Abstract
Dementia with Lewy bodies (DLB) is the second most common form of degenerative dementia. Siblings of affected individuals are at greater risk of developing DLB, but little is known about the underlying genetic basis of the disease. We set out to determine whether mutations in known highly penetrant neurodegenerative disease genes are found in patients with DLB. Whole-exome sequencing was performed on 91 neuropathologically confirmed cases of DLB, supplemented by independent APOE genotyping. Genetic variants were classified using established criteria, and additional neuropathological examination was performed for putative mutation carriers. Likely pathogenic variants previously described as causing monogenic forms of neurodegenerative disease were found in 4.4% of patients with DLB. The APOE ɛ4 allele increased the risk of disease (P=0.0001), conferred a shorter disease duration (P=0.043) and earlier age of death (P=0.0015). In conclusion, although known pathogenic mutations in neurodegenerative disease genes are uncommon in DLB, known genetic risk factors are present in >60% of cases. APOE ɛ4 not only modifies disease risk, but also modulates the rate of disease progression. The reduced penetrance of reported pathogenic alleles explains the lack of a family history in most patients, and the presence of variants previously described as causing frontotemporal dementia suggests a mechanistic overlap between DLB and other neurodegenerative diseases.
Similar content being viewed by others
Introduction
Dementia with Lewy bodies (DLB) is the second most common form of dementia. It affects 5% of the population over 75 years of age,1 and has a greater impact on healthcare provision than Alzheimer’s disease (AD).2 The neuropathological hallmark of DLB is widespread α-synuclein-positive neuronal inclusions (Lewy bodies and Lewy neurites) and in addition this is often associated with amyloid deposition.3 Siblings of affected individuals have a 2.3-fold increased risk of developing the disorder,4 but little is known about the genetic aetiology of the disease. Although genetic variants in APOE,5 GBA,6 SNCA and SCARB2 (ref. 7) have been associated with an increased risk of DLB, only a few families have been described with more than two first-degree relatives,8 and no single highly penetrant gene defects have been shown to cause familial forms of the disorder. Using exome sequencing in 91 autopsy-confirmed cases, here we determined whether confirmed or putative pathogenic mutations in genes in known neurodegenerative disease genes are found in patients with DLB.
Materials and methods
Subjects and sample preparation
We studied 91 post-mortem cases conforming to both the clinical and post-mortem diagnostic criteria for DLB.3 Two patients were first-degree relatives (mother and daughter) and two patients were siblings (brothers). The remaining 87 patients had no recorded family history of neurodegenerative disease. Age of onset, disease duration, age of death, neuropathological subtype of Lewy body disease according to McKeith/Newcastle criteria3 and Braak neurofibrillary tangle stage were recorded9 (Figure 1). In addition, we assessed Lewy body Braak stages,10 Aβ phases11 and stages of cerebral amyloid angiopathy.12 Of note, none of the cases showed intracytoplasmic TAR DNA-binding protein 43 (TDP-43) inclusions indicative for frontotemporal lobar degeneration associated with TDP-43 pathology, nor were there neuropathological features consistent with other types of frontotemporal lobar degeneration (see additional Supplementary Methods).
Clinical and pathological characteristics of the 91 dementia with Lewy body (DLB) cases. Top left: frequency of each pathological category (BS, brain stem; L, limbic; N, neocortical; UC, unclassified). Top right: BRAAK neurofibrillary tangle stage of patients (UC, unclassified). Bottom: table of the clinical and pathological data for all the 91 cases of DLB. Data are mean (s.d.). Motor features were defined by documented evidence of a Parkinsonian movement disorder by an assessing clinician.
DNA extraction and exome sequencing
DNA was extracted from cerebellum in all the cases. Illumina TruSeq 62 Mb exome capture and sequencing (Illumina Hiseq2000, 100 bp paired-end reads) was performed as described (see additional Supplementary Methods).
Known disease genes were defined as those previously shown to cause monogenic forms of Parkinson’s disease (PD), AD, frontotemporal lobar dementia and amyotrophic lateral sclerosis (Table 1). Variants were selected with a minor allele frequency of <0.01 international reference databases. Variants were defined as (1) pathogenic, (2) likely pathogenic, (3) of uncertain significance or (4) benign according to American College of Medical Genetics criteria13 (Table 1).
