Abstract
UVB radiation causes cyclobutane pyrimidine dimers (CPDs) to form on the DNA of living organisms. This study found that overexpression of the silicon absorbance gene Lsi1 reduced the accumulation of CPDs in rice, which profited from the reactivation by photolyase. The transcript abundance of deoxyribodipyrimidine photolyase (Os10g0167600) was generally correlated with the silicon content of the rice, and the up-regulation of Os10g0167600 was found to be highest in the UVB-treated Lsi1-overexpressed (Lsi1-OX) rice. A trans-acting factor, methyl-CpG binding domain protein (OsMeCP), was found to interact with the cis-element of Os10g0167600. The nucleic location of OsMeCP effectively enabled the transcriptional regulation. Compared with the WT, the level of OsMeCP was lower in the Lsi1-OX rice but higher in the Lsi1-RNAi line. Rice cultured in a high silicate-concentration solution also exhibited less OsMeCP abundance. Overexpression of OsMeCP led to lower Os10g0167600 transcript levels and a higher CPD content than in the WT, but the reverse was true in the OsMeCP-RNAi line. These findings indicate that OsMeCP acts as a negative regulator of silicon, and can mediate the repression of the transcription from Os10g0167600, which inhibits the photoreactivation of the photolyase involved in the repair of CPDs.
Similar content being viewed by others
Introduction
Because the highly active components of radiation penetrate cells, solar ultraviolet B (UVB) radiation (280 to 315 nm) generally plays a negative role in plant growth and development by causing oxidative damage and cross-links in plant cells1. Lesions form in nuclear, chloroplast, and mitochondrial DNA due to the accumulation of cyclobutane pyrimidine dimers (CPDs) and pyrimidine (6-4) pyrimidone photoproducts (6-4 PPs) on the DNA strands, which have been recognized as two major lesions on DNA2,3.
To maintain genomic integrity, repairing DNA damage is essential for an organism to survive4. Studies show that plant photoreactivation plays an essential role in repairing the CPD damage caused by UVB radiation, and that photolyase participates in this pathway by absorbing blue/UVA (320 to 400 nm) light and using the energy to monomerize the dimers5 in a process known as the photorepair pathway. In addition to photorepair, dark repair occurs in plants, including excision repair for bases or nucleotides, mismatch repair, and other DNA repair pathways. Photorepair and excision repair are both important for maintaining the stability of the genome and are essential for an organism’s survival. DNA damage repair and its relative photolyases have been extensively reported in several plant species6,7,8,9,10.
There are four genes that encode photolyases in rice, namely Os10g0167600, Os02g0204400, Os03g0343400, and Os09g0532700, and three genes that encode the photoreceptor, namely, Os02g0625000, Os02g0573200, and Os06g0661800. These genes are located in different parts of the chromosome and their expression can be activated and increased in the presence of UVA and blue light to increase photoreactivation for repairing DNA damage11,12. Overexpression of CPD photolyase in both UVB-sensitive and UVB-hypersensitive rice contributes to increased CPD photolyase activity, which increases the plants’ resistance to growth damage from UVB compared with wild-type (WT) plants13. The activity of CPD photolyase in rice can be regarded as a symbolic factor for evaluating the sensitivity of rice to UVB radiation13,14.
Beneficial elements such as silicon have been reported to contribute to the enhancement of UVB resistance in rice15,16. The ability of rice to absorb silicon from the environment is controlled by the gene that encodes the NOD26-like major intrinsic protein; Ma et al. found the gene in a low-silicon rice mutant and named it Lsi117. Fang et al. inhibited and overexpressed Lsi1 on rice to generate two types of rice with different silicon contents, and compared their gene expression profiles after exposure to UVB. The CPD photolyase gene expression was up-regulated in the transformed rice line with Lsi1-overexpression (Lsi1-OX) but down-regulated in the Lsi1-RNAi line, and these results were comparable to those for the WT. It has been suggested that the CPD photolyase gene expression was correlated with the silicon content in rice and that silicon could activate the expression of photolyase18. However, the underlying mechanism is still unknown.
We conducted a comparative study of the expression of photolyase encoded genes to select the responsible gene in types of rice with different silicon contents. The most correlative photolyase genes triggered by silicon were obtained and the positive trans-acting factor interaction with the cis-acting element of the responsible photolyase gene was revealed. The bio-interaction of the trans-acting factor with silicon and photolyase was then further investigated to illuminate the regulation pathway for repairing the CPDs on rice DNA.
Results
Silicon, CPD, and 6-4 PPs contents in the Lemont rice accession with Lsi1-OX, Lsi1-RNAi, and WT
Overexpression of Lsi1 in the Lemont rice accession (Lsi1-OX) resulted in increased silicon content in the leaves, and the silicon content in the Lsi1-RNAi transformed Lemont was notably lower than that in the WT (Fig. 1A). Exposing the rice to UVB radiation led to increased CPD and 6-4 PP contents in the leaf DNA. There was no obvious difference in the 6-4 PPs contents between the transformed rice and the WT under the same light conditions (Fig. 1B). In contrast, the CPD content was significantly lower in the leaves of the Lsi1-OX line than in the WT after exposure to UVB radiation. A 35.17% decrease was found in the Lsi1-OX line compared with the WT. However, a notable increase of 33.16% was found in the Lsi1-RNAi line compared with the WT (Fig. 1C). These results indicate that the increased Lsi1 expression in the rice enhanced the silicon content in the leaves and reduced the CPD content in the DNA of the leaves under UVB irradiation, while inhibition of Lsi1 expression in the rice samples led to the opposite result.
