Featured
-
-
Article |
 ALS-causative mutations in FUS/TLS confer gain and loss of function by altered association with SMN and U1-snRNP
ALS-causative mutations in FUS/TLS confer gain and loss of function by altered association with SMN and U1-snRNP
Dominant mutations in the RNA-binding protein FUS/TLS cause amyotrophic lateral sclerosis (ALS), an adult-onset motor neuron degenerative disease. Here, the authors show that ALS-causative FUS/TLS mutations directly bind the SMN and U1-snRNP complexes, producing both loss and gain of function effects on RNA processing.
- Shuying Sun
- , Shuo-Chien Ling
- & Don W. Cleveland
-
Article |
 Prevalent and distinct spliceosomal 3′-end processing mechanisms for fungal telomerase RNA
Prevalent and distinct spliceosomal 3′-end processing mechanisms for fungal telomerase RNA
In fission yeast, the telomerase RNA (TER) is produced through inhibition of the second step in splicing, resulting in spliceosomal cleavage. Here, the authors show that the inhibition of splicing is a conserved principle in fungal TER maturation that uses distinct molecular mechanisms across species.
- Xiaodong Qi
- , Dustin P. Rand
- & Julian J. -L. Chen
-
Article
| Open Access Diverse mechanisms for spliceosome-mediated 3′ end processing of telomerase RNA
Diverse mechanisms for spliceosome-mediated 3′ end processing of telomerase RNA
In fission yeast, the telomerase RNA (TER) is produced through spliceosomal cleavage. Here, Kannan et al. find that spliceosome-generated 3′ ends also occurs in other fungal TERs using distinct molecular mechanisms, suggesting multiple origins for this type of TER maturation pathway.
- Ram Kannan
- , Rachel M. Helston
- & Peter Baumann
-
Article |
 An artificial PPR scaffold for programmable RNA recognition
An artificial PPR scaffold for programmable RNA recognition
Pentatricopeptide repeat (PPR) proteins bind RNA and control diverse aspects of RNA metabolism in eukaryotic cells. Here, Coquille et al.present the crystal structures of several engineered PPR domains, elucidate their RNA binding mode and suggest paths to the design of modular, sequence-specific PPR domains.
- Sandrine Coquille
- , Aleksandra Filipovska
- & Oliver Rackham
-
Article |
 Unique features of the m6A methylome in Arabidopsis thaliana
Unique features of the m6A methylome in Arabidopsis thaliana
Modification of mRNA with N6-methyladenosine (m6A) is proposed to regulate transcript stability. Here, Jia et al. uncover plant-specific features in the m6A methylome of Arabidopsis, such as methylation enrichment around the start codon, and suggest a positive role in gene expression.
- Guan-Zheng Luo
- , Alice MacQueen
- & Chuan He
-
Article |
 Global 3′ UTR shortening has a limited effect on protein abundance in proliferating T cells
Global 3′ UTR shortening has a limited effect on protein abundance in proliferating T cells
The use of alternative polyadenylation sites can potentially result in mRNA being more or less susceptible to interaction with modulators of translation or stability. Here Gruber et al. find that the shortening of 3′UTRs observed in proliferating T cells does not significantly impact protein abundance.
- Andreas R. Gruber
- , Georges Martin
- & Mihaela Zavolan
-
Article |
 Autophagy supports genomic stability by degrading retrotransposon RNA
Autophagy supports genomic stability by degrading retrotransposon RNA
Retrotransposons replicate via a copy-paste mechanism involving a cytoplasmic intermediate. Guo et al.report that autophagy can suppress genetic variation by degrading cytoplasmic retrotransposon RNA, suggesting additional means by which autophagy may influence tumorigenesis.
- Huishan Guo
- , Maneka Chitiprolu
- & Derrick Gibbings
-
Article
| Open Access Caste-specific RNA editomes in the leaf-cutting ant Acromyrmex echinatior
Caste-specific RNA editomes in the leaf-cutting ant Acromyrmex echinatior
Post-translational mRNA editing has the potential to enhance the diversity of gene products and alter the functional properties of proteins. Here, Li et al. provide evidence that RNA editing is involved in generating caste-specific contrasting phenotypes in the leaf-cutting ant Acromyrmex echinatior.
