Abstract
RET/PTC1 and RET/PTC3 are the markers for papillary thyroid carcinoma. Their reported prevalence varies broadly. Nonrearranged c-RET has also been detected in a variable proportion of papillary carcinomas. The published data suggest that a wide range in expression levels may contribute to the different frequency of c-RET and, particularly, of RET/PTC detection. However, quantitative expression analysis has never been systematically carried out. We have analyzed by real-time RT-PCR 25 papillary carcinoma and 12 normal thyroid samples for RET/PTC1, RET/PTC3 and for RET exons 10–11 and 12–13, which are adjacent to the rearrangement site. The variability in mRNA levels was marked and four carcinoma groups were identified: one lacking RET/PTC rearrangement with balanced RET exon levels similar to those of the normal samples (7/25 cases, 28%), the second (6/25 cases, 24%) with balanced RET expression and very low levels of RET/PTC1, the third with unbalanced RET exons 10–11 and 12–13 expression, high RET/PTC1 levels but no RET/PTC3 (7/25 cases, 28%), and the fourth with unbalanced RET expression, high RET/PTC1 levels and low levels of RET/PTC3 (5/25 cases, 20%). Papillary carcinomas with high RET/PTC1 expression showed an association trend for large tumor size (P=0.063). Our results indicate that the variability in c-RET and RET/PTC mRNA levels contributes to the apparent inconsistencies in their reported detection rates and should be taken into account not only for diagnostic purposes but also to better understand the role of c-RET activation in thyroid tumorigenesis.
Similar content being viewed by others
Main
Oncogenic c-RET activation in thyroid tumors composed of follicular cells is the result of chromosomal rearrangements resulting in the fusion of the RET tyrosine-kinase (RET-TK) domain to the 5′-terminal region of heterologous genes. The rearrangements are a molecular marker for papillary thyroid carcinoma as their very name, RET/PTC for papillary thyroid carcinoma, implies,1 and consist of balanced inversions or translocations that involve the 3.0 kb intron 11 of c-RET. To date, at least 16 chimeric mRNAs affecting 11 different genes have been reported (Saenko et al,2 reviewed in Tallini and Asa3), of which RET/PTC1 (consisting of the fusion of RET with H4) and RET/PTC3 (consisting of the fusion of RET with RFG/ELE1) are by far the most common.3 Their prevalence varies broadly from zero to more than 60% in nonradiation-associated cases3 and is close to 90% in some series of papillary carcinomas from the Chernobyl area diagnosed after the 1986 nuclear disaster.4 While the high prevalence of RET/PTC in radiation-associated thyroid tumors is consistent with misrepair of radiation-induced double-strand DNA breaks,5, 6 the high variability in the prevalence of RET/PTC in sporadic tumors has no specific explanation. In addition to rearranged RET forms, c-RET expression has also been identified in a highly variable proportion of papillary carcinoma samples. Although its significance is unclear, c-RET may also play a role in thyroid tumorigenesis.7, 8, 9, 10, 11
Analysis of the data published in the literature suggests a wide range in the expression levels of RET/PTC and c-RET in tumor samples. This may contribute more than geographical factors to the differences in the reported frequencies of c-RET and, particularly, of RET/PTC detection. However, quantitative expression analysis has never been systematically carried out. To this purpose, we have analyzed by real-time quantitative RT-PCR RNA extracted from both papillary carcinoma and normal thyroid samples for the expression of RET/PTC1 and RET/PTC3. We have also analyzed RET exons 10–11 and 12–13 which are adjacent to the RET/PTC rearrangement site in c-RET intron 11. Thus, the primers and probe set for c-RET exons 10–11 identify nonrearranged c-RET only, whereas the set for exons 12–13 identifies both c-RET and its rearranged RET/PTC forms.
Materials and methods
Tissue Samples and Cell Lines
Thyroid samples were obtained from the Department of Pathology at the Yale University School of Medicine. Excess fresh tissue from surgically resected specimens of thyroid tumors and perilesional normal thyroid samples were snap frozen and kept at −80°C until RNA extraction. Pathologic material in each case was reviewed and tumors were classified according to established criteria.12 Before RNA extraction, representative sections of the frozen tissue were cut and examined histologically to confirm the adequacy of the sample, the diagnosis and to estimate the proportion of follicular cells vs stromal and inflammatory cells. The proportion of follicular cells in the perilesional samples of the normal set was always greater than 80%. Only tumor cases with at least 65% neoplastic follicular cells were included.
The cell lines utilized in the study were the TT medullary thyroid carcinoma cell line (CRL-1803, American Type Culture Collection-ATCC, Manassas, VA, USA), the TPC1 thyroid follicular cell line,13 the ARO undifferentiated thyroid carcinoma cell line transfected with a RET/PTC3 construct,14 and the mouse lymphoid line L1210 (Yale Cancer Center Tissue Culture Facility).
RNA Extraction and Quantitation
Frozen tissue sections and cell pellets from the cultured cells lines were incubated in 100 μl digestion buffer (0.01 M Tris pH 7.8, 0.005 M EDTA, 0.5% SDS) containing 2 μg of Proteinase K at 55oC overnight. RNA extraction was performed using Trizol (Invitrogen Corporation, Carlsbad, CA, USA) following the product insert instructions. The final RNA pellet was resuspended in RNA Storage Solution (Ambion Inc., Austin, TX, USA). The RNA in the solutions was DNAse treated using a DNA-free kit (Ambion Inc., Austin, TX, USA) following the insert instructions. The DNAse-treated RNA was checked and accurately measured using the RiboGreen RNA Quantitation kit (Molecular Probes, Inc, Eugene OR, USA). Fluorescence intensity was assessed with the Cytofluor Series 4000 (Applied Biosystems, Foster City, CA, USA).
