Article
|
Open Access
Featured
-
-
Article
| Open Access Structure of human Cdc45 and implications for CMG helicase function
Structure of human Cdc45 and implications for CMG helicase function
The cell cycle division protein Cdc45 is required for genome duplication in eukaryotes. Here, the authors determine the crystal structure of human Cdc45 and combine it with functional data to improve our understanding of its role in DNA replication.
- Aline C. Simon
- , Vincenzo Sannino
- & Luca Pellegrini
-
Article
| Open Access The topography of mutational processes in breast cancer genomes
The topography of mutational processes in breast cancer genomes
Mutational signatures provide evidence of the mechanism of action of a given mutagen and are found in cancer cells. Here, using 560 breast cancer genomes, the authors demonstrate that mutational signatures are frequently associated with genomic architecture including nucleosome positioning and replication timing.
- Sandro Morganella
- , Ludmil B. Alexandrov
- & Serena Nik-Zainal
-
Article
| Open Access 3D replicon distributions arise from stochastic initiation and domino-like DNA replication progression
3D replicon distributions arise from stochastic initiation and domino-like DNA replication progression
DNA replication in higher eukaryotes is a complex process occurring in a complex genome environment. Here the authors present a model of DNA replication that incorporates random loop chromatin folding and domino-like fork progression reproducing the spatial and temporal characteristics of S-phase.
- D. Löb
- , N. Lengert
- & B. Drossel
-
Article
| Open Access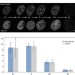 4D Visualization of replication foci in mammalian cells corresponding to individual replicons
4D Visualization of replication foci in mammalian cells corresponding to individual replicons
Whether replication happens at individual replicons or in replication factories is a controversial debate. Here the authors use super-resolution microscopy and analysis of replication fork speed to present evidence in favour of replicons.
- V. O. Chagin
- , C. S. Casas-Delucchi
- & M. C. Cardoso
-
Article
| Open Access The presence of extra chromosomes leads to genomic instability
The presence of extra chromosomes leads to genomic instability
One of the hallmarks of cancer cells is aneuploidy, however the molecular effects are poorly understood. Here the authors show that trisomic and tetrasomic cells display increased genomic instability and reduced levels of the helicase MCM2-7.
- Verena Passerini
- , Efrat Ozeri-Galai
- & Zuzana Storchová
-
Article
| Open Access Chromatin-associated degradation is defined by UBXN-3/FAF1 to safeguard DNA replication fork progression
Chromatin-associated degradation is defined by UBXN-3/FAF1 to safeguard DNA replication fork progression
Cdc48/p97 is a key component of the ubiquitin-proteasome system, acting as a ubiquitin-directed segregase to regulate multiple cellular functions. Here the authors identify UBXN-3/FAF1 as a crucial regulator of chromatin-associated protein degradation that recruits Cdc48/p97 to DNA replication forks.
- André Franz
- , Paul A. Pirson
- & Thorsten Hoppe
-
Article
| Open Access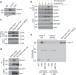 SCFCyclin F-dependent degradation of CDC6 suppresses DNA re-replication
SCFCyclin F-dependent degradation of CDC6 suppresses DNA re-replication
To ensure genome stability, cells need to restrict DNA replication to once per cell cycle. Here the authors show that Cyclin F interacts with and targets the licensing factor CDC6 for degradation, preventing re-firing of replication origins.
- David Walter
- , Saskia Hoffmann
- & Claus Storgaard Sørensen
-
Article
| Open Access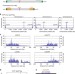 Shugoshin forms a specialized chromatin domain at subtelomeres that regulates transcription and replication timing
Shugoshin forms a specialized chromatin domain at subtelomeres that regulates transcription and replication timing
A chromosome is composed of structurally and functionally distinct domains. Here, Tashiro et al. report that the conserved centromeric protein Sgo2 localizes at the subtelomeres preferentially during G2phase and is essential for the formation of a highly condensed subtelomeric chromatin body “knob”.
- Sanki Tashiro
- , Tetsuya Handa
- & Junko Kanoh
-
Article
| Open Access Replication landscape of the human genome
Replication landscape of the human genome
The physical origin and termination sites of DNA replication in human cells have remained elusive. Here the authors use Okazaki fragment sequencing to reveal global replication patterns and show how chromatin and transcription modulate the process.