(A) The silicon content in the leaves of the three test rice lines under normal growth conditions; (B) the 6-4 PP content in the leaves of the three test rice lines for the UVB-treated group and the control group; (C) the CPD content in the leaves of the three test rice lines for the UVB-treated group and the control group. WT, wild type of Lemont rice accession; Lsi1-OX, Lsi1 overexpression transformed Lemont; Lsi1-RNAi, Lsi1 RNA interference transformed Lemont. Data are represented as the mean ± SEM. The different superscripts in the columns indicate statistical groups that are significantly different (p < 0.05) using Tukey’s range test.
Photolyase gene expression and the photoreceptor in the Lemont rice accession with Lsi1-OX, Lsi1-RNAi, and WT
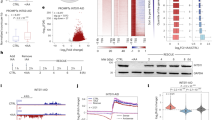
Based on our transcriptome sequencing data from the Lsi1-OX and Lsi1-RNAi transformed Lemont rice lines and the WT rice, which were irradiated with UVB for 5h, and the control group (full data not shown here), we found seven genes that encoded photolyase or the photoreceptor, and which were differentially expressed in the three UVB-treated rice lines and the control condition. Among these genes, the transcription of Os10g0167600 was up-regulated when the rice samples were exposed to UVB. Changes of 1.51, 2.59, and 1.57 times were detected in the WT, Lsi1-OX, and Lsi1-RNAi lines, respectively, when comparing the UVB group to the control group. The gene encodes deoxyribodipyrimidine photolyase to repair the CPDs on the UVB-damaged DNA. Similarly, up-regulation of the Os03g0343400 gene expression increased by 1.57, 1.12, and 1.31 times in the three rice lines, and the gene encoded a putative photolyase/blue-light receptor PHR2. In contrast, the transcript abundance of Os09g0532700, which is encoded as the deoxyribodipyrimidine photolyase protein-like family, was down-regulated in all three UVB-treated rice lines. Os02g0204400 gene expression was slightly down-regulated in the UVB-treated WT and Lsi1-RNAi transformed lines, but it was obviously up-regulated in the Lsi1-OX transformed line. Os02g0204400 encodes the (6-4) DNA photolyase that catalyzes the photoreactivation of the 6-4 PPs.
The other three genes also exhibited changes in their levels of expression. Os06g0661800 was notably up-regulated in all three rice lines, as it is encoded as cryptochrome with blue-light photoreceptor activity but lacks photolyase activity. In contrast, the transcription levels of Os02g0625000 and Os02g0573200 were down-regulated in the UVB-treated WT and Lsi1-OX lines, but were slightly up-regulated in the UVB-treated Lsi1-RNAi rice (Fig. 2). Comparison of the changes in the transcript levels of the genes suggests that Os10g0167600 is the most correlative to silicon in rice and plays a role in the photoactivation of CPDs on damaged DNA.
The transcriptional abundance of the seven genes was calculated as the reads per kilobase per million reads (RPKM), which is a method of quantifying gene expression from RNA sequencing data by normalizing the total read length and the number of sequencing reads. The RNA sequencing data were generated from the mRNA of the UVB-treated Lsi1-OX, -RNAi, and WT lines, respectively, and the control groups. The full RNA sequencing data are not shown here.
Gene dynamic expression of Os10g0167600 in the Lsi1-OX, -RNAi transformed Lemont, and WT lines
qPCR was used to detect the gene dynamic expression of Os10g0167600 in the leaves of rice exposed to UVB irradiation for 0.5, 1, 2, 3, and 5 hours. Compared with the expression of Os10g0167600 in the rice samples before UVB treatment, UVB irradiation resulted in the up-regulation of Os10g0167600 gene expression in the Lsi1-OX transformed and WT lines, and the level of up-regulation increased with time. The increase in the transcription level was higher in the Lsi1-OX transformed line than in the WT and peaked at 5 h of irradiation, at which point there was a 2.02 times increase in the UVB-irradiated Lsi1-OX transformed line compared to its control before treatment. However, no obvious change in Os10g0167600 gene expression was found in the Lsi1-RNAi transformed line after UVB treatment (Fig. 3A). Compared with the control group under the same exposure time, the gene expression was also found to be up-regulated in the UVB-treated group, with a 2.95 times increase in up-regulation in the UVB-treated Lsi1-OX transformed line (Fig. 3B). The dynamic change in Os10g0167600 gene expression detected by qPCR confirmed the results of the RNA-seq. Based on these results, Os10g0167600 was selected for further study.
(A) Gene dynamic expression of Os10g0167600 in the rice lines treated by UVB radiation for 0.5, 1, 2, 3, and 5 h compared to that of 0 h (before UVB treatment). (B) Gene dynamic expression of Os10g0167600 in the rice samples treated by UVB radiation for 0.5, 1, 2, 3, and 5 h compared to the control group with the same levels of irradiation. The gene expression level of Os10g0167600 before UVB treatment (0 h) was normalized as a value of 1. Relative mRNA expression level >1 and expression level <1 indicate the comparative up-regulation and down-regulation, respectively, of the gene expression in the UVB treatment and control samples. Data are represented as the mean ± SEM. Asterisks on the columns of 5 h UV-B treated Lsi1-OX indicate groups that are statistically significantly different (p < 0.05) using Tukey’s range test.
Promoter of Os10g0167600 and the interacted trans-acting factor
A 2154-bp fragment from the upstream region of the 5′ UTR of Os10g0167600 was amplified from Lemont (Fig. 4A) and then incubated with the natural protein (Fig. 4B) to reveal the interacting trans-acting factor according to the DNA pull-down method (Fig. 4C). Based on the results, an obvious protein band was obtained with SDS-PAGE separation. Nine proteins (see Table S2) were identified by LC-MS. Methyl-CpG binding domain containing protein (OsMeCP) was regarded as the putative interacting trans-acting factor (Table 1).