- Qiye Li
- , Zongji Wang
- & Guojie Zhang
-
Article
| Open Access Human Tra2 proteins jointly control a CHEK1 splicing switch among alternative and constitutive target exons
Human Tra2 proteins jointly control a CHEK1 splicing switch among alternative and constitutive target exons
RNA binding proteins are key regulators of alternative splicing. Here, Best et al. show that the human Tra2α and Tra2ß RNA binding proteins jointly contribute to the control of constitutive and alternative splicing events to regulate essential biological processes including the response to DNA damage.
- Andrew Best
- , Katherine James
- & David J. Elliott
-
Article
| Open Access A genome-wide map of hyper-edited RNA reveals numerous new sites
A genome-wide map of hyper-edited RNA reveals numerous new sites
Common methods to detect adenosine-to-inosine RNA editing sites rely on mapping short RNA reads to the genome while allowing only a limited number of mismatches. Here, Porath et al. present a novel RNA-seq based approach to identify hyper-edited reads that significantly expands the RNA editome.
- Hagit T. Porath
- , Shai Carmi
- & Erez Y. Levanon
-
Article
| Open Access Identification of genetic variants associated with alternative splicing using sQTLseekeR
Identification of genetic variants associated with alternative splicing using sQTLseekeR
RNA sequencing has enabled the global analysis of both gene expression levels and splicing events. Here, the authors develop a multivariate approach that is able to identify SNPs that influence splicing, and investigate the overlap of these with functional domains across the genome, including previously identified GWAS signals.
- Jean Monlong
- , Miquel Calvo
- & Roderic Guigó
-
Article |
 BCLAF1 and its splicing regulator SRSF10 regulate the tumorigenic potential of colon cancer cells
BCLAF1 and its splicing regulator SRSF10 regulate the tumorigenic potential of colon cancer cells
Alternative splicing often alters the biological function of proteins. Here, Zhou et al. show that the splicing factor SRSF10 directs the inclusion of exon5a in Bcl-2-associated transcription factor 1, and that this drives cell growth and tumorigenic potential in human colon cancer cells.
- Xuexia Zhou
- , Xuebing Li
- & Ying Feng
-
Article
| Open Access EXOSC8 mutations alter mRNA metabolism and cause hypomyelination with spinal muscular atrophy and cerebellar hypoplasia
EXOSC8 mutations alter mRNA metabolism and cause hypomyelination with spinal muscular atrophy and cerebellar hypoplasia
The exosome is responsible for mRNA degradation, which is an important step in the regulation of gene expression. Here the authors report that homozygous missense mutations in the exosome subunit, EXOSC8, may cause neurodegenerative disease in infants through the dysregulation of myelin expression.
- Veronika Boczonadi
- , Juliane S. Müller
- & Rita Horvath
-
Article |
 Mutations in the PQBP1 gene prevent its interaction with the spliceosomal protein U5–15kD
Mutations in the PQBP1 gene prevent its interaction with the spliceosomal protein U5–15kD
Frameshift mutations in the protein polyglutamine tract-binding protein 1 (PQBP1) are believed to cause X-linked mental retardation. Here, Mizuguchi et al.present the crystal structure of a C-terminal fragment of PQBP1 in complex with the spliceosomal protein U5–15kD, and show details of this interaction that can lead to mechanistic insights into the disease.
- Mineyuki Mizuguchi
- , Takayuki Obita
- & Hitoshi Okazawa
-
Article |
 Alternative splicing regulates vesicular trafficking genes in cardiomyocytes during postnatal heart development
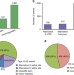
Alternative splicing regulates vesicular trafficking genes in cardiomyocytes during postnatal heart development
Alternative splicing is a process during gene expression that increases the diversity of proteins encoded by a single gene. Here, the authors perform RNA-sequencing on cardiac cells from mice and show that extensive changes in gene expression and alternative splicing occur during the first month after birth.
