Abstract
N6-methyladenosine is a prevalent internal modification in messenger RNA and non-coding RNA affecting various cellular pathways. Here we report the discovery of two additional modifications, N6-hydroxymethyladenosine (hm6A) and N6-formyladenosine (f6A), in mammalian messenger RNA. We show that FeII- and α-ketoglutarate-dependent fat mass and obesity-associated (FTO) protein oxidize N6-methyladenosine to generate N6-hydroxymethyladenosine as an intermediate modification, and N6-formyladenosine as a further oxidized product. N6-hydroxymethyladenosine and N6-formyladenosine have half-life times of ~3 h in aqueous solution under physiological relevant conditions, and are present in isolated messenger RNA from human cells as well as mouse tissues. These previously unknown modifications derived from the prevalent N6-methyladenosine in messenger RNA, formed through oxidative RNA demethylation, may dynamically modulate RNA–protein interactions to affect gene expression regulation.
Similar content being viewed by others
Introduction
RNA modifications can affect molecular interaction and structural changes extending beyond local conformation of RNA. Not until recently were some of these modifications discovered to be reversible, dynamic and responsive to cellular stimuli, which suggests that they may participate in cellular regulatory processes1,2. N6-methyladenosine (m6A) is a widespread internal modification in messenger RNA (mRNA), which constitutes 1–2% of all adenosine in mRNA3. Over 12,000 m6A sites in the transcripts of more than 7,000 human genes have been characterized4,5. Conserved preference of m6A around stop codons, in 3′-untranslated region, and within long internal exons have been revealed in both human and mouse cells4,5. This modification is essential to cell survival and development in eukaryotes, and has a fundamental role in gene expression regulation6,7. Although the methyltransferase complex is thought to contain METTL3 in mammals8, the following have been identified as components of this complex in yeast: Ime4 (yeast m6A methyltransferase), Mum2 and Slz1 (9). We have discovered that the human fat mass and obesity-associated protein FTO catalyses the removal of m6A in vitro and in vivo, providing the first example of reversible methylation in RNA2. FTO, which was identified in several genome-wide-association studies to be associated with obesity and type II diabetes10,11,12, is a member of the non-haem FeII/α-ketoglutarate (α-KG)-dependent AlkB family demethylases that mainly catalyse oxidative demethylation of N-alkylated nucleic acid bases13,14,15.
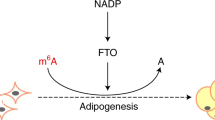
We seek to further understand the FTO-mediated oxidative demethylation of m6A (Fig. 1a) and identify potential new modifications that could be present inside mammalian cells as this family of enzymes exhibits versatile activities16. For instance, the recently discovered ten eleven translocation (TET) enzymes that belong to the general family of non-haeme FeII/α-KG-dependent dioxygenases can perform tandem oxidations17,18,19,20. Initially shown to catalyse the oxidation of 5-methylcytosine (5mC) to 5-hydroxymethylcytosine (5hmC) in mammalian genomic DNA17,18, TET proteins were found to further oxidize 5hmC to 5-formylcytosine (5fC) and 5-carboxylcytosine (5caC)19,20,21. 5hmC levels are highest in the brain extracts and 5hmC is thought to be a new DNA epigenetic marker22. The oxidation of 5hmC to 5fC and/or 5caC is a much slower process compared with the oxidation of 5mC to 5hmC (20). Nevertheless, the continuous oxidation of 5hmC by TET enzymes is essential to an active DNA demethylation pathway, and has important roles in regulating DNA epigenetics19,23.
(a) FTO demethylates m6A through oxidization of m6A to hm6A with f6A as a further oxidized product. (b) MALDI-TOF analysis of m6A-RNA after reacting with FTO. An unexpected monoisotopic peak of m6A+14 Da corresponding to f6A was observed after treating a 9mer m6A-RNA oligo (5 μM) with 10 μM of FTO at room temperature (RT) for 20 min. The m6A−2 Da peak represents the dehydration product of the putative hm6A intermediate during MALDI-TOF ionization.
Here we show that FTO oxidizes m6A to hm6A and f6A in RNA in a step-wise manner. The observation is further supported by molecular dynamic (MD) simulation, which indicates that FTO can bind to hm6A and m6A as substrates. We further demonstrate that hm6A and f6A are relatively stable under physiological conditions and can be detected in polyadenylated RNA isolated from human cells and mouse tissues. These meta-stable intermediates, generated during the course of the FTO-catalysed m6A demethylation and discovered for the first time inside mammalian cells, may have significant functional implications.