- Nataliya Petryk
- , Malik Kahli
- & Olivier Hyrien
-
Article
| Open Access T7 replisome directly overcomes DNA damage
T7 replisome directly overcomes DNA damage
Genomic instability can result from stalled or collapsed replication fork at sites of unrepaired DNA lesions. Here the authors uncover a new lesion bypass pathway for the T7 replisome, where leading strand template lesions can be overcome through interaction between the replisome's helicase and polymerase components.
- Bo Sun
- , Manjula Pandey
- & Michelle D. Wang
-
Article
| Open Access Mutagenic consequences of a single G-quadruplex demonstrate mitotic inheritance of DNA replication fork barriers
Mutagenic consequences of a single G-quadruplex demonstrate mitotic inheritance of DNA replication fork barriers
Barriers to DNA replication are potent sources of genome instability. Here, the authors provide a mechanistic model for how a single persistent G-quadruplex structure generates multiple substrates for polymerase theta-mediated end-joining, thus causing multiple deletions during animal development.
- Bennie Lemmens
- , Robin van Schendel
- & Marcel Tijsterman
-
Article
| Open Access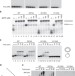 Complementation between polymerase- and exonuclease-deficient mitochondrial DNA polymerase mutants in genomically engineered flies
Complementation between polymerase- and exonuclease-deficient mitochondrial DNA polymerase mutants in genomically engineered flies
A key source of mitochondrial DNA mutations is errors introduced during genome replication. Here the authors create Drosophiliastrains with separated elongation and proofreading capabilities to explore the dynamism of mitochondrial DNA replication.
- Ana Bratic
- , Timo E. S. Kauppila
- & Nils-Göran Larsson
-
Article
| Open Access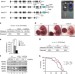 Replication stress caused by low MCM expression limits fetal erythropoiesis and hematopoietic stem cell functionality
Replication stress caused by low MCM expression limits fetal erythropoiesis and hematopoietic stem cell functionality
What causes hematopoietic stem cell loss of functionality? Here, Alvarez et al. show that loss of origin licensing factor MCM3 induces replicative stress (RS), causing aberrant erythrocyte maturation, but mice strains with higher tolerance to RS can overcome this defect.
- Silvia Alvarez
- , Marcos Díaz
- & Juan Méndez
-
Article
| Open Access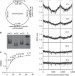 Activation of a dormant replication origin is essential for Haloferax mediterranei lacking the primary origins
Activation of a dormant replication origin is essential for Haloferax mediterranei lacking the primary origins
Archaea use multiple origins for chromosome replication, but some potential origins appear to be inactive during the genome replication. Here, Yang et al. show that when active origins are deleted from Haloferax mediterranei, a dormant origin is activated and essential, suggesting an origin-dependent replication.
- Haibo Yang
- , Zhenfang Wu
- & Hua Xiang
-
Article |
 PTEN regulates DNA replication progression and stalled fork recovery
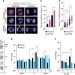
PTEN regulates DNA replication progression and stalled fork recovery
PTEN plays multiple roles in genome protection and tumour suppression. Here the authors show that PTEN depletion leads to impairment of replication progression, stalled fork recovery and diminished chromatin loading of Rad51, highlighting the interplay of PTEN with Rad51 in promoting stalled fork restart.
- Jinxue He
- , Xi Kang
- & Wen H. Shen
-
Article
| Open Access The exonuclease activity of DNA polymerase γ is required for ligation during mitochondrial DNA replication
The exonuclease activity of DNA polymerase γ is required for ligation during mitochondrial DNA replication
Mitochondrial DNA (mtDNA) polymerase γ has a 3′–5′ exonuclease proofreading activity. Here, the authors show it is required for creating ligatable ends during mtDNA replication, and inactivation of the activity in mice causes strand-specific nicks in DNA and the formation of linear mtDNA fragments.
- Bertil Macao
- , Jay P. Uhler
- & Maria Falkenberg
-
Article
| Open Access Polymerase Θ is a key driver of genome evolution and of CRISPR/Cas9-mediated mutagenesis
Polymerase Θ is a key driver of genome evolution and of CRISPR/Cas9-mediated mutagenesis
DNA double-stranded breaks can be repaired through error-prone pathways. Here, van Schendel et al. demonstrate that C. elegansacquires inheritable mutations through the use of polymerase theta-mediated end joining.
