Featured
-
-
Article
| Open Access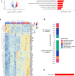 Accelerated DNA replication fork speed due to loss of R-loops in myelodysplastic syndromes with SF3B1 mutation
Accelerated DNA replication fork speed due to loss of R-loops in myelodysplastic syndromes with SF3B1 mutation
Here the authors find that erythroblasts of myelodysplastic syndromes with SF3B1 mutation leading to inefficient erythropoiesis show DNA replication stress with accelerated forks and reduced R-loops. Restoring R-loops by a histone deacetylase inhibitor rescues erythroid differentiation.
- David Rombaut
- , Carine Lefèvre
- & Michaela Fontenay
-
Article
| Open Access ZNF827 is a single-stranded DNA binding protein that regulates the ATR-CHK1 DNA damage response pathway
ZNF827 is a single-stranded DNA binding protein that regulates the ATR-CHK1 DNA damage response pathway
Here, the authors characterise the zinc finger protein ZNF827 as a single stranded DNA binding protein that accumulates at stalled replication forks to activate the ATR-CHK1 pathway and engage homologous-recombination mediated DNA repair.
- Sile F. Yang
- , Christopher B. Nelson
- & Hilda A. Pickett
-
Article
| Open Access RTF2 controls replication repriming and ribonucleotide excision at the replisome
RTF2 controls replication repriming and ribonucleotide excision at the replisome
Ribonucleotides are incorporated into DNA during every S phase. Here, the authors show that replisome protein RTF2 localizes RNase H2 to the replisome, promoting ribonucleotide removal by RNase H2 when replication is ongoing but interfering with PRIM1-dependent restart following fork stalling.
- Brooke A. Conti
- , Penelope D. Ruiz
- & Agata Smogorzewska
-
Article
| Open Access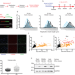 Oncogenic c-Myc induces replication stress by increasing cohesins chromatin occupancy in a CTCF-dependent manner
Oncogenic c-Myc induces replication stress by increasing cohesins chromatin occupancy in a CTCF-dependent manner
Here the authors report that oncogenic c-Myc induces replication stress via increasing the amount of cohesins bound to chromatin at CTCF sites.
- Silvia Peripolli
- , Leticia Meneguello
- & Robertus A. M. de Bruin
-
Article
| Open Access The chromatin-associated lncREST ensures effective replication stress response by promoting the assembly of fork signaling factors
The chromatin-associated lncREST ensures effective replication stress response by promoting the assembly of fork signaling factors
Replication stress represents a major threat to genome integrity of normal and cancer cells. Here, the authors find that the long non-coding RNA lncREST affects the replication stress response through interaction with nucleolin. This interaction bridges the recruitment of replication factors to stressed chromatin.
- Luisa Statello
- , José Miguel Fernandez-Justel
- & Maite Huarte
-
Article
| Open Access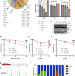 FLIP(C1orf112)-FIGNL1 complex regulates RAD51 chromatin association to promote viability after replication stress
FLIP(C1orf112)-FIGNL1 complex regulates RAD51 chromatin association to promote viability after replication stress
Recombination is essential for life. Here, the authors characterize FLIP as a novel regulator of the key recombination protein RAD51’s functions. FLIP loss caused marked sensitivity to DNA damage, increased DNA breakage and defective replication.
- Jessica D. Tischler
- , Hiroshi Tsuchida
- & Richard O. Adeyemi
-
Article
| Open Access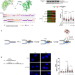 CaMKK2 and CHK1 phosphorylate human STN1 in response to replication stress to protect stalled forks from aberrant resection
CaMKK2 and CHK1 phosphorylate human STN1 in response to replication stress to protect stalled forks from aberrant resection
Here the authors show that the calcium-sensing kinase CaMKK2 phosphorylates STN1 in response to replication stress and elevated cytosolic calcium concentration to protect stalled replication forks from aberrant MRE11 degradation. Cancer-associated STN1 mutations abolish STN1 phosphorylation, resulting in fork instability.
- Rishi Kumar Jaiswal
- , Kai-Hang Lei
- & Weihang Chai
-
Article
| Open Access Nuclear actin polymerization rapidly mediates replication fork remodeling upon stress by limiting PrimPol activity
Nuclear actin polymerization rapidly mediates replication fork remodeling upon stress by limiting PrimPol activity
How nuclear architecture assists the replication stress response is still largely unknown. Here the authors show that nuclear actin polymerization rapidly extends upon mild DNA damage. By limiting Primpol activity, this response mediates fork slowing and reversal, protecting chromosome stability.