Results
New modifications in the FTO-mediated m6A oxidation
To investigate the FTO-catalysed demethylation process, we monitored the reaction with high-resolution matrix-assisted laser desorption-ionization time-of-flight mass (MALDI-TOF)/TOF mass spectrometry (MS) to detect hm6A, an expected oxidation product of m6A in RNA by FTO. Five micromolar of 9mer single-stranded RNA (ssRNA) containing m6A was treated with 10 μM of FTO at room temperature for 20 min, and analysed immediately after reaction (Fig. 1b). To our surprise, when analysing the monoisotopic peaks, other than the peak of demethylation product (m6A−14 Da), we also observed another m6A+14 Da peak representing an unknown reaction product (Fig. 1b). The putative hm6A product has a molecular weight of m6A+16 Da, which likely loses a H2O moiety (−18 Da) to form N6-methyleneadenosine (m6A−2 Da) during the process (Fig. 1b). We speculated that this unexpected +14 Da peak may correspond to N6-formyladenosine (f6A) as a result of a tandem oxidation of the m6A substrate in a process similar to that of the oxidation of 5mC to 5hmC and 5fC by the TET proteins in the DNA demethylation pathway. As the oxidation of hemiaminal to amide is a rare chemical transformation, and the enzymatic conversion of this process has never been reported, we proceeded to further confirm the identity of this unprecedented species.
We reasoned that in previous high-performance liquid chromatography (HPLC) analysis of the digested nucleosides, the basic conditions used during alkaline phosphatase digestion may accelerate the hydrolysis of this reaction product2. As FTO showed similar demethylation activity between internal and terminal m6A in RNA (Fig. 1b and Supplementary Fig. S1), we synthesized a 9mer ssRNA oligonucleotide with m6A incorporated at the 5′ end for further studies. In this case, we could analyse the first nucleoside at the 5′ position with only nuclease P1 digestion to avoid the basic conditions (Fig. 2a). Five micromolar of the substrate RNA was treated with 2.5 μM of FTO for 2 or 20 min at room temperature, and the reaction product was digested with nuclease P1 and run through a C18 reverse phase column on HPLC. Two new peaks other than m6A and A were observed (Fig. 2b). We suspected that the initially generated product was hm6A, and the latter product was f6A.
(a) A 9mer ssRNA containing a 5′ m6A is used for HPLC-MS/MS analysis. After the RNA is treated with FTO, it is digested by nuclease P1 to release the first base as a nucleoside for further HPLC-MS/MS analysis. The rest of the bases are left as nucleoside 5′-monophosphate (5′-NMP). (b) HPLC analysis of nuclease P1-digested 5′m6A-9mer RNA oligo (5 μM) after treating with FTO. m6A was converted to hm6A and f6A after the treatment with 2.5 μM of FTO for 2 min at room temperature; substantial conversion of hm6A to f6A can be observed in 20 min. These newly formed peaks coelute with hm6A and f6A standards, respectively. The last peak in hm6A standard is N6-methyleneadenosine, a dehydration product that coexists with hm6A. (c,d) Comparison of MS/MS profile of m6A-oxidation products with hm6A and f6A standards. (c) The initial product matches the elution time and MS/MS fragmentation pattern of hm6A. (d) The later-formed product matches the elution time and MS/MS fragmentation pattern of f6A. (e) Kinetic behaviour of the reaction indicates that the formation rate of f6A is slower than the oxidation rate from m6A to hm6A by a factor of 12.3±1.9. Error bars, mean±s.e.m. for n=4 experiments.
To confirm the identities of these species, we chemically synthesized hm6A and f6A standards following reported procedures (Supplementary Methods and Supplementary Fig. S2)24,25. Briefly, after treating 100 μM of adenosine with 30 mM of formaldehyde at 60 °C for 4 h, hm6A was produced, which partially dehydrated to N6-methylene-adenosine in equilibrium (Supplementary Fig. S2a)24. f6A could be prepared by treating N,N-dimethylformamidine-protected TBDMS-adenosine with weak acid N-hydroxybenzotriazole (HOBt·H2O) in MeOH (Supplementary Fig. S2b), followed by deprotection of the TBDMS group25. f6A is stable in dry organic solvent, and can be further characterized by high-resolution MS, nuclear magnetic resonance (NMR) and ultraviolet-visible (UV–vis) spectra analysis (Supplementary Figs S3 and S4). It has the highest absorbance at 274 nm with εmax (274 nm)=1.83 × 104 M−1 cm−1 (Supplementary Fig. S4) in water.
The two reaction products coelute with synthesized hm6A and f6A standards in HPLC (Fig. 2b), and have the same MS/MS fragmentation pattern as hm6A (Fig. 2c) and f6A (Fig. 2d), respectively, thus confirming their identities. Both hm6A and f6A decompose to A in aqueous solution, and the hydrolysis processes were accelerated under acidic or basic conditions (Supplementary Fig. S5). To measure the stability of hm6A and f6A under physiological conditions, hm6A- and f6A-containing RNA generated by FTO oxidation in situ were digested with nuclease P1, and diluted in buffer solutions mimicking the physiological conditions with Tris–HCl (20 mM, pH 7.4), KCl (140 mM), NaCl (15 mM) and MgCl2 (1 mM). The solutions were incubated at 37 °C and analysed by HPLC-MS/MS to monitor the degradation process. In the neutral buffered solution, long half-life times of 186±25 min for hm6A and 188±18 min for f6A were observed, respectively (Supplementary Fig. S6), which are comparable to the average half-life of mRNA in mammalian cells26. The relative long life times of hm6A and f6A under physiological conditions suggest that they are present inside living cells and may have functional roles other than as mere demethylation intermediates.