Reverse Transcription
Total RNA was reverse transcribed to cDNA with the GeneAmp RNA PCR Core kit following the manufacturer's instructions (Applied Biosystems, Foster City, CA, USA). Briefly, 1 × PCR buffer II, 2.5 U/μl of MuLV reverse transcriptase, 1 U/μl RNAase inhibitor, 5 mM MgCl2, 1 mM each dNTP and 2.5 mM random hexamers were incubated with the extracted total RNA for 10 min at 25°C, for 60 min at 42°C and for 5 min at 95°C with a PTC/200 thermocycler (MJ Research, Waltham, MA, USA). Unless otherwise specified in the result section, 1 μg of input RNA was used to evaluate exons 10–11, 12–13, RET/PTC1 and RET/PTC3 mRNA expression in tissue samples. Negative controls were carried out in parallel by replacing the RNA templates or the MuLV reverse transcriptase with DEPC water. cDNA reverse transcribed from the TPC1 cell line13 was used as positive control for RET/PTC1 rearrangement. That obtained from the ARO cell line transfected with a RET/PTC3 construct14 was used as positive controls for RET/PTC3. RNA extracted from the medullary carcinoma TT cell line was used as positive control for the amplification of c-RET exons 10/11 and 12/13.
Primers and Probes
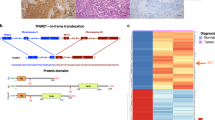
The primer and probe sequences are shown in Figure 1. They were custom designed using Primer Express software (Applied Biosystems, Foster City, CA, USA). The primer and probe set for c-RET exons 10–11 identifies nonrearranged c-RET only, whereas the set for exons 12–13 identifies both c-RET and its rearranged RET/PTC forms. The forward (nucleotides 274–288) and the reverse (nucleotides 318–339) primers as well as the probe (nucleotides 290–316) for RET/PTC1 were based on the reported NCBI database sequence accession number M31213.1 The forward (nucleotides 670–688) and the reverse (nucleotides 728–750) primers as well as the probe (nucleotides 690–718) for RET/PTC3 were based on the reported NCBI database sequence accession number X77548.15 The forward (nucleotides 1852–1867) and the reverse (nucleotides 1897–1913) primers as well as the probe (nucleotides 1872–1895) for c-RET exons 10 and 11 were based on the standard c-RET sequence (accession number X12949) with the numeration beginning from the start codon of c-RET.16 The probe for c-RET exons 10/11 spans the 10th intron of RET. The forward (nucleotides 2261–2277) and the reverse (nucleotides 2304–2324) primers as well as the probe (nucleotides 2279–2302) for c-RET exons 12 and 13 were similarly designed with the RET exon 12/13 probe spanning the 12th intron of RET. β-actin RNA was used as a quantitative reference using primers and probes previously described.17 All primers were synthetized by the program for Critical Technologies in Molecular Medicine Laboratory (Department of Pathology, Yale University School of Medicine). The probes were synthetized by the Custom Oligo Synthesis Service (Applied Biosystems, Foster City, CA, USA).
(a) Position of primers and probes relative to RET/PTC1, RET/PTC3 and c-RET. The primer and probe set for c-RET exons 10–11 identifies nonrearranged c-RET only, whereas the set for exons 12–13 identifies both c-RET and its rearranged RET/PTC forms. Ex, exon. (b) Primer and probe sequences, optimized concentrations and amplicon sizes for RET/PTC1, RET/PTC3 and c-RET.
Real-Time Quantitative PCR
For real-time quantitative PCR, 50 μl aliquots contained 5 μl of the cDNA previously transcribed, 1 × TaqMan Universal PCR Master Mix (PE Applied Biosystems, Foster City, CA, USA) and the optimized concentrations of primers and probes shown in Figure 1. Two-step PCR thermal cycling for DNA amplification and real-time data acquisition were performed with an ABI PRISM 7700 Sequence Detection System using the following cycle conditions: 50°C for 2 min × 1 cycle, 95°C for 10 min × 1 cycle, and 95°C for 15 s followed by 60°C for 1 min × 45 cycles. Each cDNA sample was assayed in triplicate. Negative cDNA controls (no cDNA or no MuLV reverse transcriptase, see above) were cycled in parallel with each run. Fluorescence data were analyzed by the ABI7700 software and expressed as Ct, the number of cycles needed to generate a fluorescent signal above a predefined threshold. Baseline and threshold values were set by the ABI7700 software.
Quantitation of Target Gene Expression
Target gene mRNA levels were expressed as (normalized for β-actin) or (normalized for β-actin and relative to a calibrator cell line) according to the equations derived by Livak and Schmittgen.18 The amount of target gene mRNA (XN), normalized to a reference gene, is given by the equation

where ΔCt is the difference in Ct values for the target and reference genes, E is the efficiency of amplification and is assumed to be equal for target and reference genes, and K is a constant. If the efficiency is close to one for both target and reference genes, then

The value of K is unknown and depends on several factors including the reporter dye used in the probe, sequence context effects on the fluorescence properties of the probe, the efficiency of probe cleavage, the purity of the probe and the setting of the fluorescence threshold.18 Each of these factors may differ between target and reference genes, and therefore K may not necessarily equal 1. If K does equal 1, then the normalized amount of target equals . In order to allow a broad comparison of c-RET and RET/PTC gene expression levels in thyroid tissues, K was assumed to be 1 for all genes, and normalized mRNA levels were expressed as with respect to β-actin.
The amount of target in a sample (q), normalized to β-actin and relative to a calibrator (cb), is given by the equation

where . Relative levels of c-RET and RET/PTC mRNA in thyroid tissues were expressed as using the TT cell line as the calibrator for exons 10–11 and 12–13, the TPC1 cell line as the calibrator for RET/PTC1, and RET/PTC3-transfected ARO cells as the calibrator for RET/PTC3.
Validation of gene expression by the and method was achieved by preparing standard curves for each gene using RNA extracted from TT cells (for exons 10–11 and 12–13), TPC1 (for RET/PTC1) and RET/PTC3-transfected ARO cells (for RET/PTC3). Efficiencies of amplification were close to 1 for all genes (see Results), justifying the expression of mRNA levels as and values.
Statistical Analysis
Intra-assay variability was assessed as the coefficient of variance (CV) of Ct values measured in triplicate in a single run. Interassay variability was assessed as the CV of mean Ct values measured on 3–10 independent runs performed on separate days. CVs were calculated as the standard deviation divided by the mean and expressed as percent. Differences between groups were analyzed by the Mann–Whitney test for nonparametric data or by one-way ANOVA, with P<0.05 considered to be significant.