- Maria Dilia Palumbieri
- , Chiara Merigliano
- & Massimo Lopes
-
Article
| Open Access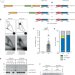 Gene duplication and deletion caused by over-replication at a fork barrier
Gene duplication and deletion caused by over-replication at a fork barrier
Gene duplications and deletions are important drivers of evolution and disease. Here, the authors show that excess DNA generated at a replication fork barrier can be integrated at a new genomic site causing both a gene duplication and a deletion.
- Judith Oehler
- , Carl A. Morrow
- & Matthew C. Whitby
-
Article
| Open Access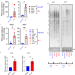 Alternative lengthening of telomeres (ALT) cells viability is dependent on C-rich telomeric RNAs
Alternative lengthening of telomeres (ALT) cells viability is dependent on C-rich telomeric RNAs
ALT cells use an alternative lengthening mechanism of telomeres and bear telomeric DNA damage with increased levels of damage-induced long non-coding RNA. Here the AUs show that antisense oligonucleotides (ASO) targeting such RNAs can induce ALT cancer cells selective cell death.
- Ilaria Rosso
- , Corey Jones-Weinert
- & Fabrizio d’Adda di Fagagna
-
Article
| Open Access Structural basis for stabilisation of the RAD51 nucleoprotein filament by BRCA2
Structural basis for stabilisation of the RAD51 nucleoprotein filament by BRCA2
Here the authors report the cryoEM structure of the RAD51 nucleoprotein filament bound to the C-terminal TR2 domain of BRCA2. The structure explains how BRCA2 stabilises the filament and uncovers a conserved mechanism of filament binding by recombination mediators.
- Robert Appleby
- , Luay Joudeh
- & Luca Pellegrini
-
Article
| Open Access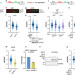 Interferon restores replication fork stability and cell viability in BRCA-defective cells via ISG15
Interferon restores replication fork stability and cell viability in BRCA-defective cells via ISG15
Here the authors show that the basal activation of the interferon/ISG15 pathway is required for the stability of nascent DNA during replication and its upregulation promotes viability, proliferation and acquisition of drug resistance in BRCA1/2 deficient cells.
- Ramona N. Moro
- , Uddipta Biswas
- & Lorenza Penengo
-
Article
| Open Access The COMPASS subunit Spp1 protects nascent DNA at the Tus/Ter replication fork barrier by limiting DNA availability to nucleases
The COMPASS subunit Spp1 protects nascent DNA at the Tus/Ter replication fork barrier by limiting DNA availability to nucleases
The Spp1 subunit of Set1C is recruited to the Tus/Ter replication fork barrier via its PHD domain and protects the replication fork by preventing excessive ssDNA formation, stressing the importance of chromatin structure in the replication stress response.
- Nagham Ghaddar
- , Yves Corda
- & Vincent Géli
-
Article
| Open Access RNAPII-dependent ATM signaling at collisions with replication forks
RNAPII-dependent ATM signaling at collisions with replication forks
Deregulation of transcription by oncogenes leads to collisions of RNA Polymerase II (RNAPII) with DNA replication machinery (transcription-replication conflicts, TRCs). This study shows that RNAPII activates ATM kinase at TRCs providing a mechanism for replication fork stalling and ATM activation at TRCs.
- Elias Einig
- , Chao Jin
- & Nikita Popov
-
Article
| Open Access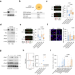 Trim33 masks a non-transcriptional function of E2f4 in replication fork progression
Trim33 masks a non-transcriptional function of E2f4 in replication fork progression
Here the authors show that under replicative stress the E2f4 transcription factor recruits the Recql DNA helicase to facilitate DNA replication. The Trim33 ubiquitin ligase targets E2f4 to limit its interactions with Recql and chromatin.
- Vanessa Rousseau
- , Elias Einig
- & Nikita Popov
-
Article
| Open Access TRAIP resolves DNA replication-transcription conflicts during the S-phase of unperturbed cells
TRAIP resolves DNA replication-transcription conflicts during the S-phase of unperturbed cells
The TRAIP E3 ubiquitin ligase is essential for genome integrity, mutations lead to primordial dwarfism in patients. Here, the authors show that TRAIP degradation in S-phase, results in cell arrest due to DNA damage caused by replication-transcription conflicts.