The biotin-labeled DNA fragment of the promoter of Os10g0167600 was mixed with the leaf total natural protein to catch the interacted proteins. The mixture was immobilized with streptavidin-coupled beads and then eluted with 1 M NaCl and forwarded to SDS-PAGE. (A) gene fragment of the promoter of Os10g0167600, M, 100 bp DNA ladder, lane 1, promoter of Os10g0167600; (B) the leaf total natural proteins of Lemont, M, protein ladder, lane 1, leaf total natural proteins; (C) the interacted protein on the promoter of Os10g0167600. M, protein ladder, lane 1 shows the proteins interacting with the DNA fragment of the promoter of Os10g0167600, lane 2 shows the negative control in which the DNA fragment is mixed with the protein extracting solution.
Subcellular localization of OsMeCP in rice
It is well known that gene regulation usually occurs in the nucleus and that the trans-acting factor acts in the nucleus. As indicated by the nuclear marker tag (Fig. 5A), the nucleic subcellular localization of OsMeCP also supports this deduction. The laser confocal imaging revealed OsMeCP with high fluorescence in the nucleus (Fig. 5B–D).
The fluorescence images were acquired at 568-nm excitation and 592-nm emission using an Olympus Fluoview FV1000 laser scanning confocal microscope. Image (A) is a monomeric far-red fluorescent protein TagFP635 (scientific name mKate) that tagged the nuclear localization signal protein localized to the nucleus, and was taken as the standard marker to indicate the location of the nucleus in the rice protoplast. Image (B) is a fluorescent field visualization of OsMeCP and image (C) is the bright field visualization of the rice protoplast. Image (D) is the overlay of images (B,C) to clearly show the OsMeCP subcellular localization.
Gene transcript levels of OsMeCP in the Lsi1-OX, -RNAi transformed Lemont, and WT lines
The gene transcription levels of OsMeCP were then detected in the three rice samples. OsMeCP was down-regulated in the Lsi1-OX rice compared with the WT, but was up-regulated in the Lsi1-RNAi line (Fig. 6A), suggesting that the silicon in the rice negatively regulated OsMeCP gene expression. Similarly, Os10g0167600 gene expression was down-regulated in the Lsi1-OX rice and slightly up-regulated in the Lsi1-RNAi line, compared with the WT (Fig. 6B). This can be attributed to the low CPD content in the rice DNA under normal light condition (Fig. 1C). When the rice samples were exposed to UVB radiation, the Lsi1-OX rice still showed down-regulation of OsMeCP gene expression compared with the WT, and the reverse was true in the Lsi1-RNAi line (Fig. 6C). However, in contrast to OsMeCP gene expression in the rice samples, Os10g0167600 gene expression was found to be up-regulated in the Lsi1-OX line but down-regulated in the Lsi1-RNAi line compared with the WT (Fig. 6D). Thus, it can be deduced that OsMeCP may have transcriptionally repressed Os10g0167600 gene expression in the rice under UVB radiation.
(A) mRNA expression of OsMeCP in the Lsi1-OX and -RNAi transformed Lemont lines compared to the WT under normal growth conditions. (B) mRNA expression of Os10g0167600 in the Lsi1-OX and -RNAi transformed Lemont lines compared to the WT under normal growth conditions. (C) mRNA expression of OsMeCP in the Lsi1-OX and -RNAi transformed Lemont lines compared to the WT under UVB radiation for 5 h. (D) mRNA expression of Os10g0167600 in the Lsi1-OX and -RNAi transformed Lemont lines compared to the WT under UVB radiation for 5 h. The gene expression levels in the WT were normalized as a value of 1. Expression level >1 and expression level <1 indicate up-regulation and down-regulation of gene expression in Lsi1-OX or -RNAi transformed rice compared to the WT under normal growth conditions and UVB radiation, respectively.
Protein expression of OsMeCP in the Lsi1-OX, -RNAi transformed Lemont, and WT lines
The protein expression levels of OsMeCP were also detected in the transformed and WT rice lines under UVB radiation. It was found that the protein expression obviously increased in the rice exposed to UVB irradiation. The overexpression of Lsi1 inhibited the expression of OsMeCP, and the reverse was true in the Lsi1-RNAi line from the control group and UVB-treated group conditions (Fig. 7). These results further confirmed that the silicon in the rice played a negative role in the regulation of OsMeCP protein expression.
“Control” indicates the rice in the control condition, with UVB and UVC eliminated by wrapping mylar film on the UV lamps; “UVB” indicates that lamps with a 0.1-mm film of cellulose diacetate were used as the source of UVB radiation. The lamps were applied at the top of the rice leaf canopy for 5 h.
OsMeCP expression and CPD content in rice cultured with different silicon concentrations
Rice cultured in different silicate concentrations of the hydroponic solution showed different levels of OsMeCP. Increasing the silicate concentration in the solution resulted in reduced protein expression levels of OsMeCP in the rice. The protein expression level of OsMeCP was increased in the UVB-treated rice compared with the control group, where the two rice samples were under the same silicate concentration. Moreover, the protein level of OsMeCP was still the highest in the rice cultured in the medium containing 0.5 mM silicate, and lowest in that containing 2.5 mM of silicate (Fig. 8A). The gene expression level of Os10g0167600 decreased slightly less in the UVB normal condition than in the rice cultured in the increasing silicate concentration. This may have been because the rice under the normal growth conditions had little DNA damage, as indicated by the lower CPD content on the rice leaves. When the three different cultured strains of rice were exposed to UVB radiation, the gene transcription of Os10g0167600 increased in all cases and was highest in the 2.5 mM silicate cultured rice (Fig. 8B). Moreover, the CPD content was highest in the 0.5 mM silicate cultured rice, significantly decreased in the rice cultured with the hydroponic nutrient solution that contained 1.5 mM silicate, and was notably lowest in the 2.5 mM condition. Detection of the CPD content in the leaves of rice cultured in the different silicate concentrations showed that increasing the silicate concentration in the solution obviously repressed OsMeCP protein expression, and resulted in enhanced Os10g0167600 gene expression in the UV-B treated rice, which strengthened the photoreactivation and significantly reduced the CPD content in the cultured rice leaves (Fig. 8C).