For completeness, exonic variants in genes previously associated with DLB (GBA, APOE, SNCA and SCARB2),5, 6, 7 AD (APOE, TREM2)14 or PD (LRRK2, GBA)15 were also identified in DLB cases and compared with 93 in-house unrelated disease control exomes.
Results
The mean exome sequencing base coverage depth was 84-fold (s.d.=13) in the 91 DLB cases and 76-fold (s.d.=12) in the 93 controls. There was no difference in the proportion of the exome target covered at >30-fold depth between DLB cases and controls (DLB 84%, s.d.=5; controls 84%, s.d.=3, P=0.588).
Known mendelian disease genes
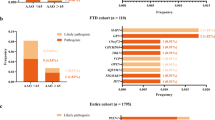
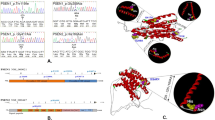
A total 18 rare heterozygous mutations in 25 patients were observed in genes previously shown to cause autosomal dominant forms of neurodegeneration (Tables 1, 2 and Supplementary Table S1). Three of these variants have been described in patients with AD, PD or frontotemporal lobar degeneration and amyotrophic lateral sclerosis (Patient A:PSEN2 p.D439A;16, 17 B:CHMP2B p.I29V;18 and C:SQSTM1 p.A33V,19, 20). In two additional cases (Patient E:EIF4G1 p.M1134V and F:SQSTM1 p.P27L), variants in known disease genes affecting highly conserved residues and predicted to be pathogenic by in silico software algorithms, were deemed of uncertain significance. Two patients also had variants of uncertain significance in GIGYF2, which is also implicated in PD (H:GIGYF2 p.S66T; G:GIGYF2 p.S1029C, Table 2). In genes causing autosomal recessive PD, AD or frontotemporal dementia and amyotrophic lateral sclerosis, only one rare compound heterozygous mutation in PARK2 was seen (Patient D, p.R275W/p.G430D).
Only patient A had a relevant family history (father affected—deceased and no tissue/DNA available). A clinical description of these cases is shown in the Supplementary Information. All showed typical DLB pathology with cortical LB being present and moderate AD pathology (Table 2).
The mean age at the presentation for the four cases with previously described pathogenic mutations (Patients A–D) was 78.25 years (s.d.=8.05). Motor symptoms developed in three cases (Patient A, B and D) at a mean of 1.33 years (s.d.=0.58) after the onset of cognitive symptoms. When patients E and F were included, the mean age of onset was 78.6 (s.d.=6.68), with motor symptoms developing in four patients (A, B, D and E), and a mean disease duration of 2.3 (s.d.=1.16) years.
Major risk alleles
GBA, TREM2 and LRRK2 had >80% coverage at 30-fold depth in both DLB cases and controls. APOE coverage was poor (DLB, 46.2%; controls 48.7% at 30-fold depth) and was therefore genotyped independently (see additional Supplementary Methods). After removing the previously described pathogenic alleles, APOE ɛ4 was significantly associated with DLB compared with controls (n=87, P=0.0001, Table 3). Ten DLB cases had one of five heterozygous GBA variants, compared with only three controls (P=0.043). Two GBA variants known to be risk factors for PD (p.L370P and p.N296S) were seen only in four patients and no controls. Two patients had variants in SCARB2 compared with six controls, and no SNCA variants were seen. There was no association between DLB and variants in SCARB2, LRRK2 or TREM2 (Supplementary Table S2).
Although there was no difference in the age of onset of DLB in APOE ɛ4 allele carriers when compared with non-APOE ɛ4 allele carriers (P=0.227), the APOE ɛ4 allele carriers had a shorter disease duration following diagnosis (P=0.036), and died at an earlier age (P=0.005) than non-APOE ɛ4 carriers (Figure 2, Table 3). There was no association between the presence of variants in GBA, SNCA, SCARB2, LRRK2, PARK2 or ATP13A2 and age of onset, disease duration, age of death, neurofibrillary Braak stage or the presence of motor symptoms.