- Jimena Giudice
- , Zheng Xia
- & Thomas A. Cooper
-
Article
| Open Access Phosphoregulation of Ire1 RNase splicing activity
Phosphoregulation of Ire1 RNase splicing activity
Ire1 is an effector of the unfolded protein response that is activated upon stress to maintain protein homeostasis. Here, Prischi et al. demonstrate that phosphorylation of Ire1 within its kinase activation loop increases its RNAse activity, thus identifying a regulatory cross-talk between the two domains.
- Filippo Prischi
- , Piotr R. Nowak
- & Maruf M. U. Ali
-
Article
| Open Access Regulation of human telomerase splicing by RNA:RNA pairing
Regulation of human telomerase splicing by RNA:RNA pairing
Telomerase activity can be regulated by alternative splicing of its catalytic subunit TERT. Here, Wong et al. demonstrate that TERTsplicing is regulated via RNA:RNA pairing of repetitive intronic sequences with the pre-mRNA, thus revealing a new function for conserved elements embedded within introns.
- Mandy S. Wong
- , Jerry W. Shay
- & Woodring E. Wright
-
Article |
 The splicing activator DAZAP1 integrates splicing control into MEK/Erk-regulated cell proliferation and migration
The splicing activator DAZAP1 integrates splicing control into MEK/Erk-regulated cell proliferation and migration
DAZAP1 is a multi-functional RNA-binding protein that affects diverse aspects of RNA metabolism. Here, Choudhury et al.demonstrate that DAZAP1 acts as a general splice activator regulated by MEK/ERK signalling and thereby serves to integrate splice control into the regulation of cellular proliferation.
- Rajarshi Choudhury
- , Sreerupa Ghose Roy
- & Zefeng Wang
-
Article |
 Quality control of spliced mRNAs requires the shuttling SR proteins Gbp2 and Hrb1
Quality control of spliced mRNAs requires the shuttling SR proteins Gbp2 and Hrb1
In eukaryotic cells, export of unprocessed pre-mRNAs is prevented by the nuclear surveillance machinery. Here Hackmann et al.identify the SR proteins Gbp2 and Hrb1 as two novel quality control factors for spliced mRNAs that determine their degradation or nuclear export.
- Alexandra Hackmann
- , Haijia Wu
- & Heike Krebber
-
Article
| Open Access Structural analysis of human 2′-O-ribose methyltransferases involved in mRNA cap structure formation
Structural analysis of human 2′-O-ribose methyltransferases involved in mRNA cap structure formation
Human mRNA transcripts possess a 5' cap structure that is modified by methylation. Here, Smietanski et al.present the structures of human methyltransferases responsible for this reaction, revealing key differences to their viral counterparts and thereby providing a framework for targeted drug design.
- Miroslaw Smietanski
- , Maria Werner
- & Janusz M. Bujnicki
-
Article |
 Cyclin D1 induction of Dicer governs microRNA processing and expression in breast cancer
Cyclin D1 induction of Dicer governs microRNA processing and expression in breast cancer
Whether microRNA processing mediated by Dicer is regulated in a cell-cycle-dependent manner is unknown. Here, Chen et al.show that Cyclin D1, which is important in the control of the cell cycle, regulates the expression of Dicer, and that Cyclin D1 and Dicer expression levels correlate in breast cancer.
- Zuoren Yu
- , Liping Wang
- & Richard G. Pestell
-
Article |
 RNA editing regulates transposon-mediated heterochromatic gene silencing
RNA editing regulates transposon-mediated heterochromatic gene silencing
The Hoppel transposable element mediates heterochromatin formation in Drosophila. Here Savva et al. report that the RNA-editing enzyme, ADAR, edits a long double-stranded RNA generated by the Hoppeltransposon, thereby regulating heterochromatin formation and gene expression.
- Yiannis A. Savva
- , James E. C. Jepson
- & Robert A. Reenan
-
Article |
 MBNL1 and RBFOX2 cooperate to establish a splicing programme involved in pluripotent stem cell differentiation
MBNL1 and RBFOX2 cooperate to establish a splicing programme involved in pluripotent stem cell differentiation
MBNL and FOX splicing factors are known to have a role in the differentiation of muscle and the nervous system during development. In this study, the authors show that MBNL1 and RBFOX2 regulate alternative splicing of genes that are required specifically for late mesoderm differentiation.