(A) OsMeCP protein expression levels in the UVB-treated rice cultured in hydroponic solutions containing 0.5 mM, 1.5 mM, and 2.5 mM of silicon, respectively, compared with the controls. (B) Os10g0167600 gene expression levels in the UVB-treated rice samples cultured in hydroponic solutions containing 0.5 mM, 1.5 mM, and 2.5 mM silicon, respectively, compared with the controls. Os10g0167600 gene expression levels in the WT control rice samples in hydroponic solution containing 0.5 mM, 1.5 mM, and 2.5 mM of silicon, normalized as a value of 1 to calculate the relative changes in expression compared with the UVB radiation condition. Expression level >1 and expression level <1 indicate up-regulation and down-regulation of the gene expression in the UVB treated group compared with the control group, respectively. (C) The CPD content in the UVB-treated rice samples compared with the controls. The different superscripts in the columns indicate statistical groups that are significantly different (p < 0.05) using Tukey’s range test.
OsMeCP-OX and -RNAi transformed rice and DNA damage
The respective OsMeCP-OX and -RNAi transformed Lemont rice lines were generated. The protein expression level of OsMeCP in the overexpression transformed line was found to be greater than in the WT, and it was obviously reduced in the OsMeCP-RNAi line (Fig. 9A). Comparison of the changes in Os10g0167600 gene expression in the two transformed rice lines and the WT line under normal growth conditions showed that Os10g0167600 gene expression was up-regulated in the OsMeCP-RNAi line but down-regulated in the OsMeCP-OX line compared with the WT (Fig. 9B). When the three lines were exposed to UVB radiation, both OsMeCP-RNAi and the WT showed up-regulation of Os10g0167600 gene expression compared with the control group, and the increase in the up-regulation was higher in the OsMeCP-RNAi transformed line than in the WT. In contrast, a slight down-regulation of Os10g0167600 gene expression was found in the OsMeCP-OX transformed line (Fig. 9C). The highest up-regulation of Os10g0167600 gene transcription in the OsMeCP-RNAi transformed line led to the lowest CPD content in the UVB-treated leaves, compared with that of the WT and the OsMeCP-OX transformed line. Moreover, the OsMeCP-OX transformed line had the highest CPD content among the three rice lines (Fig. 9D), owing to the slightly higher down-regulation of Os10g0167600 than in the WT. These results further confirmed the transcriptional repression of OsMeCP on Os10g0167600 gene expression under UVB radiation, which resulted in reduced photolyase activity in the UVB-exposed rice.
(A) OsMeCP protein expression in the OsMeCP-OX, -RNAi transformed Lemont, and WT lines detected by Western blot. (B) mRNA expression of Os10g0167600 in the OsMeCP-OX and -RNAi transformed Lemont lines compared with the WT under normal growth conditions. The gene expression level of Os10g0167600 in the WT was normalized as a value of 1. Expression level >1 and expression level <1 indicate up-regulation and down-regulation of the gene expression in the transformed line compared to the WT, respectively. (C) mRNA expression level of Os10g0167600 in the OsMeCP-OX and -RNAi transformed Lemont lines under UVB radiation compared with the control groups. The gene expression level of Os10g0167600 in the control groups was normalized to a value of 1. Expression level >1 and expression level <1 indicate up-regulation and down-regulation of the gene expression in the UVB treated group compared with the control group, respectively. (D) CPD content in the three UVB-treated rice lines and their control groups. The different superscript letters in the columns indicate groups that are statistically significantly different (p < 0.05) in terms of the leaf CPD content of the OsMeCP-OX. -RNAi, and WT lines, as indicated by Tukey’s range test. OsMeCP-OX, OsMeCP overexpression transformed Lemont. OsMeCP-RNAi, OsMeCP RNA interference transformed Lemont.
Discussion
UVB radiation causes the formation of covalent links by reactions localized on the C=C double bonds, which results in the accumulation of CPDs and 6-4PPs in DNA19. To reduce the damage, the photoreactivation of the CPD photolyase repairs the CPD in the plant chloroplasts, mitochondria, and nuclei20, which is essential for plant survival after exposure to UVB-containing sunlight.
In rice plants, Os10g0167600 encodes the deoxyribodipyrimidine photolyase, which is involved in the repair of UV radiation-induced DNA damage and catalyzes the light-dependent monomerization (300–600 nm) of CPDs, which is required for plant survival in the presence of UVB light. However, photolyase is not involved in the repair of 6-4 PPs21,22. In contrast to Os10g0167600, Os02g0204400 encodes the (6-4) DNA photolyase that catalyzes the photoreactivation of 6-4 PPs. The Os03g0343400 protein is a putative photolyase/blue-light receptor PHR2, and Os09g0532700 is deoxyribodipyrimidine photolyase family protein-like, and both have hydrolase activity. Unlike the photolyase, Os02g0625000, Os02g0573200, and Os06g0661800 encode cryptochromes with blue-light photoreceptor activity, but there is a lack of photolyase activity.