Results
RET Exons 10–11, 12–13, RET/PTC1 and RET/PTC3 mRNA Expression in Cell Lines
TT, TPC1 and RET/PTC3-transfected ARO cells were used as controls for expression of RET (exons 10–11 and 12–13), RET/PTC1 and RET/PTC3, respectively. As expected, rearranged RET forms were not detected in TT cells. However, small amounts of exons 10–11 were detected in TPC1 and RET/PTC3 transfected ARO cells indicative of low c-RET expression (Table 1). The control cell lines were used to derive standard curves describing the relationship between Ct values and the amount of input RNA. Standard curves were also obtained for the reference gene β-actin. For all genes studied, Ct values were linearly related to input RNA with R2 values >0.9 (Figure 2). Efficiencies of amplification (defined as 10−1/slope−1) were always >0.95 and were not significantly different from 1. To determine whether the cellular environment influences the efficiency of amplification, standard curves were also obtained by diluting each cell line expressing c-RET or RET/PTC in a background of nonexpressing mouse cells (L1210 cell line), and maintaining a constant input of total RNA. Linear relationships between Ct values and cell number were obtained for all genes with R2 values >0.9 (Figure 3). Efficiencies of amplification were 0.99–1.05, with no difference in efficiencies derived by RNA dilution vs those obtained by cell:cell dilution indicating that the cellular environment does not affect the efficiency of amplification of RET exons 10–11 and 12–13, RET/PTC1 and RET/PTC3.
RNA dilution standard curves for RT-PCR amplification of c-RET and RET/PTC mRNA. Serial dilutions were made of total RNA from TT cells (a), TPC1 cells (b) and RET/PTC3-transfected ARO cells (c). Linear relationships between threshold cycle (Ct) and input RNA were obtained for all target genes and the reference gene β-actin, with R2>0.9. Points represent mean±s.e.m. of n=29 for TT cells, n=25 for TPC1 cells, and n=3 for RET/PTC3-transfected ARO cells.
Cell dilution standard curves for RT-PCR amplification of c-RET and RET/PTC mRNA. TT, TPC1 and RET/PTC3-transfected ARO cells were diluted in a background of L1210 cells not expressing c-RET or its rearranged forms. Linear relationships between threshold cycle (Ct) and cell number were obtained for all target genes, with R2>0.9. Points represent mean±s.e.m. of n=4 for all cell lines.
The reproducibility of real-time RT-PCR measurements for each gene was evaluated on the respective control cell line. The intra-assay variability (%CV) of Ct values determined in triplicate within a single analytical run was consistently <2% for all genes. The interassay variability (%CV) of Ct values determined in separate analytical runs was 2.4% for exons 10–11 (n=10), 2.5% for exons 12–13 (n=10), 2.0% for RET/PTC1 (n=10), 0.4% for RET/PTC3, and 1.6% for β-actin (n=10).
RET Exons 10–11, 12–13, RET/PTC1 and RET/PTC3 mRNA Expression in Thyroid Tissue
RET exons 10–11 and 12–13 mRNA were detected in the same 10 normal samples out of the 12 examined and in all the 25 papillary carcinoma cases (Figure 4a and b, Table 1). There was no difference between normal and neoplastic samples in median and values for RET exons 10–11 indicating similar c-RET expression in the two groups. RET exons 10–11 and 12–13 mRNA levels in both groups were 3 orders of magnitude lower than in the TT cell line, as indicated by median values of ∼10−3 (Table 1). RET exons 10–11 and 12–13 expression varied among samples. For normal thyroids, the range of – values was ∼200-fold for both exons 10–11 and 12–13, whereas for papillary carcinomas, the range was ∼3000-fold for exons 10–11 and ∼16 000-fold for exons 12–13 (Table 1).
Normalized mRNA levels in normal thyroid and papillary carcinoma. mRNA levels for exons 10–11 (a), exons 12–13 (b), RET/PTC1 (c) and RET/PTC3 (d) were measured in normal thyroid (N), papillary thyroid carcinoma (PC) and calibrator cell lines (TT cells for exons 10–11 and 12–13; TPC1 cells for RET/PTC1; RET/PTC3-transfected ARO cells for RET/PTC3). mRNA levels were normalized to β-actin and expressed as . Points represent individual cases, and the bar represents the median.
RET/PTC1 was detected in 18/25 papillary carcinomas cases, but in none of the normal thyroid cases (Figure 4c, Table 1). Median RET/PTC1 expression in the positive papillary carcinoma cases was of the same order of magnitude as that measured in the TPC1 cell line (= 0.6, Table 1). However, RET/PTC1 mRNA expression varied widely, with ∼100 000-fold range in – values. In fact, two statistically distinct sets of cases could be identified with 6/18 positive cases expressing low RET/PTC1 levels (<10−3, or RET/PTC1 less than 0.1% of β-actin) and 12/18 positive cases expressing high RET/PTC1 levels (>10−3, or RET/PTC1 greater than 0.1% of β-actin) (P<0.001, Mann–Whitney test). RET/PTC1 amplicons were identified by direct gel visualization and ethidium bromide staining in 7/12 positive cases expressing high RET/PTC1 levels.
RET/PTC3 was detected in 5/25 papillary carcinoma cases, but in none of the normal thyroid cases (Figure 4d, Table 1). All five papillary carcinoma cases expressing RET/PTC3 also had high levels (>10−3) of RET/PTC1. Median RET/PTC3 expression in the positive papillary carcinomas was of the same order of magnitude as that measured in RET/PTC3-transfected ARO cells (=0.3, Table 1). RET/PTC3 mRNA expression varied over a 100-fold range for and values. RET/PTC3 levels, normalized to β-actin (), were similar to RET/PTC1 levels in the set of papillary carcinomas with low RET/PTC1 expression (<10−3, or less than 0.1% of β-actin) (Figure 4c and d). RET/PTC3 amplicons were not identified by direct gel visualization and ethidium bromide staining.
Balance of RET Exons 12–13 vs Exons 10–11 Expression in Thyroid Tissue
In normal thyroid tissues, exons 12–13 mRNA expression was linearly related to exons 10–11 mRNA expression (Figure 5; R2=0.99). The slope of the linear regression line (0.98) was not significantly different from 1, confirming that exons 12–13 mRNA expression reflects c-RET. In contrast, plotting exons 12–13 mRNA expression vs exons 10–11 mRNA expression in papillary carcinomas demonstrated the presence of two distinct sets, one with balanced expression of exons 10–11 and 12–13 containing 13/25 cases, and one with an unbalance between RET exons 10–11 and 12–13 containing 12/25 cases (Figure 5). The papillary carcinoma set with balanced RET expression also exhibited a linear relationship between exons 12–13 and 10–11 mRNA expression (R2=0.89, slope=0.81), with no significant difference in the slope of the linear regression line between it and normal thyroid cases. In contrast, the set with unbalanced RET expression, failed to show a significant relationship between exons 12–13 and 10–11 mRNA levels (R2=0.32), and exhibited exon 12–13 expression levels higher than expected for a 1:1 relationship between the two exons.