- Julian P. Venables
- , Laure Lapasset
- & Jamal Tazi
-
Article |
 Mechanism for full-length RNA processing of Arabidopsis genes containing intragenic heterochromatin
Mechanism for full-length RNA processing of Arabidopsis genes containing intragenic heterochromatin
Transposable elements found within transcribed regions of genes are often compacted into heterochromatin. Using Arabidopsisas a model, these authors show that the protein, IBM2, is required for correct processing of genes that contain intragenic heterochromatin.
- Hidetoshi Saze
- , Junko Kitayama
- & Tetsuji Kakutani
-
Article
| Open Access Endonuclease V cleaves at inosines in RNA
Endonuclease V cleaves at inosines in RNA
Bacterial endonuclease V enzymes are characterized as DNA repair proteins. Here the authors show that human endonuclease V is an inosine-specific ribonuclease, indicating a role for this enzyme in normal RNA metabolism rather than DNA repair.
- Erik Sebastian Vik
- , Meh Sameen Nawaz
- & Ingrun Alseth
-
Article
| Open Access Human endonuclease V is a ribonuclease specific for inosine-containing RNA
Human endonuclease V is a ribonuclease specific for inosine-containing RNA
In Escherichia coli, the highly conserved enzyme endonuclease V has a role in DNA repair. Here the authors show that human endonuclease V is an inosine 3' endoribonuclease and that Tudor Staphylococcal nuclease enhances this activity, suggesting a role for human endonuclease V in RNA metabolism.
- Yoko Morita
- , Toshihiro Shibutani
- & Isao Kuraoka
-
Article |
 Tertiary structural elements determine the extent and specificity of messenger RNA editing
Tertiary structural elements determine the extent and specificity of messenger RNA editing
A central, imperfect duplex RNA secondary structure is generally required for site-specific adenosine-to-inosine RNA editing by ADAR enzymes. Rieder et al. show in Drosophila that conserved and complex long-range RNA tertiary structures form in vivoand can also regulate specific RNA-editing events by ADAR enzymes.
- Leila E. Rieder
- , Cynthia J. Staber
- & Robert A. Reenan
-
Article
| Open Access The thermodynamic patterns of eukaryotic genes suggest a mechanism for intron–exon recognition
The thermodynamic patterns of eukaryotic genes suggest a mechanism for intron–exon recognition
The thermodynamics of unwinding polynucleotide duplexes can be determined from energy changes for DNA and mRNA interactions. Here the authors show that the ratio between mRNA/DNA and DNA/DNA duplex stability upstream of the 3′- spice sites is a characteristic that can contribute to intron–exon recognition.
- Marina N. Nedelcheva-Veleva
- , Mihail Sarov
- & Stoyno S. Stoynov
-
Article |
 Induction and reversal of myotonic dystrophy type 1 pre-mRNA splicing defects by small molecules
Induction and reversal of myotonic dystrophy type 1 pre-mRNA splicing defects by small molecules
Myotonic dystrophy type 1 (DM1) is caused by defects in the alternative splicing of pre-mRNA. Childs-Disney and colleagues report two small molecules that either induce or reverse DM1-associated splicing defects by modulating the binding of pre-mRNA to muscleblind-like 1 protein.
- Jessica L. Childs-Disney
- , Ewa Stepniak-Konieczna
- & Matthew D. Disney
-
Article |
 FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA
FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA
Internal modifications in mRNA and non-coding RNA are necessary for modulating various intracellular signalling pathways. In this study, the authors report novel modifications resulting from oxidative RNA demethylation, which regulate RNA–protein interactions affecting gene expression.
- Ye Fu
- , Guifang Jia
- & Chuan He
-
Article |
 Splicing factor SRSF3 is crucial for hepatocyte differentiation and metabolic function
Splicing factor SRSF3 is crucial for hepatocyte differentiation and metabolic function
Splicing factors, such as the protein SRSF3, regulate mRNA metabolism but are hard to study in vivobecause genetic kockouts are usually lethal. Here, Sen and colleagues create mice with a hepatocyte-specific knockout of Srsf3 and demonstrate its role in hepatocyte differentiation and liver function.
