform technique, enabling the detection of genetic alterations on small noninvasive samples and can be considered a promising method for population-based screening of preneoplastic lesions in the oral cavity.
Similar content being viewed by others
Main
Worldwide approximately 300 000 individuals are yearly diagnosed with oral cancer, approximately 80% with squamous cell carcinomas (OSCC) that arise in the mucosal linings.1 Despite advances in locoregional therapy, still 50% of OSCC patients die of their disease.2, 3 Screening populations for the early detection of asymptomatic carcinoma or precursor lesions is an attractive strategy to reduce the burden of OSCC. The major example in this respect is the cytological screening of women with smears to prevent cervical cancer. The introduction of screening programs has considerably declined the incidence of cervical cancer in the last three to four decades in most developed countries.4 In the 1960s and 1970s, numerous reports on the use of oral cytology as a diagnostic approach were published. However, low sensitivity and specificity precluded the general adoption of microscopic cytology for the detection of oral cancer and precancerous lesions.5
In recent years, considerable progress has been made in understanding the genetic basis of OSCC carcinogenesis. It has been well established that these cancers arise as a result of the accumulation of genetic alterations in proto-oncogenes and tumor suppressor genes, referred to as multistep carcinogenesis.6 In 2003, Braakhuis et al proposed a progression model of oral cancer in which they described that mucosal fields with genetic alterations can replace the normal epithelium in the oral cavity. Clonal evolution in these fields leads to the invasive carcinoma.7, 8, 9 Despite the large dimensions up to 10 cm in diameter, the majority of these lesions are not visible by clinical inspection. Only a minor subgroup of precursor lesions can be seen clinically and present as a white or a red lesion in the oral mucosa, designated as leukoplakia or erythroplakia, respectively.10
The knowledge that far most, if not all, OSCC, may be preceded by large precursor lesions now opens the possibility for early diagnosis of these precancerous lesions.8 At this moment, histology is the gold standard for diagnosis and grading of oral preneoplastic lesions. The lesions are histologically recognized as either nondysplastic or dysplastic, and if dysplastic, graded as mildly, moderately or severely. The potential of histological examination to identify preneoplastic lesions might suggest that histology is suitable for screening. However, screening by taking random biopsies of both clinically normal and suspect oral tissue is unpractical, since this causes serious discomfort to the patient and is not suited for repeated sampling at multiple sites. Moreover, histological grading has some value for the assessment of the risk for progression, but due to inter- and intraobserver variability, it is of limited importance for the individual patient. Grading of preneoplastic lesions by genetic methods or DNA ploidy seems more promising in this respect.11, 12, 13, 14, 15, 16
Noninvasive sampling is a more attractive alternative that would allow repeated screening for preneoplastic lesions. The cells can be brushed or scraped from the mucosa,17 and the DNA analyzed for genetic alterations. It has been generally accepted that specific genetic changes are related to the carcinogenic process, and are of value to predict the risk for progression.18 Of note, to obtain a homogeneous sample of a potential preneoplastic lesion, only small samples should be used, restricting the amount of DNA available for analysis. On the other hand, numerous chromosomal regions have been associated with early OSCC carcinogenesis and should therefore be analyzed in parallel.
Hence, an assay is needed that allows analysis of multiple markers using only small amounts of DNA. Recently, a novel assay for analysis of multiple genetic alterations using small amounts of template DNA has been developed: multiplex ligation-dependent probe amplification (MLPA).19 This assay allows the measurement of numerical chromosomal alterations at 40 target locations using only 20 ng of DNA, corresponding to approximately 3000 cells. The aim of this study was to develop a noninvasive screening test using MLPA to detect preneoplastic lesions in the oral cavity.