- Robin van Schendel
- , Sophie F. Roerink
- & Marcel Tijsterman
-
Article
| Open Access Allele-specific analysis of DNA replication origins in mammalian cells
Allele-specific analysis of DNA replication origins in mammalian cells
DNA sequences contribute to the location and timing of replication origin firings. Here by allele-specific analysis, the authors show that replication asynchrony is associated with small cumulative variations in the initiation efficiency of origins, rather than with the activation of dormant origins.
- Boris Bartholdy
- , Rituparna Mukhopadhyay
- & Eric E. Bouhassira
-
Article |
 Oncogenes create a unique landscape of fragile sites
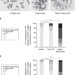
Oncogenes create a unique landscape of fragile sites
Aberrant oncogene expression can cause replication stress leading to chromosomal breaks. Here the authors map the chromosomal break loci induced by two different oncogenes and by a replication inhibitor, and show that each treatment induces a unique pattern of breaks in the same cell type.
- Karin Miron
- , Tamar Golan-Lev
- & Batsheva Kerem
-
Article |
 Sequential growth of long DNA strands with user-defined patterns for nanostructures and scaffolds
Sequential growth of long DNA strands with user-defined patterns for nanostructures and scaffolds
Assembling defined sequences of DNA is important for many applications, but the synthesis becomes more difficult as the target size increases. Here, the authors report a method for assembling DNA by combining smaller strands, with the final structure determined by the order of addition of the fragments.
- Graham D. Hamblin
- , Janane F. Rahbani
- & Hanadi F. Sleiman
-
Article |
 The DNA repair endonuclease Mus81 facilitates fast DNA replication in the absence of exogenous damage
The DNA repair endonuclease Mus81 facilitates fast DNA replication in the absence of exogenous damage
Several mechanisms are in place to ensure the accurate and timely replication of the genome as cells progress through S-phase. Here, the authors show that Mus81, an endonuclease involved in the response to DNA damage during replicative stress, also regulates the rate of DNA replication during normal growth.
- Haiqing Fu
- , Melvenia M. Martin
- & Mirit I. Aladjem
-
Article
| Open Access Cmr1/WDR76 defines a nuclear genotoxic stress body linking genome integrity and protein quality control
Cmr1/WDR76 defines a nuclear genotoxic stress body linking genome integrity and protein quality control
Defects in the DNA replication checkpoint can lead to genomic instability and cancer. Here the authors show that Cmr1/WDR76 participates in the DNA replication stress response and—along with several other components—defines a new cellular compartment that forms during cellular stress.
- Irene Gallina
- , Camilla Colding
- & Michael Lisby
-
Article |
 Structure of p15PAF–PCNA complex and implications for clamp sliding during DNA replication and repair
Structure of p15PAF–PCNA complex and implications for clamp sliding during DNA replication and repair
p15PAF regulates DNA replication and repair via interactions with the Proliferating Cell Nuclear Antigen (PCNA) sliding clamp. Here the authors present multi-faceted structural analyses of the p15-PCNA-DNA complex that suggests p15 regulates the sliding of PCNA along DNA during replication.
- Alfredo De Biasio
- , Alain Ibáñez de Opakua
- & Francisco J. Blanco
-
Article |
 FBH1 influences DNA replication fork stability and homologous recombination through ubiquitylation of RAD51
FBH1 influences DNA replication fork stability and homologous recombination through ubiquitylation of RAD51
The F-box DNA helicase 1 (FBH1) is implicated in suppression of homologous recombination (HR), but the precise mechanism is unclear. Here, the authors show that FBH1 can ubiquitylate RAD51, a central player in HR, and controls the subcellular localization of RAD51.
- Wai Kit Chu
- , Miranda J. Payne
- & Ian D. Hickson
-
Article
| Open Access Slow unloading leads to DNA-bound β2-sliding clamp accumulation in live Escherichia coli cells
Slow unloading leads to DNA-bound β2-sliding clamp accumulation in live Escherichia coli cells
DNA replication is accomplished by the replisome, a multi-protein complex that comprises the sliding clamp. Here, Moolman et al. present quantitative and dynamic measurements of the number of β2-sliding clamps at the single-cell level in live E. colicells to shed light on key aspects of DNA replication.
