Abstract
DNA methyltransferase (DNMT) inhibitors regulate target gene expression through epigenetic modifications and these compounds have primarily been studied for cancer therapy or reprogramming. However, the effect of DNMT inhibitors on the immunomodulatory capacity of human mesenchymal stem cells (hMSCs) has not been investigated. In the present study, we treated hMSCs with 5-azacytidine (5-aza), a DNMT inhibitor and confirmed that the inhibitory effects on mononuclear cell proliferation and cell migration toward activated T cells were increased. To identify the immunomodulatory factors stimulated through 5-aza treatment, we investigated the changes in promoter methylation patterns using methylation arrays and observed that the promoters of immunomodulatory factors, COX2 and PTGES and migration-related factors, CXCR2 and CXCR4, were hypomethylated after 5-aza treatment. In addition, we observed that the COX2-PGE2 pathway is one of the main pathways for the enhanced immunosuppressive activity of hMSCs through 5-aza treatment. We also determined that the migration of hMSCs toward ligands for CXCR2/CXCR4 was increased after 5-aza treatment. Moreover, using an experimental colitis model, we showed that 5-aza pre-treatment could enhance the therapeutic effect of MSCs against immune-related diseases.
Similar content being viewed by others
Introduction
Mesenchymal stem cells (MSCs) regulate inflammatory responses through cell-to-cell contact with immune cells and the secretion of anti-inflammatory soluble factors. MSC-derived soluble factors, including prostaglandin E2 (PGE2), indoleamine 2,3-dioxygenase (IDO), hepatocyte growth factor (HGF) and transforming growth factor β1 (TGF-β1), regulate inflammation1,2,3,4. Moreover, MSCs exhibit chemotaxis and migration toward various cytokines secreted from inflamed or damaged tissues. The Stromal cell-derived factor 1 (SDF-1)/Chemokine (C-X-C) motif receptor 4 (CXCR4) axis is a critical mechanism of MSC migration5. Based on these findings, a number of studies have shown that MSCs are potential therapeutics for various inflammatory diseases1,4.
MSCs can be isolated from various tissues, such as bone marrow, adipose, dermis, synovial fluid, periosteum and umbilical cord blood1. Among these, umbilical cord blood has several benefits, including its acquisition through non-invasive manipulation during tissue extraction processes and its most primitive nature among human tissues6. Umbilical cord blood-derived MSCs (UCB-MSCs) and other tissue-derived MSCs show variations in cellular characteristics and therapeutic effects depending on the primary cells. Therefore, a number of studies have found it difficult to identify representative selection markers for effective MSCs or to increase the therapeutic efficacy of MSCs in association with reproducible therapeutic effects7,8. Several studies have shown that pretreating MSCs with interferon-γ (IFN-γ), tumor necrosis factor-α (TNF-α) or interleukin-1β (IL-1β) increases immune-modulation and the therapeutic effects against inflammatory diseases9,10,11,12. Since the migration ability of MSCs is another key factor for therapeutic effect, there are efforts to increase migration ability through the enforced expression of chemokine receptors13,14,15.
DNA methyltransferases (DNMTs) and histone deacetylases (HDACs) are representative epigenetic modulators that regulate DNA methylation and the histone acetylation status of genomic DNA, respectively, to regulate the expression of target genes. As different cell types possess different methylation and histone status profiles, different responses to DNMT or HDAC inhibitors are observed according to the individual cellular characteristics. Within identical cell types, the chemical concentration or treatment duration could result in different effects16,17. Generally, DNMT or HDAC inhibitors act as tumor suppressors that increase the expression of tumor suppressor genes in various cancer cell lines18,19,20,21,22. However, DNMT or HDAC inhibitors have been reported to increase the efficiency of induced pluripotent stem cell generation or reprogramming through epigenetic modifications23,24,25,26,27. DNMT inhibitors including 5-aza-2′-deoxycytidine (5-aza) and decitabine have also been reported to have immune suppressive effects. Choi et al. showed that a DNMT inhibitor mitigated graft-versus-host disease without sacrificing the graft-versus-leukemia effect through the conversion of effector T cells to regulatory T cells28,29.