Relationship between exons 10–11 and 12–13 mRNA expression in normal thyroid and papillary carcinoma (PC) with balanced and unbalanced RET exons 10–11 vs 12–13 expression. mRNA levels were normalized to β-actin and expressed as . Data points represent individual cases. The dotted line represents the linear regression line for normal thyroid tissue (R2=0.99), with a slope of 0.98.
Exons 10–11 mRNA levels were similar in papillary carcinomas with balanced and unbalanced RET, although there was a trend for lower exons 10–11 expression in the unbalanced samples (P=0.081, Mann–Whitney test) (Figure 6a). In contrast, exons 12–13 mRNA expression was significantly higher in the unbalanced samples (P<0.0005, Mann–Whitney test) (Figure 6b). RET/PTC1 mRNA was detected in 6/12 cases with balanced RET expression (median =2.3 × 10−5, or 0.002% RET/PTC1 relative to β-actin), and in all cases with unbalanced RET expression, with a significantly higher median of 5.1 × 10−2, or 5% RET/PTC1 relative to β-actin (P<0.001, Mann–Whitney test) (Figure 6c). RET/PTC3 mRNA was not detected in papillary carcinomas with balanced RET expression but was detectable in 5/13 of cases with unbalanced RET (median =5.4 × 10−5, or 0.005% RET/PTC3 relative to β-actin) (Figure 6d).
Normalized mRNA levels in papillary carcinoma with balanced and unbalanced RET exons 10–11 vs 12–13 expression. mRNA levels for exons 10–11 (a), exons 12–13 (b), RET/PTC1 (c) and RET/PTC3 (d) were normalized to β-actin and expressed as . Points represent individual cases and the bar represents the median of positive cases.
Thus, papillary carcinoma with and without balanced RET expression differ in RET/PTC levels. Tumors with balanced RET expression are characterized by low expression of exons 12–13 mRNA, low or no RET/PTC1 mRNA and no RET/PTC3 mRNA. Those with unbalanced RET expression exhibit higher levels of exons 12–13 and RET/PTC1 mRNA, and in some cases may also express RET/PTC3 mRNA. Of note, the two papillary carcinoma sets identified by examining the relationship between RET exons 10–11 and 12–13, correspond to the two distinct clusters identified previously as cases with low and high RET/PTC1 mRNA levels. The balanced set corresponds to those cases having RET/PTC1 values lower than 10−3, whereas the unbalanced set corresponds to those cases having values higher than 10−3.
Effect of Input RNA on the Detection Limit for RET/PTC1 and RET/PTC3 in Tissue Samples
In order to determine the minimum amount of input RNA required to detect RET/PTC mRNA in papillary carcinoma samples, RNA dilutions were prepared from 10 papillary carcinoma cases with unbalanced exons 12–13 vs 10–11 expression, that is, cases displaying high levels of RET/PTC1 mRNA. As illustrated in Figure 7a, RET/PTC1 mRNA was detected in 10/10 cases using an input of 1 μg RNA, 9/10 cases with 100 ng input RNA, 7/10 cases with 10 ng input RNA, 4/10 cases with 1 ng input RNA and 1/10 cases with 0.1 ng input RNA. Of the 10 cases examined, four also expressed RET/PTC3 mRNA. RET/PTC3 mRNA was detected in 4/4 cases using an input of 1 μg RNA, 1/10 cases with 100 ng input RNA and none at further dilutions (Figure 7b).
Effect of input RNA amount on detection of RET/PTC mRNA expression in PC. mRNA levels of RET/PTC1 (a) and RET/PTC3 (b) were normalized to β-actin and expressed as . Points represent individual cases and the bar represents the median of positive cases. Numbers in brackets represent the number of cases in which mRNA was detected at each RNA dilution out of 10 cases analyzed.
Proportion of RET and RET/PTC Producing Cells in Papillary Carcinoma as TT, TPC1 or RET/PTC3-Transfected ARO Cell Equivalents
The amount of RET and RET/PTC mRNA in papillary carcinoma samples was compared to standard curves (Figure 3) derived by diluting c-RET-, RET/PTC1- and RET/PTC3-expressing cells (ie TT cells, TPC1 cells or RET/PTC3-transfected ARO cells) in a background of nonexpressing cells (L1210). This allows an estimate of the proportion of positive cells in tumor samples as TT, TPC1 or RET/PTC3-transfected ARO cell equivalents. The expression levels of papillary carcinoma samples in terms of cell equivalents are shown in Figure 8. Thus, median cell equivalents are 1:100 000-TT:L1210 cells for exons 10–11 mRNA, 1:20 000-TT:L1210 cells for exons 12–13 mRNA, 1:600-TPC1:L1210 cells for RET/PTC1 mRNA, and 1:1000-RET/PTC3-transfected ARO:L1210 cells for RET/PTC3 mRNA. However, direct comparisons of cell equivalents between RET and its rearranged forms are not possible since each cell line produces different amounts of target gene. In fact, when normalized to β-actin levels, RET/PTC3 expression by transfected ARO cells is ∼100 times lower than RET/PTC1 expression by the TPC1 cell line and ∼10 000 times lower than c-RET expression by TT cells (Figure 4).
Cell equivalents for mRNA expression in papillary carcinoma with RET/PTC rearrangements. Cell equivalents for exons 10–11 (a), exons 12–13 (b), RET/PTC1 (c) and RET/PTC3 (d) are expressed as the ratio of calibrator cells (TT, TPC1 or RET/PTC3-transfected ARO cells) to L1210 mouse cells. TT, TPC1 or RET/PTC3-transfected ARO cell equivalents were estimated from cell dilution standard curves (see Figure 3). Data are represented as individual data points for all papillary carcinoma cases, and box and whiskers for papillary carcinomas with balanced (B) and unbalanced (U) RET exons 10–11 vs 12–13 expression. Boxes extend from the 25th to 75th percentile, bars within boxes represent the median, and whiskers reflect the range.