- Shaun Scaramuzza
- , Rebecca M. Jones
- & Agnieszka Gambus
-
Article
| Open Access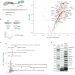 Nuclear myosin VI maintains replication fork stability
Nuclear myosin VI maintains replication fork stability
Whether actin and associated molecules have roles in the nucleus is an active area of study. Here Shi et al. report a nuclear function of the actin-based motor myosin VI in protecting stalled replication forks from nuclease-mediated degradation.
- Jie Shi
- , Kristine Hauschulte
- & Hans-Peter Wollscheid
-
Article
| Open Access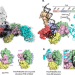 Replication fork binding triggers structural changes in the PriA helicase that govern DNA replication restart in E. coli
Replication fork binding triggers structural changes in the PriA helicase that govern DNA replication restart in E. coli
The mechanism of replication restart initiation by the bacterial DNA replication restart proteins PriA and PriB is resolved, revealing a switch-like restructuring of PriA triggered by replication fork binding that mediates PriA/PriB complex assembly.
- Alexander T. Duckworth
- , Peter L. Ducos
- & James L. Keck
-
Article
| Open Access Excessive reactive oxygen species induce transcription-dependent replication stress
Excessive reactive oxygen species induce transcription-dependent replication stress
Excessive oxidative stress is widely perceived as a key factor in cancer progression. Here, the authors reveal that oxidative stress induces transcription-dependent replication fork stalling that appears to be a major source of chromosomal rearrangements found in human cancers.
- Martin Andrs
- , Henriette Stoy
- & Pavel Janscak
-
Article
| Open Access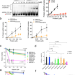 Replication gap suppression depends on the double-strand DNA binding activity of BRCA2
Replication gap suppression depends on the double-strand DNA binding activity of BRCA2
Here the authors demonstrate that the dsDNA binding function at the N-terminus of BRCA2 prevents nucleotide depletion-dependent replicative ssDNA gaps but not those induced by PARP inhibition. This function is impaired in breast-cancer variants affecting this region.
- Domagoj Vugic
- , Isaac Dumoulin
- & Aura Carreira
-
Article
| Open Access Profilin-1 regulates DNA replication forks in a context-dependent fashion by interacting with SNF2H and BOD1L
Profilin-1 regulates DNA replication forks in a context-dependent fashion by interacting with SNF2H and BOD1L
Subcellular localization plays an important yet underappreciated role in protein functions. Here the authors defined novel and context-dependent nuclear functions of the actin-binding factor profilin-1 in DNA replication fork dynamics and stability.
- Cuige Zhu
- , Mari Iwase
- & Jieya Shao
-
Article
| Open Access ISG15 conjugation to proteins on nascent DNA mitigates DNA replication stress
ISG15 conjugation to proteins on nascent DNA mitigates DNA replication stress
DNA replication stress can result in genome instability. Here the authors show that the ubiquitin like modifier protein, ISG15, important during the innate immune response, acts at replication forks to mitigate DNA replication stress.
- Christopher P. Wardlaw
- & John H. J. Petrini
-
Article
| Open Access MAD2L2 promotes replication fork protection and recovery in a shieldin-independent and REV3L-dependent manner
MAD2L2 promotes replication fork protection and recovery in a shieldin-independent and REV3L-dependent manner
MAD2L2 – as a member of the shieldin complex - counteracts resection during DNA repair. Here the authors demonstrate that MAD2L2 protects stalled replication forks from excessive resection, in a shieldin-independent and REV3L-dependent manner.
- Inés Paniagua
- , Zainab Tayeh
- & Jacqueline J. L. Jacobs
-
Article
| Open Access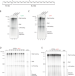 The mechanism of replication stalling and recovery within repetitive DNA
The mechanism of replication stalling and recovery within repetitive DNA
DNA replication of repetitive sequences was recreated in a test tube using purified components. DNA alone was sufficient to induce stalling. Both stalling and recovery were dictated by the capacity of DNA to fold into unusual secondary structures.
- Corella S. Casas-Delucchi
- , Manuel Daza-Martin
- & Gideon Coster
-
Article
| Open Access Genome-wide mapping of individual replication fork velocities using nanopore sequencing
Genome-wide mapping of individual replication fork velocities using nanopore sequencing
Theulot et al. introduce NanoForkSpeed, a nanopore sequencing-based method to map individual replication fork velocities on entire genomes. NFS shows that fork speed is uniform across yeast chromosomes except for a marked slowdown at pausing sites.