Although epigenetic modulators have various effects on cells and some modulators have demonstrated clinical effects, such as immune suppression, the effects of these molecules on the immune-modulatory properties of hMSCs have not been studied. Because both epigenetic modulators and hMSCs have common aspects in immune modulation, the effects of epigenetic modulators on immune modulation in hMSCs should be investigated to elucidate this potential cross-interaction. In the present study, we examined the effects of 5-aza on the immune-modulatory properties of MSCs. Moreover, we confirmed which immune modulation factors are regulated by 5-aza treatment. Finally, we also investigated the therapeutic effect of 5-aza treated hMSCs in a DSS-induced mouse colitis model.
Results
The DNMT inhibitor augments the immunomodulatory function of hMSCs
To investigate the effects of DNMT inhibitors on the immunomodulatory properties of hMSCs, a mixed leukocyte reaction (MLR) was performed after treating hMSCs with 5-aza. A significant reduction in the mitogen-induced proliferation of mononuclear cells derived from cord blood was observed. However, when cell-to-cell contact was mediated during MLR, treatment with 5-aza did not influence the suppressive effect of hMSCs on mononuclear cell (MNC) proliferation, regardless of the treatment dose (Fig. 1A). We next examined whether 5-aza could alter the secretion of immunoregulatory factors from hMSCs. First, MNCs were cultured with naïve or 5-aza-treated MSCs using transwell plates. Interestingly, 5-aza treatment enhanced the inhibitory effect of hMSCs on MNC proliferation in a dose-dependent manner (Fig. 1B). Next, MNCs were cultured with conditioned media (CM) from naïve or 5-aza-treated hMSCs and the results showed that 5-aza treatment could increase the suppressive effects of hMSC-derived CM on MNC proliferation in a dose-dependent manner (Fig. 1C). In addition, CM from 5-aza treated hMSCs significantly inhibited the proliferation of Jurkat cells, a human T-cell lymphoblast-like cell line (P = 0.007, Fig. S1). Because migration is another important function in the immunomodulation of hMSCs, we also confirmed the effects of 5-aza on the migration of hMSCs. We co-cultured hMSCs with T cell-activated MNCs using a CD3/CD28 antibody and subsequently quantified hMSC migration. Interestingly, the number of migrated cells increased significantly after 5-aza treatment (P = 1.12E-09, Fig. 1D).
5-aza stimulates the immune-modulation and migration of hMSCs.
(A–C) After treating hMSCs with 5-aza for 24 hr, an MLR assay was performed. (A) Direct MLR; hMSCs were co-cultured with hMNCs and MNC proliferation was determined using the bromodeoxyuridine kit. (B) Indirect MLR; co-culture was performed using transwell plates and MNC proliferation was detected. (C) CM-MLR; hMNCs were cultured with the CM from hMSC culture and hUCB-MNC proliferation was measured. (D) After treating hMSCs with 5-aza for 24 hr, changes in the migration toward activated T cells were determined. The migrated cells were counted in at least five fields and the average cell counts are graphically represented. Bar, 500 μm; *, p < 0.05; **, p < 0.01; ***, p < 0.001. Results show one representative experiment out of at least three. Results are shown as mean ± SD.
In a previous study, we showed that long-term treatment with 5-aza for more than 3 days increased the expression of senescence-related markers in hMSCs. Based on this finding, in the present study, we analyzed whether 5-aza treatment, which affects the immunomodulatory function of hMSCs, could influence the proliferation, senescence and apoptosis of hMSCs. Although 5-aza treatment for 24 hr slightly inhibited the proliferation of hMSCs, this treatment did not induce apoptosis (Fig. S2A–D). Furthermore, 5-aza treatment did not alter senescence-associated β-gal staining nor the expression of hMSC-specific markers and cellular senescence-related markers, such as p16INK4A and p21CIP1/WAF1 (Fig. S2E–G). Taken together, these findings suggest that 5-aza treatment for 24 hr enhances the secretory factor-mediated inhibitory effects of hMSCs on inflammatory cell proliferation and increases the migration of hMSCs toward activated T cells, without altering the expression of hMSC-specific or senescence-related markers.