As seen with the mRNA data normalized to β-actin ( values), cell equivalent expression of RET and RET/PTC also showed wide variability. RET/PTC1 expression in terms of TPC1 cell equivalents revealed two distinct sets, one with a low expression of RET/PTC1 mRNA corresponding to a cell dilution greater than 1:10 000 cells (median 1:60 000-TPC1:L1210 cell equivalents), and the other with a high expression of RET/PTC1 corresponding to a cell dilution lower than 1:10 000 cells (median 1:100-TPC1:L1210 cell equivalents) (P<0.001, Mann–Whitney test). The cases in these two sets are the same as those in the previously identified clusters characterized by balanced RET expression with low RET/PTC1 mRNA levels, and unbalanced RET expression with high RET/PTC1.
RET and RET/PTC Expression Groups and Correlation with Clinicopathologic Features in Papillary Carcinoma
In summary, quantitative analysis of RET and RET/PTC expression demonstrates four groups of papillary carcinomas of comparable numerical size. One group is characterized by the lack of RET/PTC rearrangement, and RET exons 10–11 and 12–13 levels similar to those observed in the normal samples. This group includes 7/25 cases (28%) (Table 2). The second group (6/25 cases or 24%, Table 2) also has RET expression similar to those of the normal samples, with balanced exons 10–11 and 12–13. However, it expresses RET/PTC1 at a low level in terms of β-actin normalization (<10−3, or RET/PTC1 less than 0.1% of β-actin, Figure 6c) as well as in terms of TPC1 cell equivalents (RET/PTC1 corresponding to dilutions greater than one TPC1 cell in 10 000 nonrearranged cells Figure 8c). The third and fourth groups include cases with unbalanced RET expression (Figure 5). These cases are characterized by higher RET/PTC1 in terms of β-actin normalization (>10−3, or RET/PTC1 greater than 0.1% of β-actin, Figure 6c) as well as in terms of TPC1 cell equivalents (RET/PTC1 corresponding to dilutions lower than one TPC1 cell in 10 000 nonrearranged cells Figure 8c). The third and fourth groups differ only with respect to RET/PTC3, which is absent in the third but present in the fourth (Figure 6d). They include 7/25 (28%) and 5/25 (20%) cases, respectively (Table 2).
All tumors were clinically evident with a median percentage of neoplastic follicular cells vs stromal and inflammatory cells of 80%. Median tumor size was 3.5 cm in cases with high RET/PTC1 expression (third and fourth papillary carcinoma groups), 2.7 cm in cases with low RET/PTC1 (second papillary carcinoma group) and 1.5 cm in cases with no RET/PTC1 (first papillary carcinoma group). This suggests that tumor sizes may parallel RET/PTC1 expression levels, although the difference did not reach statistical significance (P=0.063, one-way ANOVA). Expression levels of RET or of its rearranged forms did not correlate with the papillary carcinoma type, presence of lymph node metastasis or tumor stage.
Discussion
The prevalence of RET/PTC1 and RET/PTC3 reported in sporadic papillary carcinoma, varies broadly and the reason for this variability is unclear. The existence of a geographic factor is shown by comparing the prevalence of RET/PTC activation in different studies of nonradiation-associated papillary carcinomas, all of which have utilized comparable techniques (ie RT-PCR followed by hybridization of the amplified products with RET specific probes to detect RET/PTC)19, 20, 21, 22, 23 and by the observation of regional variability within one country in samples analyzed by the same laboratory.24 This points to the putative existence of unknown environmental agents favoring RET rearrangement and to the role of genetic susceptibility for papillary cancer.25 However, much of the difference in the prevalence of RET/PTC activation appears to be related to the technique used to detect the rearranged DNA or RNA. The most common techniques include Southern blotting, direct gel visualization of the RT-PCR amplicons and hybridization of blotted RT-PCR products. Meta-analysis of the data published worldwide over a greater than 10-year period (Table 3) demonstrates an average prevalence of 18% for Southern blotting (67/370 cases tested, range 0–33%),26, 27, 28, 29, 30, 31 15% for direct visualization of the RT-PCR amplicons (47/309 cases tested, range 4–45%)7, 32, 33, 34, 35 and 48% for hybridization of blotted RT-PCR products (153/314 cases tested, range 0–85%).4, 21, 22, 23, 36, 37, 38, 39 Allowing for the fact that Southern blotting permits the identification of all RET/PTC variants while the cases analyzed by RT-PCR listed above only screened the tumors for RET/PTC1, -2 and -3, there is a good correspondence between the sensitivity of Southern blotting and of direct gel visualization of RT-PCR products, as also demonstrated by parallel analysis of the same samples with the two methods.44 When RT-PCR is followed by blotting of the amplicons and hybridization with RET-specific probes the sensitivity increases dramatically, by at least 100-fold according to some investigators.11 This is not only reflected in the meta-analysis of the world literature but by the higher rate of RET/PTC detected in the same laboratory when direct gel visualization was replaced by hybridization of blotted RT-PCR products (5% compared with 61%),23, 45 or by the increase in the proportion of positive cases from 4 to 76% in two similar series from the same country.7, 37 In fact, even accounting for different amplification efficiencies of the PCR primers, technical details that increase the sensitivity of the methodology such as blotting larger amounts of RT-PCR product before hybridization, result in an increased RET/PTC detection rate.37 Although some increase in RET/PTC detection rate due to any combination of factors which affect sensitivity is certainly expected, the extent of such an increase23, 45 is surprising and points to a wide range of RET/PTC expression in papillary carcinoma.
A number of studies have noted expression of c-RET in papillary thyroid carcinoma tumor samples.7, 8, 9, 10, 11 Wild-type RET may coexist with RET/PTC9, 10 and it has been proposed as a marker for aggressive disease.10 Similar to RET/PTC, c-RET mRNA has been reported in a variable proportion of papillary carcinomas with frequencies ranging from 21%10 to 100%.8, 9
To address the issue of the expression level of RET and its rearranged forms in thyroid papillary carcinoma we have utilized real-time RT-PCR and quantitatively analyzed high-quality RNA extracted from frozen samples for the expression of RET/PTC1, RET/PTC3 and RET exons 10–11 and 12–13, which are adjacent to the RET/PTC rearrangement site. RET/PTC1 was detectable in 18/25 (72%) papillary carcinomas. RET/PTC3 coexisted with RET/PTC1 in 5/25 (20%) tumors. Exons 10–11 mRNA, indicative of c-RET expression, were detectable in all the papillary carcinomas and in 10/12 (83%) normal thyroids. Our results indicate a remarkable variability in mRNA levels. The variability is highest for RET/PTC1. It is also significant for c-RET and RET/PTC3 and it is not accounted for by the proportion of follicular cells in the samples analyzed. In fact, the range for the proportion of neoplastic follicular cells in tumor samples was within one order of magnitude and overlapped with that of the normal cases.