Although the transcript levels of these seven genes were detectable, we observed different expression levels in the three rice lines with different levels of Lsi1 abundance. Among these genes, the transcription of Os10g0167600 was correlated with the silicon content in the rice, and the highest increase was found in the Lsi1-OX transformed rice under UVB radiation. The resulting CPD content was negatively related to the gene abundance, and were lowest in the Lsi1-OX transformed rice. Such a reduction in the DNA lesion would maintain the UVB resistance of rice. Recently, Teranishi et al.13 found that CPD photolyase was overexpressed in both UVB-sensitive O. sativa Norin 1 (japonica), and UVB-hypersensitive O. sativa Surjamkhi (indica) when determining its function in relation to UVB resistance in rice. Their results showed that CPD photolyase is a vital factor for evaluating the UVB sensitivity of O. sativa13,14. Similar results were found in a study on Arabidopsis thaliana23,24.
Gene expression can be regulated by trans-acting factors, which are known as transcription factors. These regulators interact with the cis-regulatory elements to activate gene transcription. In our study, the Os10g0167600 promoter was associated with the CpG islands, and OsMeCP was found to interact with the promoter to regulate the gene transcription. Protein subcellular locations are closely linked to their biological functions, and, in this study, the subcellular location of OsMeCP was found to be in the nucleus, which confirms its role in the regulation of transcription. A decrease in the protein abundance of OsMeCP in the rice cultured under the higher silicon condition indicates that silicon acts as a negative factor in the regulation of OsMeCP expression. The methyl-CpG-binding domain, which consists of about 70 residues, possesses a unique α/β-sandwich structure with characteristic loops, and is able to bind single methylated CpG pairs as a monomer, which is involved in recruiting histone deacetylases to methyl CpG-enriched regions in the genome to repress transcription25. We found that rice with a higher silicon content exhibited lower protein abundance of OsMeCP in both the Lsi1-OX transformed line and the WT cultured in high silicate-concentration solutions. This suggests that OsMeCP is a negative feedback of silicon. However, the changes in the gene transcription levels of Os10g0167600 in the Lsi1-OX and Lsi1-RNAi transformed lines compared with the WT, and in the rice cultured in different silicon-concentration solutions, were opposite to the OsMeCP expression level in the same rice. This finding indicates the repression of OsMeCP in the transcription of Os10g0167600 in the UVB-treated rice. Nan et al.26 documented that an abundant nuclear protein, the methyl-CpG-binding protein MeCP2, interacts specifically with methylated DNA and mediates the transcriptional repression. MeCP2 binds tightly to the chromosomes in a methylation-dependent manner and contains a transcriptional repression domain that can function at a distance in vitro and in vivo.
Overexpression of OsMeCP in rice also resulted in the down-regulation of Os10g0167600 gene expression, while the gene transcription level of Os10g0167600 was up-regulated in the OsMeCP-RNAi line. These results further clarify that OsMeCP acts as a negative regulatory factor in the transcription of Os10g0167600. The rice exposed to UVB radiation showed up-regulation of Os10g0167600 gene expression, but it was still highest in the OsMeCP-RNAi line, which led to the lowest amount of CPDs on the DNA. In contrast, the highest CPD content was found in the OsMeCP-OX transformed line, which was attributable to the slight down-regulation of Os10g0167600 gene expression under UVB treatment compared with the control group. Thus, the transcriptional repression on Os10g0167600 was obviously mediated by OsMeCP.
In conclusion, increasing the exogenous silicon concentration reduced the protein expression of OsMeCP in rice, and the silicon content in rice negatively regulated the protein expression of OsMeCP. The protein abundance of OsMeCP was also lower in the Lsi1-OX transformed line than in the WT rice, whereas the reverse was true for the Lsi1-RNAi line. OsMeCP repressed the transcription of Os10g0167600 gene expression, leading to down-regulation of Os10g0167600 in the rice samples. Inhibition of OsMeCP increased Os10g016760 gene expression, which reduced the CPD content in the rice under UVB radiation, while overexpression of OsMeCP led to the reverse results. Overall, our results indicate that OsMeCP is a negative regulator of silicon uptake in rice and plays a role in repressing Os10g0167600 gene transcription, which dominates the photorepair of damaged DNA in rice following UVB radiation (Fig. 10).
Materials and Methods
Plant materials
The Lemont rice accession (UVB-tolerant; introduced from the United States) was used in this study. The Lsi1-OX and Lsi1-RNAi transformed lines, which were generated in our previous study, had significantly higher or lower levels of silicon than the WT18.
The WT and transformed rice seeds were surface-sterilized with 25% NaClO for 30 min and then soaked in sterilized ddH2O overnight. The seeds were then placed in a temperature-controlled incubator to germinate at 30 °C, and the germinated seeds were sown in separate seedling plates. At the three-leaf stage, uniform seedlings from each plant were selected and transferred onto a Styrofoam plate (with holes spaced at 5 × 6 cm). The seedlings were affixed to the plate by inserting a cotton plug into each hole. The Styrofoam plate was allowed to float in a pot (45 × 35 × 15 cm) filled with 10 L of rice (Oryza sativa L.) culture solution (silicate-containing solution) with the following composition: 482 mg (NH4)2SO4, 248 mg KH2PO4, 185 mg KNO3, 149 mg K2SO4, 864.3 mg Ca(NO3)2.4H2O, 1350.6 mg MgSO4.7H2O, 2000 mg Na2SiO3.9H2O, 457 mg FeSO4.7H2O, 484.4 mg EDTA, 14.3 mg H3BO3, 0.4 mg CuSO4.5H2O, 1.1 mg ZnSO4.7H2O, 9.05 mg MnCl2.4H2O, and 0.45 mg Na2MoO4.2H2O. The solution was changed every week. The pot was continuously aerated, and its outer side was painted black to prevent algae growth. The pH value of the solution was maintained between 5.5 and 6.0 throughout the experiment. The method follows that of Fang et al.18, which is a minor optimization of the technique of Yoshida et al.27.