Identification of the DNMT inhibitor target using microarray analysis
To identify crucial factors and elucidate the underlying mechanisms responsible for the 5-aza-mediated regulation of hMSC immune functions, we performed microarray and promoter methylation array analysis. The gene expression in hMSCs was assessed after treatment with the prominent pro-inflammatory cytokines, IFNγ and TNFα, to mimic the inflammatory environment. The expression of 150 genes was elevated more than 2-fold after stimulation with IFNγ and TNFα (Fig. 2A–B, Table S1). In addition, the promoter methylation array revealed the demethylation of the promoters for 7,734 genes and the methylation of the promoters for 5,615 genes after 5-aza treatment (Fig. 2C). Furthermore, among the 150 genes showing elevated expression after treatment with IFNγ and TNFα, 29 genes were hypomethylated and 26 genes were hyper- or hypomethylated after 5-aza treatment according to the promoter regions (Fig. 2D, Table S2).
5-aza affects promoter methylation status, regulated through the inflammatory cytokines, IFN-γ and TNF-α.
(A) Two lots of hMSCs were treated with IFN-γ and TNF-α for 24 hr and the up-regulated genes were investigated through microarray analysis. Changes in the gene expression were indicated through heat map imaging. (B) After treating hMSCs with 5-aza for 24 hr, changes in the methylation patterns in promoter regions were investigated through promoter array analysis. The changes in the methylation patterns were indicated through heat map imaging. (C) The changes of methylation status by treatment of 5-aza were indicated as a Venn diagram. Among the investigated promoters, the 10,664 promoters of 7,734 genes were hypomethylated, whereas 7,112 promoters of 5,615 genes were hypermethylated after 5-aza treatment. There was no change in methylation status of 85,086 promoters. (D) Changes in the methylation status of up-regulated genes through IFN-γ and TNF-α were investigated and the results are presented in a table.
The DNMT inhibitor regulates the expression of genes associated with MSC secretion of immunoregulatory factors and migration towards inflammatory sites
Among the 55 completely or partially hypomethylated genes, the immunomodulation-associated genes, cyclooxygenase 2 (COX2), prostaglandin E synthase (PTGES), leukemia inhibitory factor (LIF) and the cytokines IL-6 and IL-1β, were further examined. First, we determined the gene expression levels in hMSCs after treatment with inflammatory cytokines, IFNγ and TNFα, using quantitative real-time PCR. After treatment with IFNγ and TNFα, the expression of these genes was significantly increased in hMSCs isolated from different cord blood samples, designated #1 and #3 (P < 0.01, Fig. 3A). Next, we determined the gene expression levels after 5-aza treatment. The expression levels of COX-2, PTGES, IL-6 and IL-1β were significantly increased after 5-aza treatment (P < 0.01, Fig. 3B). We also examined the gene expression levels in 3 different hUCB-MSCs and the results showed that COX2 and IL-6 expression was increased after 5-aza treatment (Fig. S3A). To further determine whether pre-treatment with 5-aza affects the response of hMSCs against IFNγ and TNFα, these cells were treated with 5-aza for 24 hr, followed by treatment with IFNγ and TNFα for an additional 24 hr and the expression of the related genes was subsequently assessed. Interestingly, 5-aza pre-treatment significantly increased the expression level of PTGES compared to the sole treatment of IFNγ/TNFα in both #1 and #3 hMSCs, whereas changes in the expression of other genes varied depending on the cord blood sources (Fig. 3C). In addition, 5 different hMSCs were treated with 5-aza for 24 hr, followed by treatment with IFNγ for an additional 24 hr and subsequently COX2 expression was assessed. The pre-treatment with 5-aza increased COX2 expression compared with IFNγ treatment alone (Fig. S3B). No migration-related genes were identified among the hypomethylated genes showing increased expression after IFNγ and TNFα treatment. However, the promoter array analysis showed that the promoters of CXCR2, CXCR4, CXCR6, metallopeptidase 2 (MMP2), MMP11, MMP15 and MMP16 were hypomethylated after 5-aza treatment (Table S3). We also examined whether the expression of CXCR2, CXCR4 and CXCR6 was increased after 5-aza treatment using real-time qPCR and the results showed the increased expression of CXCR2 and CXCR4 in 5 different hUCB-MSCs (Fig. 3D, Fig. S3C). Moreover, the elevated expression of MMP11, MMP15 and MMP16 was observed after 5-aza treatment (Fig. S3D).