Statistical analysis of the quantitative data shows four groups of papillary carcinomas of similar size. One lacks RET/PTC rearrangement and has normalized RET exons 10–11 and 12–13 levels similar to those observed in the non-neoplastic samples. The second group exhibits minimal expression of RET/PTC1. RET/PTC1 expression in this group is at least 10 000 times lower as compared with β-actin, corresponds to a dilution of RET/PTC1 positive TPC1 cells greater than one in 10 000 nonrearranged cells, and it is so low that it is not associated with detectable unbalance between normalized RET exons 10–11 and 12–13 levels. In the third and fourth carcinoma groups RET exons 10–11 and 12–13 expression is clearly different from the normal samples and from the carcinoma groups one and two. The RET exons expression pattern is unbalanced, as expected in the case of RET/PTC rearrangement. Cases are characterized by high RET/PTC1 expression compared to the second carcinoma group both in terms of normalized RET/PTC1 levels and TPC1 cell equivalents. The third and fourth groups differ only with respect to RET/PTC3, which is absent in the third but present in the fourth. Of note, RET/PTC3 levels in the fourth group are low and of the same order of magnitude as RET/PTC1 levels in the second carcinoma group.
RET/PTC1 in this study was identified by direct gel visualization and ethidium bromide staining in only a proportion of our cases with high-level expression. The prevalence of RET/PTC1 identified by direct gel visualization is within the range of that reported in the literature. However, the majority of the RET/PTC1 rearranged cases and all of the RET/PTC3 rearrangements could only be identified with the TaqMan assay, confirming its superior sensitivity compared with the conventional solution phase RT-PCR.42 The overall high prevalence of RET/PTC in this study is thus consistent with the use of real-time RT-PCR technology, of high-quality RNA extracted from frozen tissue and accurately quantified, and with the high median proportion of neoplastic follicular cells in the samples analyzed. It is also in line with results obtained after hybridization of blotted RT-PCR products, although papillary carcinomas with low RET/PTC levels may escape detection even after RET hybridization of blotted RET/PTC amplicons.37
The highly variable expression of RET/PTC in tumor samples explains the increase in RET/PTC1 detection rate which parallels modifications to enhance the sensitivity of the methodology used for its detection.23, 37, 43 It is also consistent with the drop in the RET/PTC detection rate that occurs as the amount of input RNA is decreased.
The existence of papillary carcinomas with very low RET/PTC expression, the identification of which is highly dependent on any combination of factors that affect sensitivity, explains to a great extent the variation in RET/PTC prevalence rates. Furthermore, the existence of quantitative variation in the expression levels should be taken into account to investigate the correlation of RET/PTC with clinical findings. Critical review of the data available in the literature still indicates that RET/PTC has no definite role in the progression of thyroid carcinoma to aggressive forms of cancer and may indeed identify a subset of tumors with better outcome.34 However, it is possible that quantitative analysis may disclose new correlations with clinical or pathologic features, as suggested by the association trend between high RET/PTC1 levels and tumor size in this small series. The biological relevance of RET/PTC rearrangements expressed at very low levels is also unclear. It is possible that in certain tumors the rearranged RET forms are particularly unstable or that the rearrangement is only present in a subset of the neoplastic cells. Regardless, it is reasonable to question the relevance of RET/PTC1 when the rearrangement is present at levels equivalent to less than one RET/PTC1-positive TPC1 cell in 10 000 nonrearranged cells.
The presence of low-level RET/PTC3 mRNA was identified in a subset of cases. The coexistence of RET/PTC1 and RET/PTC3 rearrangements is known to occur in papillary carcinoma.22, 23, 37, 43 In fact, the presence of more than two different RET rearrangements in a single tumor indicates multiple RET/PTC rearranged clones in at least some papillary carcinomas.37 Since RET/PTC3 is only expressed at low levels in tumor cases with high RET/PTC1 expression, RET/PTC3 rearrangement appears to be a secondary phenomenon possibly due to the presence of minor neoplastic cell clones. Given the association, it is tempting to speculate that RET/PTC1 may even predispose the neoplastic cell to acquire the second RET rearrangement. RET/PTC3 without RET/PTC1 coexpression is frequently detected in tall cell papillary carcinoma, but it is uncommon in other types of papillary cancer. Tall cell papillary carcinoma is a rare but aggressive papillary carcinoma variant.12 The higher mitogenic effects of RET/PTC3 compared with RET/PTC1 have indeed been postulated as the molecular basis for the aggressive biologic behavior of tall cell papillary carcinoma.46 However, none of our cases had morphologic findings to justify such a diagnosis and none expressed high RET/PTC3 levels. In fact, tumors with both RET/PTC3 and RET/PTC1 rearrangements did not exhibit any particular clinicopathologic feature.
Our study demonstrates that c-RET is detectable in all papillary carcinomas to a level comparable with that of normal thyroid parenchyma. The expression level is variable, particularly among tumor samples, explaining the apparent inconsistency of the reported detection rates.8, 9, 10 The levels of c-RET expression are in any case ∼1000 times lower when compared to the control TT medullary carcinoma cell line. The expression level of RET exons 12–13 in individual cases is statistically higher relative to that of exons 10–11 only in those cases which express high levels of rearranged RET. This fully validates the use of methodologies that rely on the unbalance in the expression of RET exons adjacent to the RET/PTC breakpoint in c-RET intron 11, although the variability in the relative RET exon unbalance calls for great caution in the visual interpretation of gels or of hybridized of blots of RT-PCR products. The presence of detectable exons 10–11 mRNA even in those cases with high RET/PTC1 expression indicates that both c-RET and rearranged RET forms coexist in papillary carcinoma samples. The source of c-RET in such samples is debated and challenges the notion that c-RET is expressed by thyroid follicular cells only as a consequence of RET/PTC rearrangements. c-RET mRNA in papillary carcinoma may be the result of contaminating C-cells or more likely neutrophils and macrophages that express both mRNA and wild-type RET protein.47 However, low-level mRNA expression by thyroid follicular cells is a likely possibility.8, 9, 11 This is supported in our study by the lack of C-cell contamination in tumor samples as indicated by negative results for calcitonin immunohistochemistry (data not shown) and by the identification of low-level exons 10–11 in the RET/PTC1 positive TPC1 thyroid follicular cell line and in the RET/PTC3-transfected ARO cells. Interestingly, among papillary carcinomas with high-level RET/PTC1 expression there is a trend for lower RET exon 10–11 levels compared to those tumors with low-level RET/PTC1 or no rearrangement. If c-RET is indeed expressed by follicular cells, the reduction in RET exon 10–11 levels may suggest the existence of molecular mechanisms that allow RET/PTC to downregulate c-RET expression.