Reaction kinetics of FTO-mediated oxidation
To elucidate the kinetic behaviour of the production of hm6A and f6A, we measured a time-course reaction profile of FTO (2.5 μM) on the same RNA (5 μM) at 37 °C by quantitative LC-MS/MS (Fig. 2e and Supplementary Fig. S7). We found that similar to the oxidation of 5mC to 5hmC and then 5fC by Tet20, FTO catalyses the oxidation of hm6A to f6A at a relatively slower rate compared with the oxidation of m6A to hm6A (Fig. 2e). The kinetic behaviour of this two-step oxidation process suggests a non-processive oxidation pathway, which involves the releasing and rebinding of the hm6A intermediate to the FTO active site. Generation of f6A will be sensitive to the FTO concentration as the two-step reaction harbours a fast first step and a slow second step; higher local concentrations of FTO will result in the production of more f6A.
FTO recognizes m6A as well as hm6A as its substrate
To further rationalize our observations, we used MD simulation to model the binding of FTO with m6A and hm6A. Protein coordinates were retrieved from the crystal structure of FTO-N3-methylthymine (m3T) complex (PDB ID: 3LFM)27; initial base coordinates were set up in two independent ways. The first approach was to fix the sugar ring position of m3T in 3LFM, and mutate m3T to N3-methylcytosine (m3C), m6A and hm6A. These systems were labelled as FTO-m3T-cry, FTO-m3C-cry, FTO-m6A-cry and FTO-hm6A-cry, respectively. The second protocol involves aligning the AlkB–DNA complex crystal structures with FTO based on similarities in protein structure calculated with the Protein Comparison Tool28 (Supplementary Figs S8 and S9), and then using the coordinates of the lesioned bases from the aligned AlkB complexes29. We used PDB file 3O1O to derive the coordinates for m3T, 3O1M for m3C and 3O1P for εA, respectively; εA was then mutated to m6A or hm6A. These systems were labelled as FTO-m3T-alkb, FTO-m3C-alkb, FTO-m6A-alkb and FTO-hm6A-alkb, respectively (Supplementary Fig. S10).
We calculated the binding models of m6A (Supplementary Figs S11 and S12) and hm6A (Supplementary Figs S13 and S14) in FTO with more than 10-ns MD simulation, in comparison with m3T (Supplementary Figs S15 and S16) and m3C (Supplementary Figs S17 and S18) (Supporting methods). Similar to m3T, both m6A (Fig. 3a) and hm6A (Fig. 3b) are stable in the active site in two sets of MD simulations using two initial positions retrieved by two independent methods described above (left panels were calculated using the FTO-m3T crystal structure to prepare the initial coordinates of the base; right panels were calculated using aligned AlkB crystal structures to derive the initial base coordinates.). Overlays of these structures with the FTO-m3T crystal structure show that N6-methyl in m6A (Fig. 3c) and N6-methylene in hm6A (Fig. 3d) position near the iron centre in the active site of FTO similar to N3-methyl in m3T in the solved structure27. As a negative control, m3C was observed to be unstable in FTO and dissociated from the active site after 3 ns of simulation (Supplementary Figs S17 and S18), which was likely the result of the positive charge of m3C.
(a) Binding model of FTO-m6A. (b) Binding model of FTO-hm6A. m6A and hm6A is highlighted in dashed blue and red circles. (c) Binding model comparison of m6A overlay with m3T. (d) Binding model comparison of hm6A overlay with m3T. Positions of the targeting methyl in m6A and hm6A are highlighted in dashed blue and red circles, respectively; the targeting methyl in m3T is highlighted in dashed magenta circles. Protein structure is shown in cartoon and active-site residues in sticks. Left panels use the m3T coordinate as in the FTO crystal structure (PDB ID: 3LFM) to prepare the initial location of the base in the FTO simulation through the following steps: fix the sugar ring position in 3LFM and mutate m3T to m3C, m6A, and hm6A, respectively. Right panels use the base coordinates from aligned AlkB crystal structures (PDB ID: 3O1P for εA, and 3O1O for m3T): align FTO and AlkB, take the aligned positions of m3T and εA as the starting point and mutate εA to m6A and hm6A, respectively.