5-aza regulates the expression of genes associated with the hMSC secretion of immune-regulatory factors and migration into inflammatory sites.
(A) After treating hMSCs with IFN-γ and TNF-α, changes in the expression of 5 representative genes, selected via microarray analysis, were investigated in 2 lots of hMSCs (Fig 2). The expression was confirmed through real-time qPCR and the relative ratio to the control is graphically represented. (B) After treating hMSCs with 5-aza, the expression of indicated genes was detected and compared with that in control hMSCs (CTL). (C) The cells were pretreated with 5-aza (2 μM) for 24 hr and subsequently treated with IFN-γ/TNF-α for 24 hr (5-aza + IT treatment). The expression of indicated genes was determined and the results were compared with those in hMSCs treated with IT only (IT-treated). (D) After treatment with 5-aza, the expression of CXCR2, CXCR4 and CXCR6 was measured and compared with that in control hMSCs (CTL). *, p < 0.05; **, p < 0.01. Results are shown as mean ± SD.
The DNMT inhibitor augments PGE2 production in hMSCs through the up-regulation of synthesis enzymes
PGE2 is a well-known immune modulator that plays a role in the MSC-mediated regulation of immune cell activation2,30,31. To determine whether the COX2-PGE2 pathway is involved in the 5-aza-mediated enhancement of hMSC immune function, we examined the expression of COX2 and PTGES, crucial enzymes for PGE2 synthesis, after treatment with different doses of 5-aza. After treating hMSCs with 5-aza for 24 hr, the expression of COX2 and PTGES was increased on both mRNA and protein levels (Fig. 4A–B). The PGE2 concentration in the CM was also elevated after 5-aza treatment (Fig. 4C). Furthermore, COX2 inhibition through siRNA significantly restored the robust inhibitory effect of 5-aza-treated hMSCs on MNC proliferation (Fig. 4D). To determine whether the increase in COX2 and PTGES expression through 5-aza is associated with demethylation of the gene promoter, changes in the methylation pattern following 5-aza treatment were analyzed using methyl-specific PCR (Fig. 4E). The methylation of the promoters of both COX2 and PTGES was reduced after 5-aza treatment (Fig. 4F).
5-aza increases the production of PGE2 from hMSCs through the up-regulation of synthesis enzymes.
(A-B) After treating hMSCs with 5-aza for 24 hr, COX2 and PTGES expression was detected through (A) real-time qPCR and (B) western blot analysis (C) After treating hMSCs with 5-aza for 24 hr, the changes in PGE2 expression levels were measured using ELISA. PGE2 secretion from hMSCs was increased by 5-aza treatment. (D) After treatment of 5-aza with or without the inhibition of COX2 (siCOX2), indirect-MLR was performed. The suppression of MLR by hMSCs was increased by treating with 5-aza and the effect was rebounded by inhibition of COX2. (E) Schematic diagrams indicating the locations of the promoter primers. (F) After treating hMSCs with 5-aza, bisulfide conversion and methyl-specific PCR were performed. In 5-aza treated hMSCs, the promoters of COX2, PTGES1 and PTGES2 were hypomethylated compared with control MSCs. pm, promoter locus; TSS, transcription start site; *, p < 0.05; **, p < 0.01; ***, p < 0.001. Results show one representative experiment out of at least three. Results are shown as mean ± SD.
