Featured
-
-
Article
| Open Access A meet-up of two second messengers: the c-di-AMP receptor DarB controls (p)ppGpp synthesis in Bacillus subtilis
A meet-up of two second messengers: the c-di-AMP receptor DarB controls (p)ppGpp synthesis in Bacillus subtilis
In several bacteria, cyclic di-AMP mediates potassium (K+) and osmotic homeostasis. Here, the authors show that DarB, a Bacillus subtilis protein previously reported to bind cyclic di-AMP, interacts with the (p)ppGpp synthetase/hydrolase Rel in a K+-dependent manner in turn leading to Rel-dependent accumulation of pppGpp under conditions of K+ starvation.
- Larissa Krüger
- , Christina Herzberg
- & Jörg Stülke
-
Article
| Open Access Discovery of an ene-reductase for initiating flavone and flavonol catabolism in gut bacteria
Discovery of an ene-reductase for initiating flavone and flavonol catabolism in gut bacteria
Flavonoids are abundant polyphenols in plants but it is not well understood how their metabolism is initiated by microbes in the human gut. Here, the authors identify and characterise an ene-reductase from the gut bacterium, Flavonifractor plautii ATCC 49531 that catalyses the hydrogenation of the C2–C3 double bond of flavones and flavonols and present its crystal structure.
- Gaohua Yang
- , Sen Hong
- & Yang Gu
-
Article
| Open Access Uncovering de novo gene birth in yeast using deep transcriptomics
Uncovering de novo gene birth in yeast using deep transcriptomics
Genome-wide studies of de novo genes have tended to focus on genomic open reading frames (ORFs). Here, Blevins et al. use deep transcriptomics and synteny information to identify de novo transcripts in the yeast Saccharomyces cerevisiae, many of which are expressed from the alternative DNA strand.
- William R. Blevins
- , Jorge Ruiz-Orera
- & M. Mar Albà
-
Article
| Open Access Multiplexed characterization of rationally designed promoter architectures deconstructs combinatorial logic for IPTG-inducible systems
Multiplexed characterization of rationally designed promoter architectures deconstructs combinatorial logic for IPTG-inducible systems
Precisely tuning the genetic response to environmental stimuli is a key step in engineering synthetic biology systems. Here, the authors profile 8269 IPTG-induced promoters to deconstruct the relationship between sequence architecture and gene expression.
- Timothy C. Yu
- , Winnie L. Liu
- & Guillaume Urtecho
-
Article
| Open Access Varicella-zoster virus VLT-ORF63 fusion transcript induces broad viral gene expression during reactivation from neuronal latency
Varicella-zoster virus VLT-ORF63 fusion transcript induces broad viral gene expression during reactivation from neuronal latency
Varicella-zoster virus (VZV) establishes lifelong neuronal latency in humans. Here, Ouwendijk and Depledge et al. identify a fusion transcript, VLT-ORF63, which is expressed during lytic and latent infection, and demonstrate a role for the translated fusion protein in induction of lytic gene expression from latent VZV genomes.
- Werner J. D. Ouwendijk
- , Daniel P. Depledge
- & Tomohiko Sadaoka
-
Article
| Open Access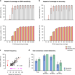 Analytical validity of nanopore sequencing for rapid SARS-CoV-2 genome analysis
Analytical validity of nanopore sequencing for rapid SARS-CoV-2 genome analysis
Nanopore sequencing (ONT) has been used in SARS-CoV-2 studies, however adoption of ONT for SARS-CoV-2 surveillance has been limited due to common concerns around sequencing accuracy. Here, the authors perform a comprehensive evaluation of ONT analytical performance on 157 matched SARS-CoV-2-positive patient specimens and synthetic RNA controls.