Relative binding free energies of m6A and hm6A to FTO were also calculated by alchemical free-energy simulations30 (Supplementary Methods, Supplementary Figs S19 and S20). All independent simulations predicted a slightly stronger binding (ranging from 0.4 to 2.8 kcal mol−1) for hm6A over m6A (Supplementary Table S1). These data support our observation of hm6A as a substrate for FTO, and suggest a similar oxidation model of m6A to hm6A for hm6A to f6A, in agreement with the kinetic behaviour of the oxidation reaction and a proposed non-processive tandem oxidation reaction pathway.
Detection of hm6A and f6A in human and mouse mRNA
To confirm the existence of these new modifications in vivo, we optimized RNA digestion conditions using RNase T1 followed by nuclease P1 at neutral pH, in which both hm6A and f6A are relatively stable. RNase T1 cleaves the phosphodiester bond after G, and exposes the following nucleotide with a free 5′-OH. Nuclease P1 releases this nucleotide for subsequent HPLC separation (Fig. 4a). Although m6A shows a preference in the consensus sequence of (G/A/U)(G>A)m6AC(U>A>C)4,5, this method can analyse hm6A and f6A present in this consensus sequence after G. Applying this method we analysed the isolated poly(A)-RNA from cultured HeLa cells and mouse liver tissues by LC-MS/MS. We observed peaks with the same elution times at the MS/MS detection channel of hm6A (298.1 fragmented to 136.0, Fig. 4b). 6A (296.1 fragmented to 164.1, Fig. 4c), respectively. We subsequently followed the f6A peak, which is more stable than hm6A. f6A in mouse liver decreased significantly upon incubating at room temperature overnight (Supplementary Fig. S21), similar to that observed with the f6A standard. Therefore, we confirmed the presence of hm6A and f6A in poly(A)-RNA in vivo. The concentration of the isolated f6A (a large portion of f6A may have decomposed during the isolation process) was estimated to be at least 0.5–1% of the total m6A (Supplementary Fig. S22). A similar amount of hm6A appears to also exist in vivo based on our results. Therefore, taking account of decomposition during isolation a noticeable portion of mRNA should carry either hm6A or f6A at any given time inside cells.
(a) Detection strategy: RNA is first digested with RNase T1 to expose the 5′ OH group of the base after G. Subsequent nuclease P1 digestion releases the first base as nucleoside, and the rest of the bases as nucleoside 5′-monophosphate (5′-NMP). (b,c) LC-MS/MS analysis of digested mRNA isolated from HeLa cell and mouse liver samples showing the presence of hm6A (b, 298.1→136.0, 3.1 min) and f6A (c, 296.1→164.1, 4.2 min).
hm6A and f6A may modulate RNA–protein interactions
As the oxidation of 5mC to 5hmC and then 5fC in DNA hinders binding to methyl-CpG-binding (MBD) proteins31, we envisioned that the extra oxygen group on hm6A and f6A may hinder their interaction with m6A-binding proteins. Recently, YTHDF2 protein has been implicated as a potential m6A-RNA-binding protein4. To evaluate its binding affinity to these RNA modifications, we cloned and expressed a GST-tagged YTHDF2 protein. The binding affinities of YTHDF2 to RNA containing hm6A or f6A, generated in situ by FTO-mediated oxidation, were measured and compared with those with m6A or A using electrophoretic mobility shift assay (Supplementary Fig. S23). About 70% of hm6A (Supplementary Fig. S24) and 60% of f6A (Supplementary Fig. S25) were estimated to be present in the electrophoretic mobility shift assay assay, respectively, as shown by LC-MS/MS analysis of an unlabelled 9mer RNA that contained 5′-m6A treated using the same procedure. We found that the YTHDF2 protein preferentially binds to m6A-RNA, while the binding affinities of YTHDF2 to hm6A or f6A were attenuated to a level similar to A (Supplementary Fig. S23).
Discussion
We show here that hm6A and f6A are new RNA modifications in mammalian cells, and that FTO can catalyse the formation of these modifications. The AlkB protein has been shown to oxidize m6A to hm6A as an oxidation intermediate in vitro with no further oxidation of hm6A observed32. FTO shows the unique activity of performing a second oxidation on the meta-stable hm6A to form an unprecedented f6A. Compared with other hemiaminal intermediates, hm6A is relatively stable because of the electron-rich nature of the exocyclic N6 nitrogen of adenosine when compared with endocyclic positions, which allows for further oxidation to f6A.
Although more than 100 RNA modifications have been discovered, the chemical diversity of the modifications is relatively limited. Recent application of more sensitive analytical methods leads to the discovery of RNA modifications with novel functional groups, such as geranylated RNA in bacteria33. The discovery of hydroxymethylated and formylated adenosines in mRNA here further expands the repertoire of chemical diversity of RNA modifications. As m6A has not only been detected in mRNA but also long non-coding RNA, it is possible that hm6A and f6A, which are derived from m6A, may also exist in long non-coding RNAs. FTO exhibits a relatively high activity towards m6A on ssRNA, but lower activity towards m6A in stem–loop structures and negligible activity against m6A on double-stranded RNA in vitro2. The in vivo activity could be affected by the secondary structure of the RNA and cellular machineries that recruit FTO to specific RNA substrates.