- Bertrand Theulot
- , Laurent Lacroix
- & Benoît Le Tallec
-
Article
| Open Access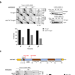 Rad51-mediated replication of damaged templates relies on monoSUMOylated DDK kinase
Rad51-mediated replication of damaged templates relies on monoSUMOylated DDK kinase
Joseph et al. reveal that monoSUMOylated DDK kinase, implicated in replication initiation, acts with Rad51 recombinase to prevent replication fork uncoupling and to mediate recombination-dependent gap-filling in the presence of genotoxic stress.
- Chinnu Rose Joseph
- , Sabrina Dusi
- & Dana Branzei
-
Article
| Open Access WRN helicase safeguards deprotected replication forks in BRCA2-mutated cancer cells
WRN helicase safeguards deprotected replication forks in BRCA2-mutated cancer cells
The tumor suppressor BRCA2 protects stalled DNA replication forks from unrestrained degradation; however the mechanism whereby unprotected stalled forks are preserved and restarted has remained elusive. Here the authors show that the WRN helicase promotes stalled fork recovery and limits fork hyper-degradation in the absence of BRCA2 protection.
- Arindam Datta
- , Kajal Biswas
- & Robert M. Brosh Jr
-
Article
| Open Access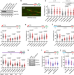 BRCA2 associates with MCM10 to suppress PRIMPOL-mediated repriming and single-stranded gap formation after DNA damage
BRCA2 associates with MCM10 to suppress PRIMPOL-mediated repriming and single-stranded gap formation after DNA damage
Tumor suppressor BRCA2 is known to stabilize and restart stalled DNA replication forks. Here the authors show that BRCA2 is recruited to the replication fork through its interaction with MCM10 and inhibits Primase-Polymerase-mediated repriming, lesion bypass and single strand DNA gap formation after DNA damage.
- Zhihua Kang
- , Pan Fu
- & Bing Xia
-
Article
| Open Access Dynamics of replication origin over-activation
Dynamics of replication origin over-activation
DNA replication processes are often dysregulated in cancer. Here the authors analyse DNA synthesis patterns in cancer cells undergoing partial genome re-replication to reveal that re-replication exhibits aberrant replication fork dynamics and a skewed distribution of replication initiation that over-duplicates early-replicating genomic regions.
- Haiqing Fu
- , Christophe E. Redon
- & Mirit I. Aladjem
-
Article
| Open Access Smc5/6 functions with Sgs1-Top3-Rmi1 to complete chromosome replication at natural pause sites
Smc5/6 functions with Sgs1-Top3-Rmi1 to complete chromosome replication at natural pause sites
Smc5/6, part of the structural maintenance of chromosomes (SMC) family, plays roles in genome structural integrity. Here the authors reveal that Smc5/6 acts jointly with Top3 within the STR complex to mediate DNA replication completion at genomic natural pausing sites (NPSs).
- Sumedha Agashe
- , Chinnu Rose Joseph
- & Dana Branzei
-
Article
| Open Access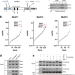 The Bloom syndrome complex senses RPA-coated single-stranded DNA to restart stalled replication forks
The Bloom syndrome complex senses RPA-coated single-stranded DNA to restart stalled replication forks
The BLM helicase interacts with the topoisomerase TOP3A and RMI1 to form the BTR complex. Here, the authors reveal that this complex contains multiple binding sites for the single-stranded DNA-binding complex RPA, and that RPA-binding stimulates BLM recruitment to stalled replication forks to promote their restart after replication stress.
- Ann-Marie K. Shorrocks
- , Samuel E. Jones
- & Andrew N. Blackford
-
Article
| Open Access Structural basis for the multi-activity factor Rad5 in replication stress tolerance
Structural basis for the multi-activity factor Rad5 in replication stress tolerance
Rad5 is a hub connecting three replication stress tolerance pathways. Here, the authors present the 3.3 Å crystal structure of a N-terminal truncated K.lactis Rad5 construct that reveals the spatial arrangement of the HIRAN, Snf2 and RING domains and structure-guided in vitro and in vivo experiments reveal multiple activities of the yeast Rad5 HIRAN domain among them a role in binding PCNA and supporting its ubiquitination.
- Miaomiao Shen
- , Nalini Dhingra
- & Song Xiang
-
Article
| Open Access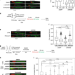 Resolution of R-loops by INO80 promotes DNA replication and maintains cancer cell proliferation and viability
Resolution of R-loops by INO80 promotes DNA replication and maintains cancer cell proliferation and viability
In mammalian cells, during transcription and replication, RNA:DNA hybrid structures known as R-loops can arise, posing as obstacles to replication fork progression. Here the authors reveal that the ATP-dependent chromatin remodelling INO80 complex promotes resolution of R-loops to prevent replication associated DNA damage in cancer cells.