- Rowena A. Bull
- , Thiruni N. Adikari
- & Ira W. Deveson
-
Article
| Open Access Pandemic Vibrio cholerae shuts down site-specific recombination to retain an interbacterial defence mechanism
Pandemic Vibrio cholerae shuts down site-specific recombination to retain an interbacterial defence mechanism
Vibrio cholerae uses a type VI secretion system (T6SS) to kill neighbouring competitors. Here, Santoriello et al. show that a T6SS gene cluster (Aux3) exists as a mobile, prophage-like element in some environmental strains, and as a stable truncated form in pandemic isolates. They propose that Aux3 acquisition increased competitive fitness of pre-pandemic V. cholerae.
- Francis J. Santoriello
- , Lina Michel
- & Stefan Pukatzki
-
Article
| Open Access No evidence for increased transmissibility from recurrent mutations in SARS-CoV-2
No evidence for increased transmissibility from recurrent mutations in SARS-CoV-2
SARS-CoV-2 has emerged recently and may still adapt to the human host. Here the authors show that none of the so far identified recurrent mutations in SARS-CoV-2 are significantly associated with increased viral transmission.
- Lucy van Dorp
- , Damien Richard
- & François Balloux
-
Article
| Open Access Horizontally acquired papGII-containing pathogenicity islands underlie the emergence of invasive uropathogenic Escherichia coli lineages
Horizontally acquired papGII-containing pathogenicity islands underlie the emergence of invasive uropathogenic Escherichia coli lineages
Escherichia coli is a major cause of urinary tract infection. Here, Biggel et al. provide a phylogenomic analysis of 907 clinical E. coli isolates and identify the P-fimbriae-encoding locus associated with invasive uropathogenic E. coli isolates.
- Michael Biggel
- , Basil B. Xavier
- & Sandra Van Puyvelde
-
Article
| Open Access High-throughput laboratory evolution reveals evolutionary constraints in Escherichia coli
High-throughput laboratory evolution reveals evolutionary constraints in Escherichia coli
Understanding evolutionary constraints in antibiotic resistance is crucial for prediction and control. Here, the authors use high-throughput laboratory evolution of Escherichia coli alongside machine learning to identify trade-off relationships associated with drug resistance.
- Tomoya Maeda
- , Junichiro Iwasawa
- & Chikara Furusawa
-
Article
| Open Access The seventh pandemic of cholera in Europe revisited by microbial genomics
The seventh pandemic of cholera in Europe revisited by microbial genomics
Since 1970, several cholera outbreaks caused by the “seventh pandemic” (7PET) lineage have been reported in Europe. Here, the authors demonstrate that the outbreaks were caused by repeated introductions of 7PET into Europe, rather than local environmental sources.
- Mihaela Oprea
- , Elisabeth Njamkepo
- & François-Xavier Weill
-
Article
| Open Access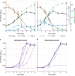 The reductive glycine pathway allows autotrophic growth of Desulfovibrio desulfuricans
The reductive glycine pathway allows autotrophic growth of Desulfovibrio desulfuricans
There are several pathways for CO2 fixation in photoautotrophic and chemoautotrophic microorganisms. Here, the authors provide experimental demonstration for the operation of the reductive glycine pathway in a natural microorganism, Desulfovibrio desulfuricans.
- Irene Sánchez-Andrea
- , Iame Alves Guedes
- & Alfons J. M. Stams
-
Article
| Open Access TssA–TssM–TagA interaction modulates type VI secretion system sheath-tube assembly in Vibrio cholerae
TssA–TssM–TagA interaction modulates type VI secretion system sheath-tube assembly in Vibrio cholerae
The molecular mechanisms by which the sheath-tube of the bacterial type VI secretion system assembles are not fully understood. Here, the authors present evidence that the priming of the sheath-tube is controlled by TssA–TssM–TagA in Vibrio cholerae.
- Maria Silvina Stietz
- , Xiaoye Liang
- & Tao G. Dong
-
Article
| Open Access Piperacillin/tazobactam resistance in a clinical isolate of Escherichia coli due to IS26-mediated amplification of blaTEM-1B
Piperacillin/tazobactam resistance in a clinical isolate of Escherichia coli due to IS26-mediated amplification of blaTEM-1B
An E. coli and K. pneumoniae phenotype resistant to piperacillin/tazobactam has recently emerged. Here, the authors show that hyperproduction of the β-lactamase driving this resistance occurs due to excision and reinsertion of a translocatable unit containing blaTEM-1B, creating a tandem array.