FTO, ALKBH5 (the second RNA demethylase recently discovered by us34), and the methyltransferase components orchestrate together to determine the modification status of mRNA. As FTO and ALKBH5 are partially localized in nuclear speckles2,34, in which the methyltransferase complex resides, the demethylated RNA could be remethylated by the methyltransferase in a direct reversible model. In contrast, the temporary existence of the hydroxymethyl and the formyl group on the N6 position of adenosine may represent a delayed reversible model, in which an additional diffusion/export step may be incorporated and coupled with the ‘slow’ hydrolysis step (Fig. 5). Therefore, these meta-stable intermediates could provide a time-window for the oxidized RNA to diffuse or be exported out of the nuclear speckles, thus eliminating the possibility of remethylation by the methyltransferase complex at the same loci in vivo (Fig. 5). It is also interesting to speculate about other functional roles of hm6A and f6A because they are not recognized by m6A-binding proteins such as YTHDF2. They could serve as another modification marker to recruit different sets of RNA-binding proteins, and differentially regulate the subsequent RNA-related pathways (Fig. 5), or they could serve as markers for nascent RNAs taking advantage of their intrinsic degradation kinetics. It is also interesting to note that the FTO-mediated formation of hm6A and f6A is similar to the TET-mediated oxidation of 5mC to 5fC and 5caC in DNA19,20,21. Although hm6A and f6A are inherently unstable, it is possible that additional protein factors could be involved to promote the FTO-mediated demethylation of m6A by catalysing the hydrolysis of hm6A and f6A; on the other hand, hm6A and f6A can also be stabilized when buried in a hydrophobic environment of their potential binding proteins. Further studies to explore the potential functions of these two modifications are required in the future to address these hypotheses.
Instead of direct demethylation, a delayed model is proposed for FTO, which possesses different kinetic behaviour to accommodate additional diffusion/export process and avoid direct remethylation. The nascent transcribed RNA is partially methylated by methyltransferase to m6A. m6A is converted to hm6A in regions with low FTO levels, and to f6A in regions with high FTO levels. Demethylated A is produced after the decomposition of hm6A and f6A, which is slower than the diffusion/export process, and which may help to avoid the remethylation by methyltransferase at the nuclear speckles. When RNA is processed and exported to the cytoplasm, different modifications on mRNA can potentially recruit or repel different protein factors, which could affect the metabolisms of mRNA. hm6A and f6A in RNA will eventually be hydrolysed to A; these modifications may also serve as markers for nascent RNA.
In summary, the identification of hm6A and f6A in cellular mRNA shows that additional modifications can be installed onto the N6-position of adenosine, which impact protein–RNA interaction to perhaps provide further dynamical tuning of the function/status of mRNA. Therefore, the discovery of these additional modifications on mRNA presents many intriguing questions and new biology to be explored.
Methods
General materials and methods
Complementary DNA clone of full-length human FTO and YTHDF2 was purchased from Open Biosystems. All primers were purchased from Eurofins MWG Operon. All RNase-free solutions were prepared from DEPC-treated MilliQ-water.
Synthesis and characterization of synthetic f6A standard
Detailed description of the synthesis of f6A standard can be found in the Supplementary Methods. The f6A standard was characterized by HPLC-QQQ-MS/MS, high-resolution mass spectra, UV–vis spectra, and both 1H and 13C NMR (Supplementary Figs S2–S4). Characterization data of f6A: 1H NMR (DMSO-d6, 500 MHz) δ 11.395 (br, 1H), 9.953 (s, 1H), 8.766 (s, 1H), 8.617 (s, 1H), 6.034 (d, J=5.5 Hz, 1H), 5.574 (d, J=6.0 Hz, 1H), 5.278 (d, J=5.0 Hz, 1H), 5.185 (t, J=5.5 Hz, 1H), 4.625 (dd, J=5.5 Hz, 5.5 Hz, 1H), 4.202 (dd, J=5.0 Hz, 4.0 Hz, 1H), 4.001 (dd, J=3.5 Hz, 4.0 Hz, 1H) 3.713 (m, 1H), 3.607 (m, 1H) p.p.m.; 13C NMR (DMSO-d6, 125 MHz) 165.193, 152.995, 152.649, 150.665, 144.237, 122.245, 88.690, 86.697, 74.769, 71.239, 62.204 p.p.m.. UV/Vis: λmax 274 nm, ε(260 nm)=1.30 × 104 M−1 cm−1, εmax(274 nm)=1.83 × 104 M−1 cm−1, LC-QQQ-MS/MS (Positive): [M+H]+=296.1, [Base+H]+=164.0, high-resolution MS (m/z) for C11H13N5O5: [M+H]+=296.0995 (calculated), 296.1009 (found).