The DNMT inhibitor increases CXCR2 and CXCR4 expression and hMSC chemotaxis
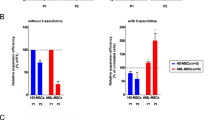
MSCs express CXCR2 and CXCR4 and show chemotactic activity towards their ligands, SDF-1/CXCL12 and IL-8/CXCL832. Moreover, the SDF-1/CXCR4 axis is important for the migration of MSCs towards target tissues with inflammatory reactivity14,33. We investigated whether the increased CXCR2 and CXCR4 expression after 5-aza treatment affects the hMSC migration. After the treatment of 5-aza on hMSCs for 24 hr, the expression of CXCR2 and CXCR4 was increased on mRNA and protein levels (Fig. 5A, B). Next, we confirmed whether the migration of hMSCs toward CXCR2 and CXCR4 ligands is altered through 5-aza treatment. The number of migrated cells toward IL-8, CXCL1 and SDF-1 was significantly increased after 5-aza treatment (P = 0.0028, 0.0001, 0.0023, respectively, Fig. 5C). To determine whether the increase in CXCR2 and CXCR4 expression through 5-aza is associated with demethylation of the gene promoter, changes in the methylation pattern following 5-aza treatment were analyzed using methyl-specific PCR (Fig. 5D). The promoters of both CXCR2 and CXCR4 were hypomethylated after 5-aza treatment (Fig. 5E).
5-aza stimulates the migration ability of hMSCs through the increase of CXCR2 and CXCR4.
(A-B) After treating hMSCs with 5-aza, CXCR2 and CXCR4 expression was confirmed through (A) real-time qPCR and (B) western blot analysis. (C) After 5-aza treatment, the migration of hMSCs was determined after treatment with the CXCR2 or CXCR4 ligands. (D) Schematic diagrams indicating the locations of the promoter primers. (E) After treating hMSCs with 5-aza, bisulfide conversion and methyl-specific PCR were performed. *, p < 0.05; **, p < 0.01; ***, p < 0.001. Results show one representative experiment out of at least three. Results are shown as mean ± SD.
The DNMT inhibitor augments the immunomodulatory and migratory properties of hMSCs in a murine inflammatory bowel disease (IBD) model
According to previous studies, PGE2 secretion from hMSCs is important for the regulation of immune reactions and regulatory T cell (Treg) populations, preventing the development of disease in a mouse IBD model2,6,14. In addition, CXCR4 up-regulated MSCs showed an increased therapeutic efficacy of transplantation in DSS-induced colitis11. Based on these results and the results of in vitro analyses, we confirmed whether 5-aza treatment increases the immunomodulation and migration of hMSCs in a DSS-induced colitis mouse model. hMSCs with or without 5-aza treatment were injected intraperitoneally at 24 hr after colitis induction with DSS. Body weight changes were monitored for 12 days and the results showed that 5-aza-treated hMSCs exerted protective effects on the reduction of body weight and lethality in mice with DSS-induced colitis (Fig. 6A). The shortening of the colon length was protected by hMSC injection and further improved by 5-aza-treated hMSC infusion (Fig. 6B). In addition, we detected and quantified the cell migration toward the inflamed colon and observed that 5-aza-treated hMSCs were significantly more infiltrated in the colon tissues (P = 0.0137, Fig. 6C, 6D, S4).
5-aza enhances the immune-modulation and migration ability of hMSCs in DSS-induced colitis mice.