- Alasdair T. M. Hubbard
- , Jenifer Mason
- & Thomas Edwards
-
Article
| Open Access Genomics of the Argentinian cholera epidemic elucidate the contrasting dynamics of epidemic and endemic Vibrio cholerae
Genomics of the Argentinian cholera epidemic elucidate the contrasting dynamics of epidemic and endemic Vibrio cholerae
Pandemic cholera was reintroduced to Argentina in 1992, leading to epidemic spread. Here, the authors use whole genome sequencing to show how, over 6 years, epidemic cholera was caused by invariant 7PET lineage Vibrio cholerae, against a background of sporadic disease caused by diverse local strains.
- Matthew J. Dorman
- , Daryl Domman
- & Nicholas R. Thomson
-
Article
| Open Access A distinct lineage of Caudovirales that encodes a deeply branching multi-subunit RNA polymerase
A distinct lineage of Caudovirales that encodes a deeply branching multi-subunit RNA polymerase
Viruses have been difficult to position in the Tree of Life using phylogenetic methods. This study uses an ancient enzyme multi-subunit RNA polymerase (RNAP) to reveal a novel viral group, the Caudovirales, and to suggest an ancient origin of RNAP in this group.
- Alaina R. Weinheimer
- & Frank O. Aylward
-
Article
| Open Access Antibiotic susceptibility signatures identify potential antimicrobial targets in the Acinetobacter baumannii cell envelope
Antibiotic susceptibility signatures identify potential antimicrobial targets in the Acinetobacter baumannii cell envelope
A unique cell envelope contributes to the antibiotic resistance of the pathogen Acinetobacter baumannii. Here, Geisinger et al. identify A. baumannii mutants with altered antibiotic susceptibility, infer the function of uncharacterized proteins involved in envelope synthesis, and predict antibiotic synergies.
- Edward Geisinger
- , Nadav J. Mortman
- & Ralph R. Isberg
-
Article
| Open Access Tracking the COVID-19 pandemic in Australia using genomics
Tracking the COVID-19 pandemic in Australia using genomics
Genome sequencing can be used to infer pathogen transmission dynamics and inform public health responses. Here, the authors sequence >1,200 SARS-CoV-2 samples from Victoria, Australia and find genomic support for the effectiveness of social restrictions in reducing transmission.
- Torsten Seemann
- , Courtney R. Lane
- & Benjamin P. Howden
-
Article
| Open Access Widespread transfer of mobile antibiotic resistance genes within individual gut microbiomes revealed through bacterial Hi-C
Widespread transfer of mobile antibiotic resistance genes within individual gut microbiomes revealed through bacterial Hi-C
Linking antibiotic resistance (AR) in the gut microbiome with their bacterial hosts remains challenging. Here, the authors apply bacterial Hi-C to map mobile genetic elements in metagenomes, and illustrate that genes are present in more diverse taxa in neutropenic patients than healthy subjects.
- Alyssa G. Kent
- , Albert C. Vill
- & Ilana Lauren Brito
-
Article
| Open Access Selective flexible packaging pathways of the segmented genome of influenza A virus
Selective flexible packaging pathways of the segmented genome of influenza A virus
The mechanism underlying packaging of the 8 segments of the influenza virus genome into virions is not well understood. Here, the authors use a multiplexed FISH assay to monitor the 8 segments in parallel in infected cells suggesting bundling routes during the packaging process.
- Ivan Haralampiev
- , Simon Prisner
- & Andreas Herrmann
-
Article
| Open Access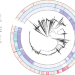 Adaptation to the cervical environment is associated with increased antibiotic susceptibility in Neisseria gonorrhoeae
Adaptation to the cervical environment is associated with increased antibiotic susceptibility in Neisseria gonorrhoeae
Antibiotic resistance in Neisseria gonorrhoeae is rising, yet sometimes strains emerge that have reverted to susceptibility. Here, the authors find that selective pressures from the host may influence susceptibility through loss-of-function mutations in genes that encode for efflux pumps.