- Supriya Sen
- , Hassan Jumaa
- & Nicholas J. G. Webster
-
Article
| Open Access An RNA architectural locus control region involved in Dscam mutually exclusive splicing
An RNA architectural locus control region involved in Dscam mutually exclusive splicing
Alternative splicing at the Drosophila Down syndrome cell adhesion molecule gene generates 38,016 isoforms, and underlies self-avoidance of growing neurons. Wang et al. identify a structure in the DSCAM mRNA that ensures mutually exclusive splicing and observe expansion of the structure with increasing number of exons during arthropod evolution.
- Xuebin Wang
- , Guoli Li
- & Yongfeng Jin
-
Article |
 TREX exposes the RNA-binding domain of Nxf1 to enable mRNA export
TREX exposes the RNA-binding domain of Nxf1 to enable mRNA export
The TREX complex and Nxf1 are involved in the export of mRNA from the nucleus but the precise molecular function of TREX is unclear. Here, the TREX components Aly and Thoc5 are shown to bind to Nxf1 resulting in a change in Nxf1 conformation that permits binding to mRNA and nuclear export.
- Nicolas Viphakone
- , Guillaume M. Hautbergue
- & Stuart A. Wilson
-
Article |
 Post-transcriptional spliceosomes are retained in nuclear speckles until splicing completion
Post-transcriptional spliceosomes are retained in nuclear speckles until splicing completion
It is unclear where in the nucleus splicing takes place and how much occurs post-transcriptionally. Using antibodies raised against a phosphorylated splicing factor, Girardet al. show that the majority of splicing occurs co-transcriptionally and that post-transcriptional splicing occurs in nuclear speckles.
- Cyrille Girard
- , Cindy L. Will
- & Reinhard Lührmann
-
Article |
 Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins
Reprogramming of tRNA modifications controls the oxidative stress response by codon-biased translation of proteins
In response to stress, yeast cells selectively translate proteins that can enhance cell survival. In this study, the authors find that tRNALEU(CAA)in yeast cells is modified in response to oxidative stress by a methyltransferase and this results in the selective translation of the mRNA for these genes.
- Clement T.Y. Chan
- , Yan Ling Joy Pang
- & Peter C. Dedon
-
Article
| Open Access SynGAP isoforms exert opposing effects on synaptic strength
SynGAP isoforms exert opposing effects on synaptic strength
Synaptic GTPase-activating protein, SynGAP, is a postsynaptic signalling protein that can regulate synaptic function. McMahonet al. express different SynGAP isoforms in neurons and find that the effect on synaptic strength depends on alternative promoter usage and alternative splicing of the C-terminus.
- A.C. McMahon
- , M.W. Barnett
- & P.C. Kind
-
Article |
 Alternative splicing of CD44 mRNA by ESRP1 enhances lung colonization of metastatic cancer cell
Alternative splicing of CD44 mRNA by ESRP1 enhances lung colonization of metastatic cancer cell
CD44 is a cell surface protein that is a marker for stem cell-like cancer cells and has a role in invasion and metastasis. Here, epithelial splicing regulatory protein 1 is shown to generate a CD44 variant protein that enhances metastasis in a mouse model and protects cells from oxidative stress.
- Toshifumi Yae
- , Kenji Tsuchihashi
- & Osamu Nagano
-
Article
| Open Access Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
Serine-7 but not serine-5 phosphorylation primes RNA polymerase II CTD for P-TEFb recognition
Phosphorylation of the carboxy-terminal domain of RNA polymerase II is important for controlling gene transcription. In this study, the transcription elongation factor Tefb is shown to phosphorylate serine-5 and its activity is enhanced when the polymerase is already phosphorylated on serine-7.
- Nadine Czudnochowski
- , Christian A. Bösken
- & Matthias Geyer
-
Article |
 Auto-regulatory RNA editing fine-tunes mRNA re-coding and complex behaviour in Drosophila
Auto-regulatory RNA editing fine-tunes mRNA re-coding and complex behaviour in Drosophila
Adars are adenosine deaminases that act on RNAs, including those encoding proteins involved in neuronal transmission and also Adar RNA. Here, Savvaet al. engineered knock-in Drosophila mutants with altered Adar autoediting and found that this changed the spectrum of adenosine deamination and Drosophilabehaviour.