Materials and methods
Patients and Tissue Samples
Normal oral mucosa was derived from 10 paraffin-embedded surgical specimens of uvulas from patients who underwent surgical treatment for snoring. In addition, 10 paraffin-embedded tumor specimens were selected from patients who underwent surgical treatment for OSCC. Dysplastic oral mucosa was obtained from 10 fresh frozen and 10 paraffin-embedded surgical margins that were taken from the specimen of surgically treated OSCC patients for routine histological assessment. All specimens were histologically examined by an experienced pathologist. Dysplasia was scored according to the standard criteria of the World Health Organisation.20
Specimens from three leukoplakia patients who underwent a transoral excision were used for comparison of the genetic changes in brushed cells, and those in the excised tissue samples. The leukoplakia lesion was brushed before excision, and the excised specimen was subsequently embedded in paraffin. The microdissected mucosal epithelium was analyzed for genetic changes as described below.
Blood samples and cells brushed from the oral mucosa obtained from 20 noncancer subjects served as control material.
The study was approved by the Institutional Review Board, and all patients were enrolled after written informed consent.
Noninvasive Sampling
For noninvasive sampling, exfoliated cells were brushed from the oral mucosa in noncancer controls and leukoplakia patients. Different sampling methods were tested, including a small disposable brush (Omnident®, Dental Union, Nieuwegein, The Netherlands), a Cytobrush plus® (Medscand Medical, Sweden) and an excavator 125–126 (Dentsply Ash, UK). To investigate the depth of sampling, cytospins were made from the different samples and subsequently immunostaining with suprabasal CK-13 and basal CK-19 antibody was performed, using IgG as negative control.
Sites of 5 × 10 mm were brushed repeatedly five times at different anatomical sites in the oral cavity in healthy controls or at the leukoplakia lesion before excision. Each brush was immediately placed into a 2 ml reaction vial containing 200 μl phosphate-buffered saline (PBS). After 15 s of vortexing, the brush was removed by a sterilized pair of tweezers. Next, the cells were pelleted by centrifuging the vials at 1500 g for 10 min. Finally, excess PBS was removed by a glass capillary and the pellets were stored at −20°C.
Microdissection
From all tissue specimens, 10-μm sections were cut and mounted on microscopic glass slides. The paraffin-embedded specimens were subsequently deparaffinized in xylene and rehydrated. The first and last tissue sections were stained with hematoxylin and eosin for histological analysis and to guide microdissection. The other tissue sections were stained with 1% toluidine blue and 0.2% methylene blue and manually microdissected under a stereomicroscope. From the uvulas and leukoplakia samples, the entire mucosal epithelium was microdissected. From the surgical margins and tumor samples, only dysplastic and neoplastic areas were microdissected, respectively.
Isolation of DNA
The pelleted exfoliated cells and blood cells as well as the microdissected tissues were treated with 1 mg/ml of proteinase K for 24 h at 52°C in 100 μl buffer containing 100 mM TRIS-HCL (pH 9.0), 10 mM NaCl, 1% sodium dodecyl sulfate and 5 mM EDTA (pH 8.2). The DNA was purified by phenol–chloroform extraction and collected by ethanol precipitation using 2 μg of glycogen as carrier. The DNA was redissolved in 20 μl LoTE buffer (3 mM TRIS-HCL, 0.2 mM EDTA, pH 7.5). DNA concentrations were measured by an ND-1000 spectrophotometer (B&L, Maarssen, The Netherlands). For comparison, DNA isolation from exfoliated cells was also performed with DNA Stat (Campro Scientific Bv, Veenendaal, The Netherlands) and QIAamp DNA mini Kit (50) (Qiagen GmbH, Hilden, Germany), carried out according to the suppliers.