- M. Charl Moolman
- , Sriram Tiruvadi Krishnan
- & Nynke H. Dekker
-
Article |
 The CDC13-STN1-TEN1 complex stimulates Pol α activity by promoting RNA priming and primase-to-polymerase switch
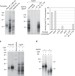
The CDC13-STN1-TEN1 complex stimulates Pol α activity by promoting RNA priming and primase-to-polymerase switch
The Cdc13-Stn1-Ten1 (CST) complex recognizes the G-strand of telomeres and functions in telomere protection and replication. Using purified components, the authors show that the Stn1 subunit of the CST complex stimulates RNA priming and the primase-to-polymerase switch by primase-Pol α in fungal and human systems.
- Neal F. Lue
- , Jamie Chan
- & Jerard Hurwitz
-
Article |
 PP2A and Aurora differentially modify Cdc13 to promote telomerase release from telomeres at G2/M phase
PP2A and Aurora differentially modify Cdc13 to promote telomerase release from telomeres at G2/M phase
Telomere maintenance requires proper termination of telomere replication at G2/M cell cycle stage. Here, Shen et al.show that termination of telomere replication requires PP2A phosphatase and Aurora kinase, which work independently but additively to remove active telomerase from telomeres.
- Zih-Jie Shen
- , Pang-Hung Hsu
- & Shu-Chun Teng
-
Article |
 Stabilization and targeting of INO80 to replication forks by BAP1 during normal DNA synthesis
Stabilization and targeting of INO80 to replication forks by BAP1 during normal DNA synthesis
The INO80 complex is involved in chromatin remodelling during transcription and has been implicated in DNA repair. Here, the authors establish a role for INO80 in normal DNA replication and reveal a mechanism that targets this remodeler to replication forks via BAP1 and H2A ubiquitination.
- Han-Sae Lee
- , Shin-Ai Lee
- & Jongbum Kwon
-
Article |
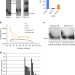 Sgs1 and Sae2 promote telomere replication by limiting accumulation of ssDNA
Sgs1 and Sae2 promote telomere replication by limiting accumulation of ssDNA
The enzymes Sae2 and Sgs1 regulate telomere maintenance in yeast cells that are telomerase-positive or -negative, but how they do this is unclear. Here the authors show that Sae2 and Sgs1 facilitate telomere replication in telomerase-positive cells, but generate single-stranded DNA at eroded telomeres in telomerase-negative cells.
- Julien Hardy
- , Dmitri Churikov
- & Marie-Noëlle Simon
-
Article |
 A role for DNA polymerase θ in the timing of DNA replication
A role for DNA polymerase θ in the timing of DNA replication
DNA polymerase θ (Pol θ) exhibits properties typical of translesion and repair synthesis; however, its physiological function remains elusive. Here, the authors show that Pol θ plays a role in the initiation and timing of DNA replication during human cell division.
- Anne Fernandez-Vidal
- , Laure Guitton-Sert
- & Jean-Sébastien Hoffmann
-
Article
| Open Access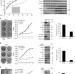 Polycomb proteins control proliferation and transformation independently of cell cycle checkpoints by regulating DNA replication
Polycomb proteins control proliferation and transformation independently of cell cycle checkpoints by regulating DNA replication
Polycomb (PcG) proteins are known to promote cell proliferation by silencing expression of the tumour suppressor Ink4A-Arf. Piunti et al.show that PcG proteins also regulate tumour progression independently of this role, revealing a requirement for PRC1 and PRC2 in replication fork progression.
- Andrea Piunti
- , Alessandra Rossi
- & Diego Pasini
-
Article |
 The Escherichia coli Tus–Ter replication fork barrier causes site-specific DNA replication perturbation in yeast
The Escherichia coli Tus–Ter replication fork barrier causes site-specific DNA replication perturbation in yeast
Analysis of DNA replication fork arrest is challenging due to lack of techniques that induce site-specific replication barriers in eukaryotic chromosomes. Here, Larsen et al. create a novel tool to trigger targeted replication fork pausing by engineering the bacterial Tus–Tersystem into the yeast genome.
- Nicolai B. Larsen
- , Ehud Sass
- & Ian D. Hickson
-
Article |
 Tousled-like kinases phosphorylate Asf1 to promote histone supply during DNA replication
Tousled-like kinases phosphorylate Asf1 to promote histone supply during DNA replication
DNA replication requires re-establishment of chromatin structure, which involves incorporation of newly synthesized histones. Here, Klimovskaia et al.show that phosphorylation of the histone chaperone Asf1 by Tousled-Like Kinase stimulates its ability to bind histones, thus promoting chromatin assembly.