- Yiannis A. Savva
- , James E.C Jepson
- & Robert A. Reenan
-
Article |
 The tRNA methyltransferase NSun2 stabilizes p16INK4 mRNA by methylating the 3′-untranslated region of p16
The tRNA methyltransferase NSun2 stabilizes p16INK4 mRNA by methylating the 3′-untranslated region of p16
The expression of the tumour suppressor p16 is frequently lost in cancer. Zhanget al. show in cultured cells that p16 mRNA levels are stabilised by methylation of the 3′-untranslated region by the tRNA methyltransferase NSun2, revealing a new mechanism for regulating p16.
- Xiaotian Zhang
- , Zhenyun Liu
- & Wengong Wang
-
Article
| Open Access A segmental genomic duplication generates a functional intron
A segmental genomic duplication generates a functional intron
The appearance of a new intron that splits an exon without disrupting the corresponding peptide sequence is a rare event in vertebrate genomes. Hellstenet al.demonstrate that, under certain circumstances, a functional intron can be produced in a single step by segmental genomic duplication.
- Uffe Hellsten
- , Julie L. Aspden
- & Daniel S. Rokhsar
-
Article
| Open Access Editing of human KV1.1 channel mRNAs disrupts binding of the N-terminus tip at the intracellular cavity
Editing of human KV1.1 channel mRNAs disrupts binding of the N-terminus tip at the intracellular cavity
RNA editing is important in regulating neuronal excitability, and a specific editing event has been shown to alter the permeation pathway of voltage-gate potassium channels. Gonzalezet al.find that the tip of the channel's inactivation gate makes a direct hydrophobic interaction with the edited position.
- Carlos Gonzalez
- , Angelica Lopez-Rodriguez
- & Miguel Holmgren
-
Article
| Open Access Predicting sites of ADAR editing in double-stranded RNA
Predicting sites of ADAR editing in double-stranded RNA
ADAR enzymes edit double-stranded RNA, converting adenosines to inosines, and are essential for neuronal function. Eggingtonet al. quantify edit sites in RNA using a Sanger sequencing protocol and use the resulting data to develop algorithms to predict RNA edit sites.
- Julie M. Eggington
- , Tom Greene
- & Brenda L. Bass
-
Article
| Open Access Chemical treatment enhances skipping of a mutated exon in the dystrophin gene
Chemical treatment enhances skipping of a mutated exon in the dystrophin gene
Duchenne muscular dystrophy is caused by a loss of thedystrophin gene, and control of dystrophin mRNA splicing could aid treatment of the disease. Nishida et al. show that a small molecule promotes skipping of exon 31 and increases production of a functional dystrophin protein in a patient.
- Atsushi Nishida
- , Naoyuki Kataoka
- & Masafumi Matsuo
-
Article |
 Two splice variants of the IDD14 transcription factor competitively form nonfunctional heterodimers which may regulate starch metabolism
Two splice variants of the IDD14 transcription factor competitively form nonfunctional heterodimers which may regulate starch metabolism
The alternative splicing of genes increases the number and diversity of proteins produced within a cell. Seoet al. demonstrate that the beta form of the alternatively spliced Arabidopsis gene, IDD14, is produced under cold conditions and may have a role in regulating starch accumulation.
- Pil Joon Seo
- , Mi Jung Kim
- & Chung-Mo Park
-
Article |
 A role for TREX components in the release of spliced mRNA from nuclear speckle domains
A role for TREX components in the release of spliced mRNA from nuclear speckle domains
The pre-mRNA splicing and TREX mRNA export machineries are found in nuclear speckle domains. Diaset al. microinject CMV-DNA constructs into cells and find that transcripts containing functional splice sites accumulate in nuclear speckles and that the TREX complex is required to release the mRNA once processed.
- Anusha P. Dias
- , Kobina Dufu
- & Robin Reed

 PRMT9 is a Type II methyltransferase that methylates the splicing factor SAP145
PRMT9 is a Type II methyltransferase that methylates the splicing factor SAP145