Measurement of Numerical Chromosomal Alterations by MLPA
MLPA kits were used as described by the supplier.19 Three different probe sets each were evaluated with 40–42 probes detecting primarily oncogenes and tumor suppressor genes: SALSA P005, P006 and P007 (MRC-Holland, Amsterdam, The Netherlands). In addition, the SALSA P014 chromosome 8p kit was used. In short, 5 μl of genomic DNA diluted in LoTE at a concentration of 10 ng DNA/μl was denatured at 98°C for 5 min, cooled to 25°C and 3 μl of a 1:1 mixture of MLPA buffer and SALSA probe-mix was added. After hybridization for 16 h at 60°C, 32 μl ligation mix was added and the reaction was incubated for 15 min at 54°C followed by 5 min at 98°C. Subsequently, 40 μl of the SALSA PCR-mix (FAM label) was added to 10 μl of ligation product and this was amplified by PCR in 33 cycles (30 s 95°C, 30 s 60°C, 60 s 72°C). For analysis, 12 μl of deionized formamide was combined with 0.5 μl of Genescan-500 (Rox) size standard (Applied Biosystems, Nieuwerkerk a/d IJssel, The Netherlands) and 1 μl of PCR product in a Genetic Analyzer plate. This was denatured at 96°C for 2 min, loaded on an automated ABI PRISM sequence analyzer and run following the supplier's protocol (3100 Genetic Analyser; Applied Biosystems). All assays were performed in triplicate.
The data were analyzed with Genescan Analysis software (version 3.7; Applied Biosystems). All peak areas were exported to an Excel spreadsheet. The data were standardized on the basis of the total peak areas. For each peak, a ratio was calculated by dividing the median of the normalized peak area of the tissue sample by the normalized peak area of control DNA from peripheral blood samples, or as otherwise indicated. A number of quality controls were used. The total signal of all peaks had to pass a certain threshold value. When one or more MLPA reactions failed to pass this threshold value, the triplicate assay was repeated. Finally, in every MLPA run a tumor DNA sample and a normal DNA sample were included as control.
Microsatellite Analysis
Allelic loss was assessed using 23 microsatellite markers located at chromosomes 3p, 9p, 17p, 8p, 13q and 18q. These markers were selected because they frequently demonstrate loss of heterozygosity (LOH) in head and neck squamous cell carcinoma and preneoplastic lesions.6, 8 The following markers were used: D3S1274, D3S1284, D3S1217, D3S1766, D3S1029, D3S1293, D9S171, D9S1748, D9S1751, IFNA, D9S162, D9S157, CHRNB1, TP53, D17S1866, D13S294, D13S168, D13S170, D13S158, D18S34, D18S57, D8S1130 and LPL-GZ. Primer sequences were obtained from the Genomic Database for all of these markers (http://gdbwww.gdb.org/). LOH analysis was performed as described previously.8, 9
Statistical Analysis
All MLPA experiments were performed in triplicate. For analysis, the median was used to control for outliers. Fisher's exact test was used to assess the difference of the number of genetic alterations between the different experimental groups. P-values <0.05 were considered statistically significant. To determine the relation between MLPA and LOH analysis on the dysplastic tissues, the Pearson correlation coefficient was calculated, using SPSS (SPSS for windows, release 10.1; SPSS, Chicago, IL, USA). To determine more specific cutoff values for genetic alterations in MLPA, the 99.99% confidence intervals to set the criteria for ‘normal’ were calculated for each probe on the log-transformed data.
Results
Noninvasive Sampling
Noninvasive sampling of 5 × 10 mm areas in the oral cavity of healthy controls showed a minimal yield of 30 000 exfoliated cells per brushed site. We found that the yield varied with site, with the lowest yields at the palate and the highest yields at the gingiva and lateral border of the tongue. Measured DNA amounts showed a range of 150–600 ng/sample.
For noninvasive sampling, we chose to use a small disposable brush. Other sampling methods, the Cytobrush plus® and excavator 125–126, gave less reproducible and lower yields. All three different sampling methods showed comparable numbers of positive cells for CK-13 and CK-19, suggesting that the same depth was obtained (data not shown).
DNA isolation from exfoliated cells with proteinase K isolation showed approximately five times higher DNA yield compared to DNA isolation with DNA Stat and QIAamp DNA mini Kit.