Synthesis of m6A-RNA oligo
m6A phosphoramidite (N6-methyl-5′-O-(4,4′-dimethoxytrityl)-2′-O-t-butyldimethylsilyl adenosine 3′-O-(2-cyanoethyl-N,N-diisopropyl)phosphoramidite) was synthesized according to a previously reported procedure35. RNA oligo containing m6A was synthesized using m6A RNA phosphoramidite and standard RNA phosphoramidite (Glen Research) under ultra-mild conditions using 1H-tetrazole as activator reagent in an Expedite Nucleic Acid Synthesis System (PerSeptive Biosystems) with DMT-ON protocol. The oligo was cleaved from the beads and purified in a Glen-Pak RNA purification cartridge (Glen Research) according to the standard protocol provided by the manufacturer. The quality and purity of synthesized RNA was monitored by high-resolution MALDI-TOF-MS. Synthesized RNA sequence: RNA oligo with internal m6A: CUGGm6ACUGG, RNA oligo with 5′ m6A: m6ACUGACUAG.
Biochemistry assay of FTO activity in vitro
N-terminal his-tagged truncated human FTO protein (his6-FTO-NΔ31) was expressed and purified as previously reported27. The demethylation activity assay was performed in 20–100 μl of reaction mixture containing RNA with m6A, FTO, 75 μM of (NH4)2Fe(SO4)2·6H2O, 300 μM of α-KG, 2 mM of L-ascorbic acid, 150 mM of KCl and 50 mM of HEPES buffer (pH 7.0). The reaction was incubated at room temperature or 37 °C, and quenched by the addition of 1 mM of EDTA. The reaction was then frozen in liquid N2 immediately, and thawed only before analysis was performed. For HPLC or HPLC-MS/MS analysis, the reaction mixture was digested by 1 U of nuclease P1 at 37 °C for 15 min. HPLC analysis was performed on a HPLC system equipped with an Agilent Eclipse XDB-C18 analysis column (150 × 4.6 mm) eluted with buffer A (50 mM ammonium acetate in H2O) and buffer B (50 mM ammonium acetate in 60% of acetonitrile in H2O) with a flow rate of 1 ml min−1 at room temperature. The detection wavelength was set at 260 nm and 280 nm. HPLC-MS/MS was carried out by reverse-phase ultra-performance liquid chromatography on an Agilent ZORBAX Eclipse XDB-C18 column (Rapid Resolution HT, 50 × 2.1 mm) eluted with buffer A (0.1% formic acid in H2O) and buffer B (0.1% formic acid in methanol) with a flow rate of 0.5 ml min−1 at 35 °C, with online mass spectrometry detection using Agilent 6410 triple-quadrupole (QQQ) LC mass spectrometer in multiple reaction monitoring–positive electrospray ionization mode. The nucleosides were quantified using the nucleoside to base ion mass transitions of 282.1 to 150.1 (m6A), 268.0 to 136.0 (A), 296.1 to 164.1 (f6A) and 198.1 to 136.1 (hm6A). Quantification was performed by comparison with the standard curve obtained from synthetic nucleoside standards running at the same batch of samples.
MALDI-TOF-MS analysis of RNA oligo
The RNA/FTO reaction solution was first desalted by mixing with 50 μl of ammonium-charged AG 50 W-X8 Cation Exchange Resins (Bio-Rad). One microlitre of the desalted solution was then mixed with an equal amount of MALDI matrix, which was composed by 9:1 (v:v) ratio of 2′,4′,6′-trihydroxyacetophenone (THAP, 10 mg ml−1 in 50% CH3CN/H2O):diammonium citrate (50 mg ml−1 in H2O). The mixture was then spotted on a MALDI sample plate, dried under vacuum and analysed by a Bruker Ultraflextreme MALDI-TOF-TOF Mass Spectrometers in reflector, positive mode.
Analysis of hm6A and f6A in mRNA using HPLC-MS/MS
Total RNA was isolated from cultured cell or mouse tissues using TRIzol reagent according to the manufacture procedure. mRNA was then isolated with biotin-oligo(dT) and streptavidin beads from Promega, following a modified procedure, which eliminated all heating steps. Eight micrograms of mRNA was digested with RNase T1 in 20 μl for 15 min at 37 °C, followed by nuclease P1 (1 U) for 30 min at 37 °C. The solution was diluted five times, and 20 μl of the solution was analysed by HPLC-QQQ-MS/MS. The nucleosides were quantified using the nucleoside to base ion mass transitions of 282.1 to 150.1 (m6A), 268.0 to 136.0 (A), 296.1 to 164.1 (f6A) and 198.1 to 136.1 (hm6A) in multiple reaction monitoring–positive electrospray ionization mode.