Kaplan–Meier survival curves for DLB patients by APOE allele. Kaplan–Meier survival curves for DLB patients by APOE allele carrying at least one APOE ɛ4 allele (n=43, blue line), compared with non-APOE ɛ4 carriers (n=39, green line). Despite there being no significant difference in the age of onset of the DLB (see Results), APOE ɛ4 carriers (a) lived for a shorter period of time following diagnosis (P=0.036, log rank, Mantel–Cox test), and thus (b) died at a younger age (P=0.005, log rank, Mantel–Cox test) that non-APOE ɛ4 carriers. DLB, dementia with Lewy body.
Discussion
Exome sequencing of 91 cases of pathologically confirmed DLB identified four patients harbouring previously described pathogenic mutations neurodegenerative disease genes based on current diagnostic criteria (PSEN2, CHMP2B, SQSTM1, PARK2); possible pathogenic mutations in two (EIF4G1 and SQSTM1); and two further cases with mutations in GIGYF2, which has previously been associated with autosomal dominant PD. The central question is: are these variants causing DLB, or are they co-incidental findings? The role of GIGYF2 in PD remains contentious,21 and the p.D439A variant in PSEN2 may have incomplete penetrance,17 and is thus found in control databases along with the CHMP2B and SQSTM1 variants. Providing definitive proof of pathogenicity is therefore challenging, and there are arguments in both directions.
On one hand, the variants detected in PSEN2, CHMP2B, SQSTM1 and PARK2 are exceptionally rare in the general population.22 Given the clinical, pathological and mechanistic overlap between DLB and the neurodegenerative disorders where these disease genes were first described, it is plausible that they are contributing to the neuropathology. For example, in families with familial AD due to PSEN2 mutations, up to 64% of cases have extensive Lewy body deposition at autopsy.23 The CHMP2B protein has been shown to be found in association with Lewy bodies in post-mortem cases of DLB,24 and SQSTM1 deficiency has been shown to enhance α-synuclein accumulation in mice.25 The SQSTM1 p.A33V variant was previously described in five cases of frontotemporal dementia.19, 20 Recently, this allele was also detected in a patient with young-onset AD.26 Although seen in 0.0012% of controls, the p.A33V variant has now been seen in 8/1060 (0.007%) of patients with a neurodegenerative disease (including our study)19, 20,26 suggesting a broad association with neurodegenerative disorders (P=0.0037, chi squared with Yate’s correction). These findings support the notion that rare, incompletely penetrant pathogenic alleles cause overlapping syndromes of neurodegeneration, perhaps explaining why previously ascribed variants for frontotemporal dementia were also found in our DLB cases. Pathogenic mutations with a reduced penetrance will also be detected in healthy individuals (as for PSEN2 p.D439A17), and their presence in a control cohort does not preclude their potential to cause disease.22 This may explain why none of the four patients harbouring established pathogenic mutations reported a relevant family history.
On the other hand, the clinical and pathological phenotype of these five cases was wholly typical of DLB: how can this be reconciled with known pathogenic compound heterozygous mutations in PARK2, which typically presents with dystonia in early adult life? These findings highlight the challenges of using exome or whole-genome sequencing in a clinical context: is rare pathogenic mutation in a known disease gene more likely to be causing a variant phenotype, or is the phenotype so unusual that the variants must be a co-incidental finding? This will be difficult to resolve in individual cases, but the ongoing reporting of rare putative disease alleles, linked to rich phenotypic data, is an essential step in generating global data sets, which will ultimately provide definitive evidence of pathogenicity.22
Although the size of our study cohort limited the potential to discover new disease genes and risk loci, and did not include exclusion of repeat expansions such as C9orf72, we saw enrichment of GBA alleles and APOE ɛ4 alleles in DLB. In total, 48 patients (55.2%) possessed an APOE ɛ4 allele, with 5 (5.7%) having a variant in GBA, together with four (4.4%) having likely pathogenic alleles (potentially with incomplete penetrance). Therefore, 62.6% of patients harbour a risk factor or potentially pathogenic allele. This could explain why DLB is a relatively common disorder in the population, with an increased risk of disease within families, but few pedigrees suggestive of highly penetrant alleles. Finally, the association between APOE genotype and clinical progression has, to our knowledge, not been previously described, and has implications for cohort stratification in treatment studies.