Suitability of MLPA
MLPA was performed on DNA isolated from 10 normal oral mucosa and 10 OSCC (paraffin-embedded) samples, using all available probes, to set initial cutoff values to decide whether the DNA copy number was normal or showed gain or loss. The genetic status of the OSCC samples was determined previously with LOH analysis using microsatellite markers and this information was exploited.8, 9 The following arbitrary criteria for an aberration were defined: a ratio <0.75 was considered a loss and a ratio >1.33 was considered a gain. Within these limits, all normal samples were free of losses and gains (0/1160 probes), whereas tumor tissues showed losses and gains in 29.5% (342/1160 probes), which compared well with the previous LOH data. Also, 10 paraffin-embedded surgical margins that were dysplastic and known to contain genetic alterations were analyzed by MLPA, showing losses and gains in 23.0% (267/1160 probes). The MLPA results of normal vs dysplastic tissues and dysplastic vs tumor tissues showed significant differences in the number of alterations with P-values <0.001. The unprocessed MLPA data, displayed as peaks, showed distinct differences between normal and dysplastic mucosal epithelium. A typical example is shown in Figure 1.
Unprocessed MLPA data of DNA derived from normal oral mucosa (upper panels) and from a dysplastic surgical margin (lower panels), all paraffin-embedded tissue. Duplicate MLPA analyses are indicated as exp. 1 and exp. 2. On the x-axis, the number of base pairs is depicted, and on the y-axis, the fluorescence intensity. The lower left arrow indicates a gain of 11q and the lower right arrow indicates a loss of 17p in the DNA of the dysplastic sample. Besides these two genetic changes, a considerable number of other alterations can be observed in the dysplastic sample.
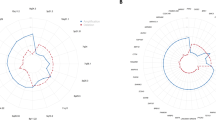
A widely used and more accepted assay for genetic analysis of oral premalignant lesions is detection of LOH using microsatellite markers. Previous studies have even reported that LOH at different chromosome arms is associated with an increased risk for OSCC.13, 14 To see whether MLPA analysis corresponds to LOH analysis on the same tissues, we compared the results of both assays on 20 surgical margins, focusing on those chromosomal regions for which both LOH markers and MLPA probes were available. The MLPA assay on fresh frozen tissue does not differ from the MLPA assay on paraffin-embedded tissue. The overall percentage LOH and overall percentage genetic alterations (gains and losses) as assessed with MLPA was determined for each sample and their Pearson correlation coefficient was calculated to be 0.474, with a two-tailed P-value of 0.035, indicating a moderate but nevertheless significant correlation between MLPA and LOH results (Figure 2).
Results of MLPA analysis and LOH analysis on 20 dysplastic surgical margins. On the x-axis, the % MLPA alterations (gains and losses) are shown, and on the y-axis, the % LOH alterations for each sample. In all, 14 MLPA markers were used: 03p21.3 (2x), 03p22, 08p23.1, 08p22 (2x), 09p21 (2x), 13q14.3, 17p13.1 (4x) and 18q11.2. A total of 13 LOH markers were used: D3S1029, D3S1293, D9S171, D9S1748, D9S1751, D9S162, TP53, D13S294, D13S168, D18S34, D18S57, LPL-GZ and D8S1130. Pearson correlation coefficient=0.474, with a two-tailed P-value of 0.035.
Selection of Probes and Cutoff Values for an OSCC Set
Since the DNA amount of the small noninvasive samples is limited and not all available probes are relevant for the detection of oral preneoplastic lesions, the most suitable probes were combined in one probe set. Probes, suitable for the detection of oral preneoplastic lesions, were selected based on the following characteristics: (1) genetic alterations in a high percentage of dysplastic oral lesions, (2) reproducible measurements, and (3) no genetic alteration in healthy controls using the arbitrary cutoff values of 1.33 and 0.75. Probes at chromosomes 2 (IL1A-DO1, TANK-DO1) and 17q (ERBB2-DO2), which did not show genetic alterations in dysplastic mucosa and OSCC, were added as control probes enabling independent standardization of the data.