MD and alchemical free-energy simulations
The initial coordinates were taken from the crystal structure of FTO bound with Fe2+, N-oxalylglycine and m3T (PDB ID: 3LFM). We replaced the N-oxalylglycine (antagonist of FTO) with the active cofactor α-KG. The coordinates of the modified bases were generated in two different ways, and two independent sets of simulations were carried out accordingly. The first approach was to fix the sugar ring position of m3T in 3LFM, and then mutate the original m3T to m3C, m6A or hm6A, respectively. These systems were labelled as FTO-m3T-cry, FTO-m3C-cry, FTO-m6A-cry and FTO-hm6A-cry, respectively. The second protocol was to align the AlkB–DNA complex crystal structures with FTO based on three-dimensional protein structure similarity calculated with the Protein Comparison Tool and then take the coordinates of the modified bases from the aligned AlkB complexes. We used PDB file 3O1O to derive the coordinates for m3T, 3O1M for m3C and 3O1P for εA; εA was then mutated to m6A or hm6A. These systems were labelled as FTO-m3T-alkb, FTO-m3C-alkb, FTO-m6A-alkb and FTO-hm6A-alkb, respectively. Detailed procedure for MD simulation set-up and alchemical free-energy simulations for relative binding affinities are provided in Supplementary Methods.
Additional information
How to cite this article: Fu, Y. et al. FTO-Mediated Formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nat. Commun. 4:1798 doi: 10.1038/ncomms2822 (2013).
References
Chan, C. T. et al. A quantitative systems approach reveals dynamic control of tRNA modifications during cellular stress. PLoS Genet 6, e1001247 (2010).
Jia, G. et al. N6-Methyladenosine in nuclear RNA is a major substrate of the obesity-associated FTO. Nat. Chem. Biol. 7, 885–887 (2011).
Perry, R. P., Kelley, D. E., Friderici, K. & Rottman, F. The methylated constituents of L cell messenger RNA: evidence for an unusual cluster at the 5′ terminus. Cell 4, 387–394 (1975).
Dominissini, D. et al. Topology of the human and mouse m6A RNA methylomes revealed by m6A-seq. Nature 485, 201–206 (2012).
Meyer, K. D. et al. Comprehensive analysis of mRNA methylation reveals enrichment in 3' UTRs and near stop codons. Cell 149, 1635–1646 (2012).
Bodi, Z., Button, J. D., Grierson, D. & Fray, R. G. Yeast targets for mRNA methylation. Nucleic Acids. Res. 38, 5327–5335 (2010).
Hongay, C. F. & Orr-Weaver, T. L. Drosophila inducer of meiosis 4 (IME4) is required for Notch signaling during oogenesis. Proc. Natl Acad. Sci. USA. 108, 14855–14860 (2011).
Bokar, J. A., Shambaugh, M. E., Polayes, D., Matera, A. G. & Rottman, F. M. Purification and cDNA cloning of the AdoMet-binding subunit of the human mRNA (N6-adenosine)-methyltransferase. RNA 3, 1233–1247 (1997).
Agarwala, S. D., Blitzblau, H. G., Hochwagen, A. & Fink, G. R. RNA methylation by the MIS complex regulates a cell fate decision in yeast. PLoS Genet. 8, e1002732 (2012).
Dina, C. et al. Variation in FTO contributes to childhood obesity and severe adult obesity. Nat. Genet. 39, 724–726 (2007).
Frayling, T. M. et al. A common variant in the FTO gene is associated with body mass index and predisposes to childhood and adult obesity. Science 316, 889–894 (2007).
Scuteri, A. et al. Genome-wide association scan shows genetic variants in the FTO gene are associated with obesity-related traits. PLoS Genet. 3, e115 (2007).
Falnes, P. O., Johansen, R. F. & Seeberg, E. AlkB-mediated oxidative demethylation reverses DNA damage in Escherichia coli. Nature 419, 178–182 (2002).
Trewick, S. C., Henshaw, T. F., Hausinger, R. P., Lindahl, T. & Sedgwick, B. Oxidative demethylation by Escherichia coli AlkB directly reverts DNA base damage. Nature 419, 174–178 (2002).
Gerken, T. et al. The obesity-associated FTO gene encodes a 2-oxoglutarate-dependent nucleic acid demethylase. Science 318, 1469–1472 (2007).
Loenarz, C. & Schofield, C. J. Physiological and biochemical aspects of hydroxylations and demethylations catalyzed by human 2-oxoglutarate oxygenases. Trends Biochem. Sci. 36, 7–18 (2011).
Kriaucionis, S. & Heintz, N. The nuclear DNA base 5-hydroxymethylcytosine is present in Purkinje neurons and the brain. Science 324, 929–930 (2009).
Tahiliani, M. et al. Conversion of 5-methylcytosine to 5-hydroxymethylcytosine in mammalian DNA by MLL partner TET1. Science 324, 930–935 (2009).
He, Y. F. et al. Tet-mediated formation of 5-carboxylcytosine and its excision by TDG in mammalian DNA. Science 333, 1303–1307 (2011).