- Lisa Prendergast
- , Urszula L. McClurg
- & Manolis Papamichos-Chronakis
-
Article
| Open Access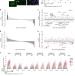 Sequential role of RAD51 paralog complexes in replication fork remodeling and restart
Sequential role of RAD51 paralog complexes in replication fork remodeling and restart
Replication stress has been associated with transient remodelling of replication intermediates into reversed forks, followed by efficient fork restart. Here the authors systematically analyse the role of RAD51 paralogs in these transactions, providing insights on the mechanistic role of different complexes of these proteins.
- Matteo Berti
- , Federico Teloni
- & Massimo Lopes
-
Article
| Open Access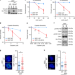 Ubiquitinated-PCNA protects replication forks from DNA2-mediated degradation by regulating Okazaki fragment maturation and chromatin assembly
Ubiquitinated-PCNA protects replication forks from DNA2-mediated degradation by regulating Okazaki fragment maturation and chromatin assembly
PCNA is essential for DNA replication and cellular proliferation. Here, the authors reveal that PCNA ubiquitination protects stalled replication forks from DNA2-mediated degradation via regulation of Okazaki fragment maturation and chromatin assembly.
- Tanay Thakar
- , Wendy Leung
- & George-Lucian Moldovan
-
Article
| Open Access The nuclear pore complex prevents sister chromatid recombination during replicative senescence
The nuclear pore complex prevents sister chromatid recombination during replicative senescence
The Nuclear Pore Complex has been linked to DNA damage processing. Here the authors reveal that the Nup1 C-terminus is critical for the relocalization of eroded telomeres to nuclear pores and that modification of Nup1 promotes sister chromatid recombination and unleashes a new telomere maintenance mechanism.
- Paula Aguilera
- , Jenna Whalen
- & Vincent Géli
-
Article
| Open Access ATAD5 promotes replication restart by regulating RAD51 and PCNA in response to replication stress
ATAD5 promotes replication restart by regulating RAD51 and PCNA in response to replication stress
How the replisome machinery contributes to fork stability under replication stress is currently not clear. Here the authors reveal a role for ATAD5 in maintaining genome integrity during replication stress by promoting replication restart through RAD51/PCNA regulation.
- Su Hyung Park
- , Nalae Kang
- & Kyungjae Myung
-
Article
| Open Access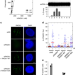 Non-enzymatic roles of human RAD51 at stalled replication forks
Non-enzymatic roles of human RAD51 at stalled replication forks
RAD51 has been implicated in replication fork processing and restart in response to replication stress. Here, authors reveal mechanistic aspects of non-enzymatic roles of RAD51 for fork reversal and cooperation with FBH1.
- Jennifer M. Mason
- , Yuen-Ling Chan
- & Douglas K. Bishop
-
Article
| Open Access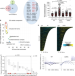 MRE11-RAD50-NBS1 promotes Fanconi Anemia R-loop suppression at transcription–replication conflicts
MRE11-RAD50-NBS1 promotes Fanconi Anemia R-loop suppression at transcription–replication conflicts
Accumulations of R-loops can lead to genome instability. Here the authors reveal a role for the MRN complex in suppressing R-loops and associated DNA damage at transcription–replication conflicts.
- Emily Yun-Chia Chang
- , Shuhe Tsai
- & Peter C. Stirling
-
Article
| Open Access qDSB-Seq is a general method for genome-wide quantification of DNA double-strand breaks using sequencing
qDSB-Seq is a general method for genome-wide quantification of DNA double-strand breaks using sequencing
Measuring relative frequencies of DNA double-strand breaks between loci does not provide the full physiological relevance of those breaks. Here Rowicka and colleagues present qDSB-Seq method which uses spike-in double-strand breaks induced by a restriction enzyme to accurately quantify DNA damage.
- Yingjie Zhu
- , Anna Biernacka
- & Maga Rowicka
-
Article
| Open Access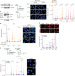 Rad52 prevents excessive replication fork reversal and protects from nascent strand degradation
Rad52 prevents excessive replication fork reversal and protects from nascent strand degradation
Stabilisation of stalled replication forks prevents excessive fork reversal and genome instability. Here authors reveal a RAD52-dependent replication fork protection mechanism.