Initially, cutoff values of 1.33 and 0.75 were used to define a gain or loss, respectively. These values were chosen more or less arbitrarily, and do not take variation between probes into account. Therefore, for each MLPA probe in the OSCC probe set, a 99.99% confidence interval was calculated, using the median values of 20 DNA samples of blood from noncancer controls. The upper limit of the confidence interval was used to define gain, while the lower limit defines loss. The mean coefficient of variation for the probes over these 20 samples was determined to be 7.2%, with a range of 4.0–17.6%. The MLPA assay, using the OSCC probe set with specific cutoff values, was performed on 10 normal, 10 dysplastic and 10 tumor tissues, all paraffin embedded. The normal tissues showed 0% (0/420), the dysplastic tissues 21.4% (90/240) and the tumor tissues 29.3% (123/420) genetic alterations (Figure 3). The number of genetic alterations between the normal and the dysplastic tissues, as well as between the dysplastic and tumor tissues remained significantly different (P<0.0001 and P=0.011, respectively). The 99.99% confidence interval corresponding to a nonaltered DNA content of all probes is presented in Table 1.
MLPA results from DNA obtained from microdissected normal, dysplastic and OSCC tissue samples, using the OSCC probe set. Columns from left to right: probe name, chromosome position, 10 normal, 10 dysplastic and 10 OSCC tissue samples. For calculations, the 99.99% confidence interval limits were used. Explanation of used colors: gray=gain, black=loss and white=no genetic alteration. The normal tissues showed 0% (0/420), the dysplastic tissues 21.4% (90/420) and the tumor tissues 29.3% (123/420) genetic alterations.
Feasibility of Noninvasive Screening by MLPA
In order to investigate the feasibility of noninvasive screening by MLPA, we first brushed the oral mucosa of 20 noncancer subjects. None of these samples showed genetic changes. Then, we compared the MLPA results from noninvasive samples, obtained by brushing the mucosal surface of three different leukoplakia lesions, with the MLPA results from the microdissected mucosal epithelium of the paraffin-embedded surgical specimen. All surgical specimens were found to be nondysplastic and showed no genetic alterations. The MLPA results of the exfoliated cells showed in all three cases no genetic alterations.
Discussion
This study was performed to develop and evaluate a noninvasive screening test for oral preneoplastic lesions, based on genetic markers. Early diagnosis of oral preneoplastic lesions might be of importance for clinical management, particularly in high-risk populations, considering the poor prognosis of advanced stage tumors. Patients who are known to be at high risk for developing OSCC and would qualify for screening are those with oral leukoplakia or erythroplakia, those who have been treated for OSCC and those with excessive tobacco and alcohol abuse. Besides these well-known risk groups, there are also patients who are genetically predisposed, suffering from Fanconi anemia, or relatives from OSCC patients who should be considered as risk groups.21, 22
Methods such as toluidine blue staining23 and autofluorescence imaging have been investigated to improve the clinical identification of oral premalignant lesions as the majority other than leukoplakia and erythroplakia cannot be recognized by visual inspection. These methods would allow frequent surveillance of lesions by imaging or inspection and, when indicated, histological examination of biopsies, but to date there is not much evidence for the efficacy of these visualization methods. Considering that these premalignant lesions are characterized by genetic alterations, and that genetic markers can be assayed on minimal amounts of tissue, we developed a genetic test that allows analysis of noninvasive samples.
To ensure homogeneous sampling within a precursor lesion, we used small brushes. These gave the most consistent results, even more reliable than scraping cells with an excavator. This simple sampling method might even allow self-sampling in the future. As expected, the amount of DNA isolated from these samples was limited, and we therefore decided to exploit the recently developed MLPA test. Although the lower limit of DNA for analysis has been indicated as 20 ng according to the supplier, we found in numerous experiments that 40–60 ng of input DNA gave most reliable results. Furthermore, we consider triplicate assays a prerequisite to ensure sufficient accuracy.