- Ilnaz M. Klimovskaia
- , Clifford Young
- & Anja Groth
-
Article |
 Poly(ADP-ribose) binding to Chk1 at stalled replication forks is required for S-phase checkpoint activation
Poly(ADP-ribose) binding to Chk1 at stalled replication forks is required for S-phase checkpoint activation
DNA damage at stalled replication forks activates Chk1 kinase and poly(ADP-ribose) (PAR) polymerase 1. Min et al.find that retention of Chk1 to stalled replication forks depends on its direct interaction with PAR, and show that PAR chain length fine-tunes Chk1 and S-phase checkpoint activation.
- WooKee Min
- , Christopher Bruhn
- & Zhao-Qi Wang
-
Article |
 Stepwise histone modifications are mediated by multiple enzymes that rapidly associate with nascent DNA during replication
Stepwise histone modifications are mediated by multiple enzymes that rapidly associate with nascent DNA during replication
Chromatin marks have to be re-established after DNA replication. Here Petruk et al. show that many histone-modifying enzymes are found in close proximity to newly replicated DNA in cells of Drosophilaembryos before the corresponding histone marks are re-established.
- Svetlana Petruk
- , Kathryn L. Black
- & Alexander Mazo
-
Article
| Open Access RecG and UvsW catalyse robust DNA rewinding critical for stalled DNA replication fork rescue
RecG and UvsW catalyse robust DNA rewinding critical for stalled DNA replication fork rescue
The helicases UvsW and RecG have both unwinding and rewinding activities and are involved in the rescue of stalled DNA replication forks. Here Manosas et al. use single-molecule techniques to characterize the rewinding activities of the two helicases, concluding that rewinding is actively catalysed.
- Maria Manosas
- , Senthil K. Perumal
- & Vincent Croquette
-
Article
| Open Access Conformational landscapes of DNA polymerase I and mutator derivatives establish fidelity checkpoints for nucleotide insertion
Conformational landscapes of DNA polymerase I and mutator derivatives establish fidelity checkpoints for nucleotide insertion
The fidelity of DNA polymerases depends on conformational changes that promote the rejection of incorrect nucleotides. Here, by using an intramolecular single-molecule FRET assay, the authors establish and characterize the partially closed conformation as a crucial fidelity checkpoint.
- Johannes Hohlbein
- , Louise Aigrain
- & Achillefs N. Kapanidis
-
Article |
 A spontaneous Cdt1 mutation in 129 mouse strains reveals a regulatory domain restraining replication licensing
A spontaneous Cdt1 mutation in 129 mouse strains reveals a regulatory domain restraining replication licensing
Cdt1 is part of a protein complex that regulates the initiation of DNA replication. Here Coulombe et al. identify a PEST-like regulatory domain in the N terminus of Cdt1 that prevents premature initiation of DNA synthesis during the cell cycle.
- Philippe Coulombe
- , Damien Grégoire
- & Marcel Méchali
-
Article
| Open Access Fork sensing and strand switching control antagonistic activities of RecQ helicases
Fork sensing and strand switching control antagonistic activities of RecQ helicases
RecQ helicases are enzymes that play a central role in maintaining genome stability in the DNA repair cascade. Klaue et al. show that RecQ2 and RecQ3 from Arabidopsis thalianaprocess DNA by, respectively, unwinding and rewinding forked DNA substrates, using a frequent strand switching mechanism.
- Daniel Klaue
- , Daniela Kobbe
- & Ralf Seidel
-
Article
| Open Access Aurora-A controls pre-replicative complex assembly and DNA replication by stabilizing geminin in mitosis
Aurora-A controls pre-replicative complex assembly and DNA replication by stabilizing geminin in mitosis
Geminin blocks the inappropriate assembly of pre-replication complexes on DNA, and this activity is inhibited in G1 by its proteasomal degradation. Tsunematsu et al.demonstrate that geminin is stabilized during mitosis due to its phosphorylation by the mitotic kinase Aurora-A.