In summary, this is the first report of quantitative expression analysis of RET and its rearranged forms in papillary thyroid carcinoma. Our results indicate that c-RET and RET/PTC levels are highly variable. This variability contributes to the apparent inconsistencies in their reported detection rates and should be taken into account not only for diagnostic purposes but also to better understand the role of RET activation in thyroid follicular cell tumorigenesis.
References
Grieco M, Santoro M, Berlingieri MT, et al. PTC is a novel rearranged form of the ret proto-oncogene and is frequently detected in vivo in human thyroid papillary carcinomas. Cell 1990;60:557–563.
Saenko V, Rogounovitch T, Shimizu-Yoshida Y, et al. Novel tumorigenic rearrangement, Delta rfp/ret, in a papillary thyroid carcinoma from externally irradiated patient. Mutat Res 2003;527:81–90.
Tallini G, Asa SL . RET oncogene activation in papillary thyroid carcinoma. Adv Anatom Pathol 2001;8:345–354.
Nikiforov YE, Rowland JM, Bove KE, et al. Distinct pattern of ret oncogene rearrangements in morphological variants of radiation-induced and sporadic thyroid papillary carcinomas in children. Cancer Res 1997;57:1690–1694.
Mizuno T, Kyoizumi S, Suzuki T, et al. Continued expression of a tissue specific activated oncogene in the early steps of radiation-induced human thyroid carcinogenesis. Oncogene 1997;15:1455–1460.
Nikiforova MN, Stringer JR, Blough R, et al. Proximity of chromosomal loci that participate in radiation-induced rearrangements in human cells. Science 2000;290:138–141.
Learoyd DL, Messina M, Zedenius J, et al. RET/PTC and RET tyrosine kinase expression in adult papillary thyroid carcinomas. J Clin Endocrinol Metab 1998;83: 3631–3635.
Bunone G, Uggeri M, Mondellini P, et al. RET receptor expression in thyroid follicular epithelial cell-derived tumors. Cancer Res 2000;60:2845–2849.
Fluge Ø, Haugen DR, Akslen LA, et al. Expression and alternative splicing of c-RET RNA in papillary thyroid carcinomas. Oncogene 2001;20:885–892.
Kjellman P, Learoyd DL, Messina M, et al. Expression of the RET proto-oncogene in papillary thyroid carcinoma and its correlation with clinical outcome. Br J Surg 2001;88:557–563.
Nikiforova MN, Caudill CM, Biddinger P, et al. Prevalence of RET/PTC rearrangements in Hashimoto's thyroiditis and papillary thyroid carcinomas. Int J Surg Pathol 2002;10:15–22.
Rosai J, Carcangiu ML, DeLellis RA . Atlas of Tumor Pathology—Tumors of the thyroid gland, 3rd series. Washington: Armed Force Institute of Pathology, 1992.
Ishizaka Y, Ushijima T, Sugimura T, et al. cDNA cloning and characterization of ret activated in a human papillary thyroid carcinoma cell line. Biochem Biophys Res Comm 1990;168:402–408.
Santoro M, Papotti M, Chiappetta G, et al. RET activation and clinicopathologic features in poorly differentiated thyroid tumors. J Clin Endocrinol Metab 2002;87:370–379.
Santoro M, Dathan NA, Berlingieri MT, et al. Molecular characterization of RET/PTC3: a novel rearranged version of the RET proto-oncogene in a human thyroid papillary carcinoma. Oncogene 1994;9:509–516.
Takahashi M, Buma Y, Iwamoto T, et al. Cloning and expression of the ret protooncogene encoding a tyrosine kinase with two potential transmembrane domains. Oncogene 1988;3:571–578.
Kreuzer KA, Lass U, Landt O, et al. Highly sensitive and specific fluorescence reverse transcription-PCR assay for the pseudogene-free detection of beta-actin transcripts as quantitative reference. Clin Chem 1999; 45:297–300.
Livak KL, Schmittgen TD . Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 2001;25:402–408.
Ishizaka Y, Kobayashi S, Ushijima T, et al. Detection of ret/PTC transcripts in thyroid adenomas and adenomatous goiter by an RT-PCR method. Oncogene 1991;6:1667–1672.
Zou M, Shi Y, Farid NR . Low rate of ret proto-oncogene activation (PTC/retTPC) in papillary thyroid carcinomas from Saudi Arabia. Cancer 1994;73:176–180.
Motomura T, Nikiforov YE, Namba H, et al. Ret rearrangements in Japanese pediatric and adult papillary thyroid cancers. Thyroid 1998;8:485–489.
Tallini G, Santoro M, Helie M, et al. RET/PTC oncogene activation defines a subset of papillary thryoid carcinomas lacking evidence of progression to poorly differentiated or undifferentiated tumor phenotypes. Clin Cancer Res 1998;4:287–294.
Sugg SL, Ezzat S, Rosen IB, et al. Distinct multiple RET/PTC gene rearrangements in multifocal papillary thyroid neoplasia. J Clin Endocrinol Metab 1998;83: 4116–4122.
Pierotti MA, Vigneri P, Bongarzone I . Rearrangements of RET and NTRK1 tyrosine kinase receptors in papillary thyroid carcinomas. Recent Results Cancer Res 1998;154:237–247.
Lesueur F, Corbex M, McKay JD, et al. Specific haplotypes of the RET proto-oncogene are over-represented in patients with sporadic papillary thyroid carcinoma. J Med Genet 2002;39:260–265.