Previously, we used microsatellite markers to assess the presence of mucosal precursor lesions, and we analyzed whether the genetic changes determined by allelic loss correlate to those determined by MLPA. Although both techniques determine essentially genetic alterations, numerical methods such as MLPA do not allow one to detect copy number neutral events, while allelic loss analysis by microsatellites is less suited for detecting gains. In addition, the exact locations of microsatellite markers and MLPA probes did not exactly correspond. Despite these limitations, the data collected by both assays correlated significantly, indicating that major changes detected by LOH analysis are in agreement with major changes as detected by MLPA.
Based on the results obtained with tumor, dysplastic and normal mucosa, we selected a number of markers from all available genomic probe sets that could be combined in a single OSCC probe set. Initially, we used arbitrary cutoff levels to indicate a loss or gain (a ratio of 0.75 or 1.33, respectively), and using these limits, there were significant genetic differences between normal and dysplastic mucosa, indicating that MLPA with this OSCC probe set has the ability to discriminate between normal and preneoplastic tissue. However, screening exfoliated samples is different from screening microdissected mucosal epithelium. It is to be expected that exfoliated samples of precursor lesions are contaminated with few wild-type cells, necessitating a higher sensitivity to detect genetic alterations. After selection of the probes for the OSCC screening set, we decided to refine the cutoff level to improve the accuracy and included variation per probe. The 99.99% confidence interval for normality of each separate probe was determined on 20 normal DNA samples isolated from blood of noncancer subjects. Within these limits, all tissue samples from noncancer controls showed no genetic alterations. It should be noted that there is a variation in the confidence intervals making some markers more reliable than others.
To determine proof of principle of noninvasive screening by MLPA, we analyzed noninvasive samples of leukoplakia biopsies and their corresponding noninvasive samples. In addition, we analyzed 20 brushed noninvasive samples of noncancer subjects. All samples revealed reliable data and no alterations were found as expected. These results indicated the feasibility of noninvasive screening by MLPA. The next step is a prospective screening study to determine the clinical sensitivity and specificity by analyzing a large number of matched brushed and biopsy samples in leukoplakia and erythroplakia patients. Subsequently, the prognostic value of MLPA-based detection of oral preneoplastic lesions should be ascertained in a large cohort of subjects at risk. As the number and type of specific genetic alterations is most likely associated with the risk for progression of a lesion,12, 13, 14, 17 criteria must be defined that allow identification of specifically those lesions with a high risk for progression.
In summary, the MLPA technique with a panel of specific markers has proven to be sufficiently sensitive, accurate, high-throughput and easy-to-perform on small noninvasive samples, and can be considered a promising method to improve the early diagnosis of preneoplastic lesions in the oral cavity.
References
Parkin DM . Global cancer statistics in the year 2000. Lancet Oncol 2001;2:533–543.
Vokes EE, Weichselbaum RR, Lippman SM, et al. Head and neck cancer. N Engl J Med 1993;328:184–194.
Leemans CR, Tiwari R, Naura JJ, et al. Recurrence at the primary site in head and neck cancer and the significance of neck lymph node metastases as a prognostic factor. Cancer 1994;73:187–190.
Wells M, Östör AG, Crum CP, et al. Tumours of the uterine cervix. In: Fattaneh AT, Peter D (eds). World Health Organization Classification of Tumours, Pathology and Genetics, Tumours of the Breast and Female Genital Organs. IARC Press: Lyon, 2003, pp 262–266.
King Jr OH . Cytology—its value in the diagnosis of oral cancer. Dent Clin N Am 1971;15:817–826.
Califano J, van der Riet P, Westra W, et al. Genetic progression model for head and neck cancer: implications for field cancerization. Cancer Res 1996;56:2488–2492.
Braakhuis BJM, Tabor MP, Kummer JA, et al. A genetic explanation of Slaughter's concept of field cancerization: evidence and clinical implications. Cancer Res 2003;63:1727–1730.