- Takaaki Tsunematsu
- , Yoshihiro Takihara
- & Yasusei Kudo
-
Article
| Open Access SUMO2/3 modification of cyclin E contributes to the control of replication origin firing
SUMO2/3 modification of cyclin E contributes to the control of replication origin firing
The organized initiation of DNA replication at sites throughout the genome must be carefully choreographed to maintain genome stability. Bonne-Andrea and colleagues show that protein SUMOylation controls the density of origin firing, and identify cyclin E as an important substrate in this context.
- Catherine Bonne-Andrea
- , Malik Kahli
- & Olivier Coux
-
Article |
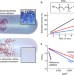 Impact of macromolecular crowding on DNA replication
Impact of macromolecular crowding on DNA replication
Macromolecular crowding significantly affects interactions between macromolecules such as proteins, DNA and RNA. Akabayov and colleagues use a SAXS reconstitution assay to show that the effect of macromolecular crowding on T7 DNA replication causes structural changes of the replisome.
- Barak Akabayov
- , Sabine R. Akabayov
- & Charles C. Richardson
-
Article |
 DNA replication timing and higher-order nuclear organization determine single-nucleotide substitution patterns in cancer genomes
DNA replication timing and higher-order nuclear organization determine single-nucleotide substitution patterns in cancer genomes
Human cancer genomes often contain large amounts of single-nucleotide substitutions (SNS). Liu et al. catalogued SNS signatures across various cancer and normal genomes, demonstrating coordinative effects between replication timing and nuclear architecture on SNS patterns in cancer genomes.
- Lin Liu
- , Subhajyoti De
- & Franziska Michor
-
Article |
 FBH1 co-operates with MUS81 in inducing DNA double-strand breaks and cell death following replication stress
FBH1 co-operates with MUS81 in inducing DNA double-strand breaks and cell death following replication stress
DNA replication stress promotes genome instability and cell death. Here Fugger et al.describe how FBH1, via its helicase activity, is required to eliminate cells with excessive DNA replication stress, through the generation of MUS81-induced DNA double-strand breaks.
- Kasper Fugger
- , Wai Kit Chu
- & Claus Storgaard Sørensen
-
Article |
 DNA replication timing and selection shape the landscape of nucleotide variation in cancer genomes
DNA replication timing and selection shape the landscape of nucleotide variation in cancer genomes
Cancer cells form by somatic mutations and natural selection, but how these factors affect tumorigenesis is not clear. Here, somatic mutations are characterized in human cancer genomes, revealing that DNA replication timing influences the frequency of single-nucleotide variants in different genomic regions.
- Yong H Woo
- & Wen-Hsiung Li
-
Article |
 Pyrimidine pool imbalance induced by BLM helicase deficiency contributes to genetic instability in Bloom syndrome
Pyrimidine pool imbalance induced by BLM helicase deficiency contributes to genetic instability in Bloom syndrome
Mutations in the DNA helicaseBLM cause Bloom syndrome, which is characterized by slow replication fork progression and genetic instability. Here, cells lacking BLMare shown to have a defect in cytidine deaminase, which alters the pyrimidine pool and results in replication fork progression with altered velocity.
- Pauline Chabosseau
- , Géraldine Buhagiar-Labarchède
- & Mounira Amor-Guéret
-
Article
| Open Access Histone acetylation controls the inactive X chromosome replication dynamics
Histone acetylation controls the inactive X chromosome replication dynamics
How one copy of the X chromosome is silenced in replicating female somatic cells is poorly understood. Here, the authors demonstrate that the inactive X chromosome is replicated before constitutive heterochromatin and that histone hypoacetylation has a role in controlling replication of the inactive X chromosome.
- Corella S. Casas-Delucchi
- , Alessandro Brero
- & M. Cristina Cardoso
-
Article
| Open Access Herpesviruses carrying a Brainbow cassette reveal replication and expression of limited numbers of incoming genomes
Herpesviruses carrying a Brainbow cassette reveal replication and expression of limited numbers of incoming genomes
The replication of viral genomes in infected cells is required for successful infection. In this study, using Cre-conditional expression of multiple coloured fluorophores, the authors demonstrate that the number of viral genomes expressed and replicated in a cell is surprisingly limited.
- Oren Kobiler
- , Yaron Lipman
- & Lynn W. Enquist

 RPA and Rad51 constitute a cell intrinsic mechanism to protect the cytosol from self DNA
RPA and Rad51 constitute a cell intrinsic mechanism to protect the cytosol from self DNA