References
Rahkonen T, Eloniemi-Sulkava U, Rissanen S, Vatanen A, Viramo P, Sulkava R . Dementia with Lewy bodies according to the consensus criteria in a general population aged 75 years or older. J Neurol Neurosurg Psychiatry 2003; 74: 720–724.
Bostrom F, Jonsson L, Minthon L, Londos E . Patients with Lewy body dementia use more resources than those with Alzheimer's disease. Int J Geriatr Psychiatry 2007; 22: 713–719.
McKeith IG, Dickson DW, Lowe J, Emre M, O'Brien JT, Feldman H et al. Diagnosis and management of dementia with Lewy bodies: third report of the DLB Consortium. Neurology 2005; 65: 1863–1872.
Nervi A, Reitz C, Tang MX, Santana V, Piriz A, Reyes D et al. Familial aggregation of dementia with Lewy bodies. Arch Neurol 2011; 68: 90–93.
Lane R, He Y, Morris C, Leverenz JB, Emre M, Ballard C . BuChE-K and APOE epsilon4 allele frequencies in Lewy body dementias, and influence of genotype and hyperhomocysteinemia on cognitive decline. Mov Disord 2009; 24: 392–400.
Clark LN, Kartsaklis LA, Wolf Gilbert R, Dorado B, Ross BM, Kisselev S et al. Association of glucocerebrosidase mutations with dementia with lewy bodies. Arch Neurol 2009; 66: 578–583.
Bras J, Guerreiro R, Darwent L, Parkkinen L, Ansorge O, Escott-Price V et al. Genetic analysis implicates APOE, SNCA and suggests lysosomal dysfunction in the etiology of dementia with Lewy bodies. Hum Mol Genet 2014; 23: 6139–6146.
Harding AJ, Das A, Kril JJ, Brooks WS, Duffy D, Halliday GM . Identification of families with cortical Lewy body disease. Am J Med Genet B Neuropsychiatr Genet 2004; 128: 118–122.
Braak H, Alafuzoff I, Arzberger T, Kretzschmar H, Del Tredici K . Staging of Alzheimer disease-associated neurofibrillary pathology using paraffin sections and immunocytochemistry. Acta Neuropathol 2006; 112: 389–404.
Alafuzoff I, Ince PG, Arzberger T, Al-Sarraj S, Bell J, Bodi I et al. Staging/typing of Lewy body related alpha-synuclein pathology: a study of the BrainNet Europe Consortium. Acta Neuropathol 2009; 117: 635–652.
Thal DR, Rub U, Orantes M, Braak H . Phases of A beta-deposition in the human brain and its relevance for the development of AD. Neurology 2002; 58: 1791–1800.
Thal DR, Ghebremedhin E, Orantes M, Wiestler OD . Vascular pathology in Alzheimer disease: correlation of cerebral amyloid angiopathy and arteriosclerosis/lipohyalinosis with cognitive decline. J Neuropathol Exp Neurol 2003; 62: 1287–1301.
Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med 2015; 17: 405–423.
Guerreiro R, Wojtas A, Bras J, Carrasquillo M, Rogaeva E, Majounie E et al. TREM2 variants in Alzheimer's disease. N Engl J Med 2013; 368: 117–127.
An XK, Peng R, Li T, Burgunder JM, Wu Y, Chen WJ et al. LRRK2 Gly2385Arg variant is a risk factor of Parkinson's disease among Han-Chinese from mainland China. Eur J Neurol 2008; 15: 301–305.
Lleo A, Blesa R, Gendre J, Castellvi M, Pastor P, Queralt R et al. A novel presenilin 2 gene mutation (D439A) in a patient with early-onset Alzheimer's disease. Neurology 2001; 57: 1926–1928.
Sassi C, Guerreiro R, Gibbs R, Ding J, Lupton MK, Troakes C et al. Investigating the role of rare coding variability in Mendelian dementia genes (APP, PSEN1, PSEN2, GRN, MAPT, and PRNP) in late-onset Alzheimer's disease. Neurobiol Aging 2014; 35: 2881.e1–2881.e6.
Parkinson N, Ince PG, Smith MO, Highley R, Skibinski G, Andersen PM et al. ALS phenotypes with mutations in CHMP2B (charged multivesicular body protein 2B). Neurology 2006; 67: 1074–1077.