At the five-leaf stage, the transformed rice line and WT rice seedlings were exposed to enhanced UVB radiation following the method described by Fang et al.18,28. The procedure was as follows. Fluorescent lamps (40 W, Beijing Electric Light Sources Research Institute, China) were used as the source of UVB radiation. The lamps were suspended above the rice plants, and UV radiation of wavelengths less than 280 nm (i.e., UVC) was eliminated by wrapping the lamps in a 0.1-mm film of cellulose diacetate (West Design Product Co., Ltd, United Kingdom). The distance between the rice canopy and the lamps was 30 cm, and four lamps were used. UVB radiation was applied at the top of the leaf canopy from 10 am to 3 pm at an intensity of 18.6 kJ • m−2 • d−1. The rice seedlings used as a control were wrapped in a mylar film to filter UVB and UVC and placed under the lamps. The UVB-exposed and control rice leaves were sampled at 0.5, 1, 2, 3, and 5 h, and were immediately frozen in liquid nitrogen and stored at −80 °C for isolation of the total RNA to detect Os10g0167600 gene expression. The sample of leaves treated for 5 h was also used to determine the silicon content and extract natural protein for the DNA-pull down.
Detection of the rice silicon content
The fresh rice leaves were ground into powder in liquid nitrogen and oven-dried at 70 °C. Each 0.1 g portion of the dry powder was digested in 3 ml of 50% NaOH (W/V) in an autoclave sterilizer at 121 °C for 30 min. The mixture was centrifuged, and the supernatant liquor was diluted with ddH2O to 50 mL. One milliliter of the solution was added to 30 ml of 20% acetic acid (V/V) and 10 ml of ammonium molybdate solution (54 g/L; pH 7.0). The mixture was blended and allowed to sit for 5 min; 5 ml of 25% tartaric acid solution (W/V) and 1 ml of reducing agent was then added, and the mixture was finally diluted with 20% acetic acid to 50 ml. To prepare the reducing agent, 25 ml of ddH2O was dissolved with 2 g of Na2SO3 and 0.4 g of 1-amino-2-naphthol-4-sulfonic acid, and another 200 ml of ddH2O with 25 g NaHSO3 was combined and diluted with ddH2O to 250 mL.
The absorbance of the rice leaf solution was detected with a spectrophotometer at a wavelength of 650 nm. For the standard curve of the silicon content, a gradient of Na2SiO3 concentrations from 0, 1, 2, 3, 4, 5, 6, 7, 8, 9 mg/ml was taken to conduct the same reaction and then detected at OD260.
Detection of CPDs and 6-4 PPs in transformed and WT rice
The DNA of the Lsi1-OX and Lsi1-RNAi transformed lines and the WT rice leaves were extracted with a NuClean PlantGen DNA Kit (Beijing ComWin Biotech Co. Ltd.), and the CPDs and 6-4 PPs on the DNA were detected with an OxiSelect™ UV-Induced DNA Damage ELISA Kit (CPD and 6-4 PP quantitation, respectively; Cell Biolabs, Inc., USA) following the manufacturers’ instructions.
Os10g0167600 gene expression in transformed and WT rice
The transcriptional abundance of the seven genes that encode photolyase or the photoreceptor was compared with our RNA-seq data to select the gene with the highest correlation with silicon. To detect the target photolyase gene expression in the UVB-irradiated Lsi1-OX and Lsi1-RNAi transformed lines and the WT, we designed gene-specific primers for Os10g0167600 (see Table S1), and used a real-time quantitative polymerase chain reaction (qPCR) to detect the relative mRNA expression levels of the genes in the UVB-treated group and the control group. The reaction was performed in an Eppendorf realplex4 cycle, and the threshold cycle values (Ct) were recorded for each candidate mRNA in both the control and test samples. The 2−△△Ct method was used to analyze the relative quantification of the gene expression29, and the gene expression level of β-actin was taken as a reference to normalize the candidate genes in the test sample.
Cloning of the specific photolyase gene promoter and pulling the interacted proteins
Based on the comparative study of photolyase gene expression in the UVB-treated Lsi1-OX and -RNAi lines and the WT, Os10g0167600 was selected as the available gene that was correlated with the silicon content of the rice strains and enhanced their capacity to repair the UVB damage on DNA. A 2134-bp upstream fragment of the 5′ end of the ORF of the gene was cloned and labeled with biotin (see Table S1). The natural proteins of the Lemont rice leaf were extracted with phosphate-buffered saline solution. The purified DNA fragment was mixed with the protein to reveal the interacted proteins. The DNA fragment was immobilized with streptavidin-coupled beads from the Dynabeads® kilobaseBINDERTM Kit (Thermo Fisher Scientific Inc., USA), and the interacted proteins on the DNA fragment were then eluted with 1 M NaCl and forwarded to sodium dodecyl sulfate-polyacrylamide gel electrophoresis (SDS-PAGE). Liquid chromatography-mass spectrometry (LC-MS; LTQ, Thermo Scientific) was performed to identify the interacted proteins.
Gene transcript levels of the DNA interacted proteins
A protein that contains the DNA binding domain, methyl-CpG binding domain protein (OsMeCP), was selected to detect the transcription levels in the Lsi1-OX, -RNAi transformed, and WT rice lines under UVB radiation for 5 h and the control. The specific primers for the encoding genes were as follows: OsMeCP-F: 5′-CTC ATC TGG CAG AAA GGG-3′ and OsMeCP-R: 5′-GGA AGG CTC AGT TGG GTT-3′. qPCR was used to detect the gene expression levels in the test rice.