- Eva Malacaria
- , Giusj Monia Pugliese
- & Pietro Pichierri
-
Article
| Open Access Overexpression of Claspin and Timeless protects cancer cells from replication stress in a checkpoint-independent manner
Overexpression of Claspin and Timeless protects cancer cells from replication stress in a checkpoint-independent manner
Oncogene-induced replication stress (RS) promotes cancer development. Here, the authors report that cancer cells adapt to oncogene-induced RS by overexpressing downstream components of ATR-CHK1 pathway, Claspin and Timeless, which have protective role at the replication forks independent of their checkpoint function.
- Julien N. Bianco
- , Valérie Bergoglio
- & Philippe Pasero
-
Article
| Open Access Lnk/Sh2b3 deficiency restores hematopoietic stem cell function and genome integrity in Fancd2 deficient Fanconi anemia
Lnk/Sh2b3 deficiency restores hematopoietic stem cell function and genome integrity in Fancd2 deficient Fanconi anemia
Loss of Fancd2 leads to replication stress intolerance and Fanconi Anemia, where haematopoietic stem cell (HSC) function is compromised. Here, the authors show that Lnk/Sh2b3 loss restores HSC proliferation and survival in Fancd2 knockout mice and ameliorates replication stress in a cytokine/JAK2 signaling dependent manner.
- Joanna Balcerek
- , Jing Jiang
- & Wei Tong
-
Article
| Open Access The Swr1 chromatin-remodeling complex prevents genome instability induced by replication fork progression defects
The Swr1 chromatin-remodeling complex prevents genome instability induced by replication fork progression defects
SWR-C and its substrate the histone variant Htz1 are considered important for genome maintenance. Here the authors reveal that SWR-C/Htz1 plays a critical role during replication stress caused by absence of the replication fork progression proteins Mrc1/Tof1/Csm3.
- Anjana Srivatsan
- , Bin-Zhong Li
- & Richard D. Kolodner
-
Article
| Open Access SUMO2 conjugation of PCNA facilitates chromatin remodeling to resolve transcription-replication conflicts
SUMO2 conjugation of PCNA facilitates chromatin remodeling to resolve transcription-replication conflicts
Transcription-replication conflicts need to be resolved to minimize genome instability. Here the authors show that SUMO2-conjugated PCNA destabilizes RNAPII from chromatin, enhances replication progression and limits transcription-induced DNA damage at common fragile sites.
- Min Li
- , Xiaohua Xu
- & Yilun Liu
-
Article
| Open Access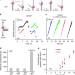 Helicase promotes replication re-initiation from an RNA transcript
Helicase promotes replication re-initiation from an RNA transcript
During DNA replication, replicative helicases play an essential role for DNA unwinding to occur. Here the authors find that bacteriophage T7 helicase is also involved in replication re-initiation by interacting with a non-replicating DNAP and increasing unwinding rate.
- Bo Sun
- , Anupam Singh
- & Michelle D. Wang
-
Article
| Open Access Chromatin conformation regulates the coordination between DNA replication and transcription
Chromatin conformation regulates the coordination between DNA replication and transcription
The maintenance of chromatin integrity during replication is critical for cell viability. Here the authors study how dividing cells respond to alterations in chromatin structure and find that these elicit a range of responses in the dynamics of DNA replication and consequences on replicative stress.
- Ricardo Almeida
- , José Miguel Fernández-Justel
- & María Gómez
-
Article
| Open Access Loss of PBRM1 rescues VHL dependent replication stress to promote renal carcinogenesis
Loss of PBRM1 rescues VHL dependent replication stress to promote renal carcinogenesis
Mutations in VHL have been linked to clear cell renal cancer, but the molecular mechanisms involved remain unclear. Here the authors generate a mouse model closely mimicking the human disease and show that VHL loss induces DNA replication stress that is rescued by the concomitant loss of PBRM1 permitting transformation.
- Judit Espana-Agusti
- , Anne Warren
- & Athena Matakidou
-
Article
| Open Access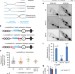 The end-joining factor Ku acts in the end-resection of double strand break-free arrested replication forks
The end-joining factor Ku acts in the end-resection of double strand break-free arrested replication forks
Terminally arrested replication forks are restarted through homologous recombination after processing single-stranded DNA gaps. Here the authors show that resection is regulated by the NHEJ factor Ku, helping to fine-tune recombination at forks.
- Ana Teixeira-Silva
- , Anissia Ait Saada
- & Sarah A. E. Lambert

 H2AX promotes replication fork degradation and chemosensitivity in BRCA-deficient tumours
H2AX promotes replication fork degradation and chemosensitivity in BRCA-deficient tumours