Tabor MP, Brakenhoff RH, van Houten VMM, et al. Persistence of genetically altered fields in head and neck cancer patients: biological and clinical implications. Clin Cancer Res 2001;7:1523–1532.
Tabor MP, Brakenhoff RH, Ruijter-Schippers HJ, et al. Multiple head and neck tumors frequently originate from a single preneoplastic lesion. Am J Pathol 2002;161:1051–1060.
Van der Waal I, Schepman KP, Van der Meij EH, et al. Oral leukoplakia: a clinicopathological review. Oral Oncol 1997;33:291–301.
Tabor MP, Braakhuis BJM, van der Wal JE, et al. Comparative molecular and histological grading of epithelial dysplasia of the oral cavity and the oropharynx. J Pathol 2003;199:354–360.
Reibel J . Prognosis of oral premalignant lesions: significance of clinical, histopathological, and molecular biological characteristics. Crit Rev Oral Biol Med 2003;14:47–62.
Mao L, Lee JS, Fan YH, et al. Frequent microsatellite alterations at chromosomes 9p21 and 3p14 in oral premalignant lesions and their value in cancer risk assessment. Nature Med 1996;2:682–685.
Rosin MP, Cheng X, Poh C, et al. Use of allelic loss to predict malignant risk for low-grade oral epithelial dysplasia. Clin Cancer Res 2000;6:357–362.
Partridge M, Pateromichelakis S, Phillips E, et al. A case–control study confirms that microsatellite assay can identify patients at risk of developing oral squamous cell carcinoma within a field of cancerization. Cancer Res 2000;60:3893–3898.
Sudbø J, Kildal W, Risberg B, et al. DNA content as a prognostic marker in patients with oral leukoplakia. N Engl J Med 2001;344:1270–1278.
Spafford MF, Koch WM, Reed AL, et al. Detection of head and neck squamous cell carcinoma among exfoliated oral mucosal cells by microsattelite analysis. Clin Cancer Res 2001;7:607–612.
Rosin MP, Lam WL, Poh C, et al. 3p14 and 9p21 loss is a simple tool for predicting second oral malignancy at previously treated oral cancer sites. Cancer Res 2002;62:6447–6450.
Schouten JP, McElgunn CJ, Waaijer R, et al. Relative quantification of 40 nucleic acid sequences by multiplex ligation-dependent probe amplification. Nucleic Acid Res 2002;30:e57.
Pindborg JJ, Reichart PA, Smith CJ, et al. Histological Typing of Cancer and Precancer of the Oral Mucosa. World Health Organisation. Springer-Verlag: Berlin, 1997, pp 21–31.
Kutler DI, Auerbach AD, Satagopan J, et al. High incidence of head and neck squamous cell carcinoma in patients with Fanconi anemia. Arch Otolaryngol Head Neck Surg 2003;129:106–112.
Copper MP, Jovanovic A, Nauta JJ, et al. Role of genetic factors in the etiology of squamous cell carcinoma of the head and neck. Arch Otolaryngol Head Neck Surg 1995;121:157–160.
Guo Z, Yamaguchi K, Sanchez-Cespedes M, et al. Allelic loss in oratest-directed biopsies of patients with prior upper aerodigestive tract malignancy. Clin Cancer Res 2001;7:1963–1968.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Bremmer, J., Braakhuis, B., Ruijter-Schippers, H. et al. A noninvasive genetic screening test to detect oral preneoplastic lesions. Lab Invest 85, 1481–1488 (2005). https://doi.org/10.1038/labinvest.3700342
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700342
Keywords
This article is cited by
-
Diagnostic aids for detection of oral precancerous conditions
International Journal of Oral Science (2013)
-
Lower expression of ATM and gene deletion is more frequent in adrenocortical carcinomas than adrenocortical adenomas
Endocrine (2012)
-
The molecular biology of head and neck cancer
Nature Reviews Cancer (2011)