Protein expression detected by Western blot
To investigate the protein expression levels of OsMeCP in the different UVB-treated rice lines and the control we used monoclonal antibodies of OsMeCP (Abmart Inc., USA). The leaf proteins of the three UVB-treated rice samples and the control were extracted and separated by SDS-PAGE, and the monoclonal antibodies were found to be immune to the total protein. A Clarity Western ECL Kit (Bio-Rad Laboratories, Inc., USA) was used for the staining, and the protein expression level was checked with the ChemiDoc™ MP System (Bio-Rad Laboratories) under chemiluminescence.
OsMeCP protein expression levels in rice cultured in different silicon concentrations of hydroponic solution
To detect the OsMeCP protein abundance, samples of the Lsi1-OX transformed Lemont rice were cultured in Yoshida culture solution with different silicate concentrations: 0.5 mM, 1.5 mM, and 2.5 mM. The rice under the three silicon concentrations was exposed to UVB radiation at the five-leaf stage, and rice wrapped in mylar film before being placed under the lamps was again used as the control. After 5 h, the respective rice leaves were sampled to detect OsMeCP protein abundance, Os10g0167600 gene expression, and the CPD content.
Protein subcellular localization of OsMeCP
The protoplast of rice was prepared to study the protein location. The full-length CDS of OsMeCP was amplified with gene-specific forward primer location CPG-F: 5′-ATC GC T CTA GA A TGG CCA CGG CCG GCG ACG A-3′ and location CPG-R: 5′-TAT AGG ACT AGT GGT GCA CTT CAC GGC AGA GG-3′ containing Xba I and Spe I sites (underlined). The gene was inserted into a plant binary expression vector, pCambia-eYFP-2300s, to create a recombinant plant transient expression vector and then transformed into the rice protoplast. The cell was cultured for two days to completely express the protein, and an Olympus Fluoview FV1000 laser scanning confocal microscope (Olympus, Tokyo, Japan) was used to observe the protein subcellular localization. The fluorescence images were acquired at 568-nm excitation and 592-nm emission. A monomeric far-red fluorescent protein TagFP635 (scientific name mKate) that tagged the nuclear localization signal protein localized to the nucleus was taken as the standard marker.
Overexpression and RNA interference of OsMeCP in rice
To further confirm the gene function of OsMeCP, the full-length CDS of the gene was again amplified with primers containing different recognition sites for Sac I and Bam H I, respectively, in forward primer OsMeCP-OX-F: 5′-TAT AC G AGC TC A TCC CCA AAT CCC CAC ACG TC-3′ and reverse primer OsMeCP-OX-R: 5′-TAG CG G GAT C CC CAG CAA CGT CAG TTC CTT GG-3′. The gene was then inserted into another plant binary expression vector, pCambia-1300. A 347 base pair (bp) partial coding region downstream of the ATG of rice OsMeCP was cloned with a forward primer 5′-CG GGATCC AAATGAAGAAGC GAAAGACG-3′ and a reverse primer 5′-GG GGTACC AAGGCTCAGTTGGGTTGC-3′ containing BamH I and Kpn I sites (underlined), respectively. The same fragment was again amplified with the primers containing different recognition sites for Sac I and Spe I, respectively, in forward primer 5′-C GAGCTC AAATGAAGAAGCGAAAGACG-3′ and reverse primer 5′-GG ACTAGT AAG GCTCAGTTGGGTTGC-3′. Both fragments were inserted into pTCK303 to create an OsMeCP-RNAi stability vector. The two recombinant vectors were then transformed into the Agrobacterium strain EHA105, and the process of rice genetic transformation followed the methods described by Fang et al.18. After the positively transformed rice was confirmed and selected, the transformed rice and the WT were exposed to UVB radiation for 5 h, and the second top leaves of the rice plants were sampled to extract DNA and RNA, respectively, to detect the CPD contents and Os10g0167600 gene expression in the different types of rice under UVB irradiation.
Additional Information
How to cite this article: Fang, C. et al. Methyl-CpG binding domain protein acts to regulate the repair of cyclobutane pyrimidine dimers on rice DNA. Sci. Rep. 6, 34569; doi: 10.1038/srep34569 (2016).
References
Nawkar, G. M. et al. UV-induced cell death in plants. Int. J. Mol. Sci. 14, 1608–1628 (2013).
Bray, C. M. & West, C. E. DNA repair mechanisms in plants: crucial sensors and effectors for the maintenance of genome integrity. New Phytol. 168, 511–528 (2005).
Sinha, R. P. & Häder, D. P. UV-induced DNA damage and repair: a review. Photoch. Photobio. Sci. 1, 225–236 (2012).
Landry, L. G. et al. An Arabidopsis photolyase mutant is hypersensitive to ultraviolet-B radiation. Proc. Natl. Acad. Sci. USA 94, 328–332 (1997).
Sancar, A. Structure and function of DNA photolyase and cryptochrome blue-light photoreceptors. Chem. Rev. 103, 2203–2237 (2003).
Gao, L. et al. The tomato DDI2, a PCNA ortholog, associating with DDB1-CUL4 complex is required for UV-damaged DNA repair and plant tolerance to UV stress. Plant Sci. 235, 101–110 (2015).
Fujimori, N., Suzuki, N., Nakajima, Y. & Suzuki, S. Plant DNA-damage repair/toleration 100 protein repairs UV-B-induced DNA damage. DNA repair 21, 171–176 (2014).