- Kevin C. Ma
- , Tatum D. Mortimer
- & Yonatan H. Grad
-
Article
| Open Access Efflux pump activity potentiates the evolution of antibiotic resistance across S. aureus isolates
Efflux pump activity potentiates the evolution of antibiotic resistance across S. aureus isolates
Some bacterial lineages appear to be pre-disposed to evolving antibiotic resistance. Here, the authors show that differential expression of an efflux pump causes widespread variation in evolvability across Staphylococcus aureus isolates, and chemical inhibition of the pump prevents resistance evolution.
- Andrei Papkou
- , Jessica Hedge
- & R. Craig MacLean
-
Article
| Open Access Pathways for horizontal gene transfer in bacteria revealed by a global map of their plasmids
Pathways for horizontal gene transfer in bacteria revealed by a global map of their plasmids
Plasmids can mediate gene transfer across bacterial populations. Here, the authors describe a global map of the prokaryotic plasmidome, where plasmids organize into discrete ‘plasmid taxonomic units’ based on their genomic composition and pairwise sequence identity.
- Santiago Redondo-Salvo
- , Raúl Fernández-López
- & Fernando de la Cruz
-
Article
| Open Access “Gene accordions” cause genotypic and phenotypic heterogeneity in clonal populations of Staphylococcus aureus
“Gene accordions” cause genotypic and phenotypic heterogeneity in clonal populations of Staphylococcus aureus
Gene tandem amplifications can drive bacterial evolution. Here, Belikova et al. identify copy number variations of lipoprotein-encoding genes in Staphylococcus aureus clinical isolates, and show that the loci expand and contract during bacterial growth in vitro and in mice, leading to changes in immunostimulatory capacity.
- Darya Belikova
- , Angelika Jochim
- & Simon Heilbronner
-
Article
| Open Access Within-host microevolution of Streptococcus pneumoniae is rapid and adaptive during natural colonisation
Within-host microevolution of Streptococcus pneumoniae is rapid and adaptive during natural colonisation
Streptococcus pneumoniae is an opportunistic pathogen and asymptomatic colonization is a precursor for invasive disease. Here the authors show rapid within-host evolution of naturally acquired pneumococci in ninety-eight infants driven by high nucleotide substitution rates and intra-host homologous recombination.
- Chrispin Chaguza
- , Madikay Senghore
- & Brenda A. Kwambana-Adams
-
Article
| Open Access A biochemically-interpretable machine learning classifier for microbial GWAS
A biochemically-interpretable machine learning classifier for microbial GWAS
Current machine learning classifiers have been applied to whole-genome sequencing data to identify determinants of antimicrobial resistance, but they lack interpretability. Here the authors present a metabolic machine learning classifier that uses flux balance analysis to estimate the biochemical effects of alleles.
- Erol S. Kavvas
- , Laurence Yang
- & Bernhard O. Palsson
-
Article
| Open Access Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0
Precise phylogenetic analysis of microbial isolates and genomes from metagenomes using PhyloPhlAn 3.0
The increasing amount of sequenced microbial genomes and metagenomes requires platforms for efficient integrated analysis. Here, Asnicar et al. present PhyloPhlAn 3.0, a pipeline allowing large-scale microbial genome characterization and phylogenetic contextualization at multiple levels of resolution.
- Francesco Asnicar
- , Andrew Maltez Thomas
- & Nicola Segata
-
Article
| Open Access Sphingolipids produced by gut bacteria enter host metabolic pathways impacting ceramide levels
Sphingolipids produced by gut bacteria enter host metabolic pathways impacting ceramide levels
Ceramides are a type of sphingolipid (SL) that have been shown to play a role in several metabolic disorders. Here, the authors investigate the effect of SL-production by gut Bacteroides on host SL homeostasis and show that microbiome-derived SLs enter host circulation and alter ceramide production.