Ito, S. et al. Tet proteins can convert 5-methylcytosine to 5-formylcytosine and 5-carboxylcytosine. Science 333, 1300–1303 (2011).
Pfaffeneder, T. et al. The discovery of 5-formylcytosine in embryonic stem cell DNA. Angew. Chem. Int. Ed. Engl. 50, 7008–7012 (2011).
Munzel, M. et al. Quantification of the sixth DNA base hydroxymethylcytosine in the brain. Angew. Chem. Int. Ed. Engl. 49, 5375–5377 (2010).
Zhang, L. et al. Thymine DNA glycosylase specifically recognizes 5-carboxylcytosine-modified DNA. Nat. Chem. Biol. 8, 328–330 (2012).
Alderson, T. Mechanism of formaldehyde-induced mutagenesis. The uniqueness of adenylic acid in the mediation of the mutagenic activity of formaldehyde. Nature 187, 485–489 (1960).
Ohkubo, A. et al. Oligonucleotide synthesis involving deprotection of amidine-type protecting groups for nucleobases under acidic conditions. Org. Lett. 12, 2496–2499 (2010).
Ross, J. mRNA stability in mammalian cells. Microbiol. Rev. 59, 423–450 (1995).
Han, Z. et al. Crystal structure of the FTO protein reveals basis for its substrate specificity. Nature 464, 1205–1209 (2010).
Prlic, A. et al. Pre-calculated protein structure alignments at the RCSB PDB website. Bioinformatics 26, 2983–2985 (2010).
Yi, C. et al. Iron-catalysed oxidation intermediates captured in a DNA repair dioxygenase. Nature 468, 330–333 (2010).
Straatsma, T. P. & Mccammon, J. A. Multiconfiguration thermodynamic integration. J. Chem. Phys. 95, 1175–1188 (1991).
Hashimoto, H. et al. Recognition and potential mechanisms for replication and erasure of cytosine hydroxymethylation. Nucleic Acids. Res. 40, 4841–4849 (2012).
Li, D. et al. Exocyclic carbons adjacent to the N(6) of adenine are targets for oxidation by the Escherichia coli adaptive response protein AlkB. J. Am. Chem. Soc. 134, 8896–8901 (2012).
Dumelin, C. E., Chen, Y., Leconte, A. M., Chen, Y. G. & Liu, D. R. Discovery and biological characterization of geranylated RNA in bacteria. Nat. Chem. Biol. 8, 913–919 (2012).
Zheng, G. et al. ALKBH5 is a mammalian RNA demethylase that impacts RNA metabolism and mouse fertility. Mol. Cell 49, 18–29 (2013).
Dai, Q. et al. Identification of recognition residues for ligation-based detection and quantitation of pseudouridine and N6-methyladenosine. Nucleic Acids Res. 35, 6322–6329 (2007).
Acknowledgements
We thank Dr Q. Jin for help with MALDI-TOF-MS and HPLC-QQQ-MS/MS, and G. Hou for discussions of technical issues on MD simulations. This work is supported by National Institute of Health (GM071440 to C.H., and GM084028 to Q.C.) and by a NIH EUREKA award (GM088599 to C.H.). The Mass Spectrometry Facility of the University of Chicago is funded by National Science Foundation (CHE-1048528). Y.F. is supported by Weil Endowed Fellowship. Y.F. and X.P. acknowledges support from the China Scholarship Council. We thank S. F. Reichard, MA, for editing the manuscript.
Author information
Authors and Affiliations
Contributions
Y.F., G.J. and C.H. designed and performed the experiments with the help of R.W., X.W., C.L.; and Q.D., X.P., K.H. and Q.C. designed and performed the molecular dynamics simulations. K.B., S.S. and M.N. provided mouse tissue samples. Y.F., Q.C. and C.H. wrote the paper. All authors discussed the results and commented on the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Information
Supplementary Figures S1-S25, Supplementary Table S1, Supplementary Methods and Supplementary References (PDF 17273 kb)
Rights and permissions
About this article
Cite this article
Fu, Y., Jia, G., Pang, X. et al. FTO-mediated formation of N6-hydroxymethyladenosine and N6-formyladenosine in mammalian RNA. Nat Commun 4, 1798 (2013). https://doi.org/10.1038/ncomms2822
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/ncomms2822
This article is cited by
-
The emerging roles of N6-methyladenosine RNA modifications in thyroid cancer
European Journal of Medical Research (2023)
-
dTrmt10A impacts Hsp70 chaperone m6A levels and the stress response in the Drosophila brain
Scientific Reports (2023)
-
The interplay between N6-methyladenosine and precancerous liver disease: molecular functions and mechanisms
Discover Oncology (2023)
-
N6-methyladenosine RNA methylation in liver diseases: from mechanism to treatment
Journal of Gastroenterology (2023)
-
The role of N6-methyladenosine methylation in PAHs-induced cancers
Environmental Science and Pollution Research (2023)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.