Namba H, Yamashita S, Pei HC, et al. Lack of PTC gene rearrangement in human thyroid tumors. Endocrinol Japan 1991;38:627–632.
Wajjwalku W, Nakamura S, Hasegawa Y, et al. Low frequency of rearrangement of the ret and trk proto-oncogenes in Japanese thyroid papillary carcinomas. Jpn J Cancer Res 1992;83:671–675.
Jhiang SM, Caruso DR, Gilmore E, et al. Detection of the PTC/ret oncogene in human thyroid cancers. Oncogene 1992;7:1331–1337.
Santoro M, Carlomagno F, Hay ID, et al. RET oncogene activation in human thyroid neoplasms is restricted to the papillary carcinoma subtype. J Clin Invest 1992; 89:1517–1522.
Soares P, Fonseca E, Wynford-Thomas D, et al. Sporadic ret-rearranged papillary carcinoma of the thyroid: a subset of slow growing, less aggressive thyroid neoplasms? J Pathol 1998;185:71–78.
Bongarzone I, Vigneri P, Mariani L et al. RET/NTRK1 rearrangements in thyroid gland tumors of the papillary carcinoma family: correlation with clinicopathological features. Clin Cancer Res 1998;4:223–228.
Lee C-H, Hsu LS, Chi CW, et al. High frequency of rearrangement of the RET protooncogene (RET/PTC) in Chinese papillary thyroid carcinomas. J Clin Endocrinol Metab 1998;83:1629–1632.
Mayr B, Potter E, Goretzki P, et al. Expression of Ret/PTC1, -2, -3, -Δ3, and -4 in German papillary thyroid carcinoma. Br J Cancer 1998;77:903–906.
Musholt TJ, Musholt PB, Khaladj N, et al. Prognostic significance of RET and NTRK1 rearrangements in sporadic papillary thyroid carcinoma. Surgery 2000;128:984–993.
Fenton CL, Lukes Y, Nicholson D, et al. The ret/PTC mutations are common in sporadic papillary thyroid carcinoma of children and young adults. J Clin Endocrinol Metab 2000;85:1170–1175.
Park KY, Koh JM, Kim YI, et al. Prevalences of Gs alpha, ras, p53 mutations and ret/PTC rearrangement in differentiated thyroid tumours in a Korean population. Clin Endocrinol 1998;49:317–323.
Chua EL, Wu WM, Tran KT, et al. Prevalence and distribution of ret/ptc 1, 2, and 3 in papillary thyroid carcinoma in New Caledonia and Australia. J Clin Endocrinol Metab 2000;8:2733–2739.
Elisei R, Romei C, Vorontsova T, et al. RET/PTC rearrangements in thyroid nodules: studies in irradiated and not irradiated, malignant and benign thyroid lesions in children and adults. J Clin Endocrinol Metab 2001;86:3211–3216.
Puxeddu E, Moretti S, Giannico A, et al. Ret/PTC activation does not influence clinical and pathological features of adult papillary thyroid carcinomas. Eur J Endocrinol 2003;148:505–513.
Williams GH, Rooney S, Thomas GA, et al. RET activation in adult and childhood papillary thyroid carcinoma using a reverse transcriptase-n-polymerase chain reaction approach on archival-nested material. Br J Cancer 1996;74:585–589.
Cinti R, Yin L, Ilc K, et al. RET rearrangements in papillary thyroid carcinomas and adenomas detected by interphase FISH. Cytogenet Cell Genet 2000;88:56–61.
Sheils OM, O'Leary JJ, Sweeney EC . Assessment of ret/PTC-1 rearrangements in neoplastic thyroid tissue using TaqMan RT-PCR. J Pathol 2000;192:32–36.
Finn SP, Smyth P, O'Leary J, et al. Ret/PTC chimeric transcripts in an Irish cohort of sporadic papillary thyroid carcinoma. J Clin Endocrinol Metab 2003;88: 938–941.
Bongarzone I, Fugazzola L, Vigneri P, et al. Age-related activation of the tyrosine kinase receptor protooncogenes RET and NTRK1 in papillary thyroid carcinoma. J Clin Endocrinol Metab 1996;81:2006–2009.
Sugg SL, Zheng L, Rosen IB, et al. Ret/PTC-1, -2, and -3 oncogene rearrangements in human thyroid carcinomas: implications for metastatic potential? J Clin Endocrinol Metab 1996;81:3360–3365.
Basolo F, Giannini R, Monaco C et al. Potent mitogenicity of the RET/PTC3 oncogene correlates with its prevalence in tall-cell variant of papillary thyroid carcinoma. Am J Pathol 2002;160:247–254.
Gattei V, Celetti A, Cerrato A, et al. Expression of the RET receptor tyrosine kinase and GDNFR-alpha in normal and leukemic human hematopoietic cells and stromal cells of the bone marrow microenvironment. Blood 1997;89:2925–2937.
Acknowledgements
This work was supported by the Departments of Pathology and Laboratory Medicine, Yale University School of Medicine, New Haven, CT, USA.
Author information
Authors and Affiliations
Corresponding author
Rights and permissions
About this article
Cite this article
Rhoden, K., Johnson, C., Brandao, G. et al. Real-time quantitative RT-PCR identifies distinct c-RET, RET/PTC1 and RET/PTC3 expression patterns in papillary thyroid carcinoma. Lab Invest 84, 1557–1570 (2004). https://doi.org/10.1038/labinvest.3700198
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/labinvest.3700198
Keywords
This article is cited by
-
A study to evaluate association of nuclear grooving in benign thyroid lesions with RET/PTC1 and RET/PTC3 gene translocation
Thyroid Research (2023)
-
Malignant struma ovarii: next-generation sequencing of six cases revealed Nras, Braf, and Jak3 mutations
Endocrine (2021)
-
Values of molecular markers in the differential diagnosis of thyroid abnormalities
Journal of Cancer Research and Clinical Oncology (2017)
-
A mutation screening of oncogenes, tumor suppressor gene TP53 and nuclear encoded mitochondrial complex I genes in oncocytic thyroid tumors
BMC Cancer (2015)
-
Prevalence of RET/PTC1 and RET/PTC3 gene rearrangements in Chennai population and its correlation with clinical parameters
Tumor Biology (2014)