- Elizabeth L. Johnson
- , Stacey L. Heaver
- & Ruth E. Ley
-
Article
| Open Access The route to transcription initiation determines the mode of transcriptional bursting in E. coli
The route to transcription initiation determines the mode of transcriptional bursting in E. coli
Transcription noise in bacteria is often attributed to burstiness, but the mechanisms are unclear. Here, the authors show that the transition from low to high expression can be regulated via burst size or burst frequency, depending on the mode of transcription initiation determined by different sigma factors.
- Christoph Engl
- , Goran Jovanovic
- & Martin Buck
-
Article
| Open Access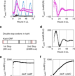 Damped circadian oscillation in the absence of KaiA in Synechococcus
Damped circadian oscillation in the absence of KaiA in Synechococcus
Proteins KaiA, KaiB and KaiC constitute a biochemical circadian oscillator in Synechococcus cyanobacteria. Here, Kawamoto et al. show that kaiBC promoter activity exhibits a damped, low-amplitude circadian oscillation in the absence of KaiA, which could explain the circadian rhythms observed in other bacteria that lack a kaiA homologue.
- Naohiro Kawamoto
- , Hiroshi Ito
- & Hideo Iwasaki
-
Article
| Open Access Stability and nuclear localization of yeast telomerase depend on protein components of RNase P/MRP
Stability and nuclear localization of yeast telomerase depend on protein components of RNase P/MRP
Pop1 and 6 are subunits of RNase P and RNase MRP, which process ribosomal and tRNAs. The authors show that when Pop1 and 6 are impaired, the telomerase subunit Est1 binds telomerase RNA at normal levels, but the binding is unstable. As a result, nuclear import of the telomerase holoenzyme is inhibited.
- P. Daniela Garcia
- , Robert W. Leach
- & Virginia A. Zakian
-
Article
| Open Access An acid-tolerance response system protecting exponentially growing Escherichia coli
An acid-tolerance response system protecting exponentially growing Escherichia coli
The ability to grow at acidic pH is crucial for E. coli colonization of the host’s intestine. Here, the authors identify an acid-tolerance response system that is important for E. coli exponential growth at pH 4.2, survival in the mouse intestine, and production of 3-hydroxypropionate during fermentation.
- Ying Xu
- , Zhe Zhao
- & Guang Zhao
-
Article
| Open Access Dominant resistance and negative epistasis can limit the co-selection of de novo resistance mutations and antibiotic resistance genes
Dominant resistance and negative epistasis can limit the co-selection of de novo resistance mutations and antibiotic resistance genes
The authors study the interactions between chromosomal mutations and horizontally acquired genes in the evolution of antibiotic resistance in experimental evolution assays. They identify constraints that may allow better prediction and control of antibiotic resistance evolution.
- Andreas Porse
- , Leonie J. Jahn
- & Morten O. A. Sommer
-
Article
| Open Access De novo emergence of adaptive membrane proteins from thymine-rich genomic sequences
De novo emergence of adaptive membrane proteins from thymine-rich genomic sequences
There is increasing evidence that protein-coding genes can emerge de novo from noncoding genomic regions. Vakirlis et al. propose that sequences encoding transmembrane polypeptides can emerge de novo in thymine-rich genomic regions and provide organisms with fitness benefits.
- Nikolaos Vakirlis
- , Omer Acar
- & Anne-Ruxandra Carvunis
-
Article
| Open Access Abundance and diversity of resistomes differ between healthy human oral cavities and gut
Abundance and diversity of resistomes differ between healthy human oral cavities and gut
Antimicrobial resistance (AMR) represents a global health threat. Here, the authors analyse the oral and gut resistomes from metagenomes of diverse populations and find that the oral resistome harbours higher abundance but lower diversity of antimicrobial resistance genes than the gut resistome.