Kimura, S. et al. DNA repair in higher plants; photoreactivation is the major DNA repair pathway in non-proliferating cells while excision repair (nucleotide excision repair and base excision repair) is active in proliferating cells. Nucleic Acids Res. 32, 2760–2767 (2004).
Liu, Z., Hossain, G. S., Islas-Osuna, M. A., Mitchell, D. L. & Mount, D. W. Repair of UV damage in plants by nucleotide excision repair: Arabidopsis UVH1 DNA repair gene is a homolog of Saccharomyces cerevisiae Rad1. Plant J. 21, 519–528 (2000).
Stapleton, A. E., Thornber, C. S. & Walbot, V. UV-B component of sunlight causes measurable damage in field-grown maize (Zea mays L.): developmental and cellular heterogeneity of damage and repair. Plant Cell Environ. 20, 279–290 (1997).
Carell, T., Burgdorf, L. T., Kundu, L. M. & Cichon, M. The mechanism of action of DNA photolyases. Curr. Opin. Chem.Biol. 5, 491–498 (2001).
Jansen, M. A. K., Gaba, V. & Greenberg, B. M. Higher plants and UV-B radiation: balancing damage, repair and acclimation. Trends Plant Sci. 3, 131–135 (1998).
Teranishi, M., Taguchi, T., Ono, T. & Hidema, J. Augmentation of CPD photolyase activity in japonica and indica rice increases their UVB resistance but still leaves the difference in their sensitivities. Photoch. Photobio. Sci. 11, 812–820 (2012).
Hidema, J. et al. Increase in CPD photolyase activity functions effectively to prevent growth inhibition caused by UVB radiation. Plant J. 50, 70–79 (2007).
Li, W. B., Shi, X. H., Wang, H. & Zhang, F. S. Effects of silicon on rice leaves resistance to ultraviolet-B. Acta Bot. Sin. 46, 691–697 (2004).
Goto, M. et al. Protective effect of silicon on phenolic biosynthesis and ultraviolet spectral stress in rice crop. Plant Sci. 164, 349–356 (2003).
Ma, J. F. et al. A silicon transporter in rice. Nature 440, 688–691 (2006).
Fang, C. X. et al. Suppression and overexpression of Lsi1 induce differential gene expression in rice under ultraviolet radiation. Plant Growth Regul. 65, 1–10 (2011).
Whitmore, S. E., Potten, C. S., Chadwick, C. A., Strickland, P. T. & Morison, W. L. Effect of photoreactivating light on UV radiation-induced alterations in human skin. Photodermatol. Photo. 17, 213–217 (2001).
Takahashi, M. et al. Cyclobutane pyrimidine dimer (CPD) photolyase repairs ultraviolet-B-induced CPDs in rice chloroplast and mitochondrial DNA. Plant J. 66, 433–442 (2011).
Hirouchi, T. et al. A gene for a Class II DNA photolyase from Oryza sativa: cloning of the cDNA by dilution-amplification. Mol. Genet. Genomics 269, 508–516 (2003).
Teranishi, M., Nakamura, K., Morioka, H., Yamamoto, K. & Hidema, J. The native cyclobutane pyrimidine dimer photolyase of rice is phosphorylated. Plant Physiol. 146, 1941–1951 (2008).
Kaiser, G., Kleiner, O., Beisswenger, C. & Batschauer, A. Increased DNA repair in Arabidopsis plants overexpressing CPD photolyase. Planta 230, 505–515 (2009).
Kalbina, I. & Strid, Å. Supplementary ultraviolet-B irradiation reveals differences in stress responses between Arabidopsis thaliana ecotypes. Plant Cell Environ. 29, 754–763 (2006).
Ballestar, E. & Wolffe, A. P. Methyl-CpG-binding proteins. E ur. J. Biochem. 268, 1–6 (2001).
Nan, X. et al. Transcriptional repression by the methyl-CpG-binding protein MeCP2 involves a histone deacetylase complex. Nature 393, 386–389 (1998).
Yoshida, S., Forno, D., Cock, J. & Gomez, K. Laboratory manual for physiological studies of rice. The International Rice Research Institute, Manila, the Philippines 83 (1976).
Fang, C. X. et al. UV-induced differential gene expression in rice cultivars analyzed by SSH. Plant Growth Regul. 59, 245–253 (2009).
Livak, K. J. & Schmittgen, T. D. Analysis of relative gene expression data using real-time quantitative PCR and the 2−ΔΔCT method. Methods 25, 402–408 (2001).
Acknowledgements
This work was supported by the National Natural Science Foundation of China (grant No. 31300336); Provincial Natural Science Foundation of Fujian, China (grant No. 2015J01079); Foundation of The China Scholarship Council (grant No. 201508350004); and Fujian-Taiwan Joint Innovative Centre for Germplasm Resources and Cultivation of Crop (grant No. 2015-75. FJ 2011 Program, China).
Author information
Authors and Affiliations
Contributions
C.F. and W.L. conceived and designed the experiments; C.F., W.C., C.L., X.J., Y.L. and H.L. performed the experiments; C.F., W.C. and W.L. analyzed the data. C.F. and W.L. wrote the main manuscript text. Funding Acquisition, C.F. and W.L. All authors reviewed the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Rights and permissions
This work is licensed under a Creative Commons Attribution 4.0 International License. The images or other third party material in this article are included in the article’s Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by/4.0/
About this article
Cite this article
Fang, C., Chen, W., Li, C. et al. Methyl-CpG binding domain protein acts to regulate the repair of cyclobutane pyrimidine dimers on rice DNA. Sci Rep 6, 34569 (2016). https://doi.org/10.1038/srep34569
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep34569
This article is cited by
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.