Fecto F, Yan J, Vemula SP, Liu E, Yang Y, Chen W et al. SQSTM1 mutations in familial and sporadic amyotrophic lateral sclerosis. Arch Neurol 2011; 68: 1440–1446.
Le Ber I, Camuzat A, Guerreiro R, Bouya-Ahmed K, Bras J, Nicolas G et al. SQSTM1 mutations in French patients with frontotemporal dementia or frontotemporal dementia with amyotrophic lateral sclerosis. JAMA Neurol 2013; 70: 1403–1410.
Bras J, Simon-Sanchez J, Federoff M, Morgadinho A, Januario C, Ribeiro M et al. Lack of replication of association between GIGYF2 variants and Parkinson disease. Hum Mol Genet 2009; 18: 341–346.
MacArthur DG, Manolio TA, Dimmock DP, Rehm HL, Shendure J, Abecasis GR et al. Guidelines for investigating causality of sequence variants in human disease. Nature 2014; 508: 469–476.
Leverenz JB, Fishel MA, Peskind ER, Montine TJ, Nochlin D, Steinbart E et al. Lewy body pathology in familial Alzheimer disease: evidence for disease- and mutation-specific pathologic phenotype. Arch Neurol 2006; 63: 370–376.
Tanikawa S, Mori F, Tanji K, Kakita A, Takahashi H, Wakabayashi K . Endosomal sorting related protein CHMP2B is localized in Lewy bodies and glial cytoplasmic inclusions in alpha-synucleinopathy. Neurosci Lett 2012; 527: 16–21.
Tanji K, Odagiri S, Miki Y, Maruyama A, Nikaido Y, Mimura J et al. p62 deficiency enhances alpha-synuclein pathology in mice. Brain Pathol 2014; 25: 552–564.
Cuyvers E, van der Zee J, Bettens K, Engelborghs S, Vandenbulcke M, Robberecht C et al. Genetic variability in SQSTM1 and risk of early-onset Alzheimer dementia: a European early-onset dementia consortium study. Neurobiol Aging 2015; 36: 2005.e15–2005.e22.
Acknowledgements
This study was funded by the NHS National Institute of Health Research Biomedical Research Unit for Lewy body dementia at Newcastle upon Tyne Hospitals NHS Foundation Trust and Newcastle University. Tissue for this study was provided by Newcastle Brain Tissue Resource, which is funded in part by a grant from the UK Medical Research Council and by Brains for Dementia Research, a joint venture between Alzheimer’s Society and Alzheimer’s Research UK. MJK is a Wellcome Trust Clinical Research Training Fellow. PFC is a Wellcome Trust Senior Fellow in Clinical Science and National Institute for Health Research Senior Investigator. He receives funding from the Medical Research Council and the National Institute for Health Research Biomedical Research Centre for Ageing and Age-Related Disease award to the Newcastle upon Tyne Foundation Hospitals National Health Service Trust. The funding sources had no role in study design, data collection/analysis, the writing of the paper or the decision of when or where to publish it. The views expressed here are the views of the authors and not necessarily those of the NHS, NIHR or the Department of Health.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no conflict of interest.
Additional information
Supplementary Information accompanies the paper on the Translational Psychiatry website
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Keogh, M., Kurzawa-Akanbi, M., Griffin, H. et al. Exome sequencing in dementia with Lewy bodies. Transl Psychiatry 6, e728 (2016). https://doi.org/10.1038/tp.2015.220
Received:
Revised:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/tp.2015.220
This article is cited by
-
The presence and co-incidence of geriatric syndromes in older patients with mild-moderate Lewy body dementia
BMC Neurology (2022)
-
Effect of GBA gene variants on clinical characteristics of dementia with Lewy bodies: a review and meta-analyses
Neurological Sciences (2022)
-
Altered ceramide metabolism is a feature in the extracellular vesicle-mediated spread of alpha-synuclein in Lewy body disorders
Acta Neuropathologica (2021)
-
Dementia with Lewy bodies: an update and outlook
Molecular Neurodegeneration (2019)
-
Dementia with Lewy bodies — from scientific knowledge to clinical insights
Nature Reviews Neurology (2019)