- Victoria R. Carr
- , Elizabeth A. Witherden
- & David L. Moyes
-
Article
| Open Access Plasmid-mediated metronidazole resistance in Clostridioides difficile
Plasmid-mediated metronidazole resistance in Clostridioides difficile
Cases of C. difficile (CD) resistant to metronidazole have been reported but the mechanism remains enigmatic. Here the authors identify a plasmid, which correlates with metronidazole resistance status in a large international collection of CD isolates, and demonstrate that the plasmid can confer metronidazole resistance.
- Ilse M. Boekhoud
- , Bastian V. H. Hornung
- & Wiep Klaas Smits
-
Article
| Open Access Integrating multiple genomic technologies to investigate an outbreak of carbapenemase-producing Enterobacter hormaechei
Integrating multiple genomic technologies to investigate an outbreak of carbapenemase-producing Enterobacter hormaechei
Antibiotic-resistant bacteria are an urgent threat to human health. Here, Roberts et al. characterise and monitor an ongoing hospital outbreak of carbapenemase-producing Enterobacter hormaechei by integrating several technologies for whole-genome sequencing and shotgun metagenomics.
- Leah W. Roberts
- , Patrick N. A. Harris
- & Scott A. Beatson
-
Article
| Open Access Acquisition, transmission and strain diversity of human gut-colonizing crAss-like phages
Acquisition, transmission and strain diversity of human gut-colonizing crAss-like phages
CrAss-like phages are bacterial viruses often found in the human gut. Here, Siranosian et al. analyze gut metagenomic data to evaluate the patterns of acquisition, transmission and strain diversity of these phages in mother-infant pairs and in patients undergoing fecal microbiota transplantation.
- Benjamin A. Siranosian
- , Fiona B. Tamburini
- & Ami S. Bhatt
-
Article
| Open Access Recording mobile DNA in the gut microbiota using an Escherichia coli CRISPR-Cas spacer acquisition platform
Recording mobile DNA in the gut microbiota using an Escherichia coli CRISPR-Cas spacer acquisition platform
Horizontal gene transfer (HGT) in complex bacterial communities has been mostly studied using metagenomic analyses. Here, the authors develop an E. coli CRISPR-Cas spacer acquisition platform that allows real-time recording of HGT events at nucleotide-resolution, identifying diverse DNA transfer events in human clinical fecal samples.
- Christian Munck
- , Ravi U. Sheth
- & Harris H. Wang
-
Article
| Open Access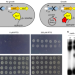 YtfK activates the stringent response by triggering the alarmone synthetase SpoT in Escherichia coli
YtfK activates the stringent response by triggering the alarmone synthetase SpoT in Escherichia coli
The enzyme SpoT is important for accumulation of the alarmone (p)ppGpp, which triggers the stringent response in E. coli. Here, Germain et al. show that the protein YtfK promotes SpoT-dependent accumulation of (p)ppGpp and is required for activation of the stringent response during phosphate and fatty acid starvation.
- Elsa Germain
- , Paul Guiraud
- & Etienne Maisonneuve
-
Article
| Open Access Droplet Tn-Seq combines microfluidics with Tn-Seq for identifying complex single-cell phenotypes
Droplet Tn-Seq combines microfluidics with Tn-Seq for identifying complex single-cell phenotypes
Culturing transposon-mutant libraries in pools can mask complex phenotypes. Here the authors present microfluidics mediated droplet Tn-Seq, which encapsulates individual mutants, promotes isolated growth and enables cell-cell interaction analyses.
- Derek Thibault
- , Paul A. Jensen
- & Tim van Opijnen
-
Article
| Open Access A bacterial gene-drive system efficiently edits and inactivates a high copy number antibiotic resistance locus
A bacterial gene-drive system efficiently edits and inactivates a high copy number antibiotic resistance locus
Genedrives bias the inheritance of alleles in diploid organisms. Here, the authors develop a gene-drive analogous system for bacteria, selectively editing and clearing plasmids.
- J. Andrés Valderrama
- , Surashree S. Kulkarni
- & Ethan Bier
-
Article
| Open Access Gene gain and loss push prokaryotes beyond the homologous recombination barrier and accelerate genome sequence divergence
Gene gain and loss push prokaryotes beyond the homologous recombination barrier and accelerate genome sequence divergence
A significant proportion of the molecular evolution of bacteria and archaea occurs through gene gain and loss. Here Iranzo et al. develop a mathematical model that explains observed differential patterns of sequence evolution vs. gene content evolution as a consequence of homologous recombination.
- Jaime Iranzo
- , Yuri I. Wolf
- & Itamar Sela
-
Article
| Open Access Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis
Evaluation of 16S rRNA gene sequencing for species and strain-level microbiome analysis
Here, the authors explore the potential of the 16S gene for discriminating bacterial taxa and show that full-length sequencing combined with appropriate clustering of intragenomic sequence variation can provide accurate representation of bacterial species in microbiome datasets.
- Jethro S. Johnson
- , Daniel J. Spakowicz
- & George M. Weinstock
-
Article
| Open Access Conditional quorum-sensing induction of a cyanide-insensitive terminal oxidase stabilizes cooperating populations of Pseudomonas aeruginosa
Conditional quorum-sensing induction of a cyanide-insensitive terminal oxidase stabilizes cooperating populations of Pseudomonas aeruginosa
Quorum sensing (QS) regulates production of ‘public goods’ by Pseudomonas aeruginosa, which releases toxic hydrogen cyanide to constrain QS-deficient cheaters. Here, Yan et al. show that QS-proficient strains protect themselves by producing a cyanide-insensitive enzyme in response to reactive oxygen species released by cheaters.
- Huicong Yan
- , Kyle L. Asfahl
- & Meizhen Wang
-
Article
| Open Access Plasticity of the Mycobacterium tuberculosis respiratory chain and its impact on tuberculosis drug development
Plasticity of the Mycobacterium tuberculosis respiratory chain and its impact on tuberculosis drug development
New tuberculosis therapies, targeting respiratory chain components of Mycobacterium tuberculosis, are under development. Here the authors show that, contrary to common belief, some of these components are not essential for pathogen viability and/or virulence in animal models of infection.
- Tiago Beites
- , Kathryn O’Brien
- & Dirk Schnappinger
-
Article
| Open Access Prophages and satellite prophages are widespread in Streptococcus and may play a role in pneumococcal pathogenesis
Prophages and satellite prophages are widespread in Streptococcus and may play a role in pneumococcal pathogenesis
Prophages are viral genomes integrated within bacterial genomes. Here, Rezaei Javan et al. identify nearly 800 prophages and satellite prophages in > 1300 Streptococcus genomes, and show that a satellite prophage is associated with virulence in a mouse model of pneumococcal infection.
- Reza Rezaei Javan
- , Elisa Ramos-Sevillano
- & Angela B. Brueggemann
-
Article
| Open Access Dissecting the molecular evolution of fluoroquinolone-resistant Shigella sonnei
Dissecting the molecular evolution of fluoroquinolone-resistant Shigella sonnei
Shigella sonnei is one of the main species causing shigellosis worldwide. Here the authors analyse nearly 400 S. sonnei genome sequences and carry out experimental evolution experiments to shed light into the evolutionary processes underlying the recent emergence of fluoroquinolone resistance in this pathogen.
- Hao Chung The
- , Christine Boinett
- & Stephen Baker
-
Article
| Open Access Insights into the ecological roles and evolution of methyl-coenzyme M reductase-containing hot spring Archaea
Insights into the ecological roles and evolution of methyl-coenzyme M reductase-containing hot spring Archaea
Methane metabolism by some lineages of Archaea contributes to the cycling of carbon on Earth. Here, the authors show high diversity of methyl-coenzyme M reductase (Mcr), a key enzyme associated with archaeal methane/alkane metabolism, in hot spring Archaea, and investigate their ecological roles and evolution.
- Zheng-Shuang Hua
- , Yu-Lin Wang
- & Wen-Jun Li

 Control of a programmed cell death pathway in Pseudomonas aeruginosa by an antiterminator
Control of a programmed cell death pathway in Pseudomonas aeruginosa by an antiterminator