(A) hMSCs with or without 5-aza treatment were injected into mice with DSS-induced colitis. Body weight loss and survival rate were monitored. Number of mice for each group: (-) = 7, (+) = 9, hMSC = 10, 5-aza treated hMSC = 8. (B) The colon length was measured and shown in a graph. Number of mice for each group: (-) = 7, (+) = 9, hMSC = 10, 5-aza treated hMSC = 8. (C) CFDA-SE-labeled hMSCs were detected through confocal image analysis. Bar, 100 μm. (D) The hMSC distribution in colon tissues was analyzed using real-time qPCR of the human-specific ALU gene. *, p < 0.05; **, p < 0.01; ***, p < 0.001. Results are shown as mean ± SD.
The DNMT inhibitor-mediated COX-2 activation in hMSCs is critical for the amelioration of colitis
Given that DNMT inhibitor treatment enhanced the expression of COX2 in hMSCs, we explored the physiological role of COX2 activation in the immunoregulatory capability of 5-aza-treated hMSCs. As expected, COX2 inhibition by specific siRNA abrogated the ability of 5-aza-treated hMSCs to protect mice from body weight loss and lethality (Fig. 7A). Although the administration of non-treated (naïve) hMSCs exerted a limited protective effect in body weight loss, naïve hMSCs significantly decreased the disease activity index and 5-aza pretreatment further improved this effect (Fig. 7B). In contrast, COX2 inhibition in 5-aza-treated hMSCs resulted in the loss of this protective effect (Fig. 7B). On histological examination, destruction of colonic epithelium and infiltration of inflammatory cells were observed in DSS-induced colitic mice. In hMSC-injected mice, mucosal destruction and lymphocyte infiltration were partially reduced. Importantly, the administration of 5-aza-treated hMSCs greatly suppressed the histological damage and the treatment of siRNA for COX2 on 5-aza-treated hMSCs neither prevented the epithelial destruction nor the lymphocyte infiltration (Fig. 7C and D).
5-aza-mediated immunomodulatory effect of hMSCs is COX-2 dependent.
siRNA for COX-2 was transfected into hMSCs. siRNA-transfected hMSCs with or without 5-aza treatment intraperitoneally injected into DSS-induced colitic mice. (A) Body weight loss and survival rate were monitored. Number of mice for each group: (-) = 12, (+) = 19, siCTL-hMSC = 18, siCTL-hMSC + 5-aza = 18, siCOX2-hMSC + 5-aza = 18. (B) Disease activity index for colitis severity was evaluated. (C-D) Histopathological analysis of colon, Bar, 200 μm, Five to seven mice/group were used. *, p < 0.05; **, p < 0.01; ***, p < 0.001. Results are shown as mean ± SD.
Discussion
In the present study, we investigated the effect of the DNMT inhibitor, 5-aza, on the immune-modulatory and migratory properties of hMSCs. We observed that 5-aza treatment augmented immune-modulation and migration through the up-regulation of immune modulators and chemotaxis-related factors in hMSCs. In addition, 5-aza treatment increased the immunosuppressive effects of hMSCs in both MLR, using transwell plates or CM. These findings suggest that 5-aza increases the secretion of immunomodulatory soluble factors from hMSCs, regardless of the interaction between hMSCs and lymphocytes. However, when cell-to-cell contact between hMSCs and lymphocytes was facilitated, naïve hMSCs suppressed lymphocyte proliferation to a similar extent as unactivated lymphocytes and the additive effect of 5-aza treatment could not be determined, suggesting that additional factors, such as adhesion molecules, are involved in the direct MLR.
The up-regulated expression and demethylation of the COX2 and PTGES promoters and increased PGE2 secretion from hMSCs were also observed after 5-aza treatment. In the present study, we observed that the increased PGE2 secretion after 5-aza treatment augmented the suppressive effect of hMSCs on lymphocyte proliferation. PGE2 is a powerful immunomodulator secreted by hMSCs. Previous studies have reported that hMSCs inhibit Th1/Th17 differentiation and induce regulatory phenotypes through COX2-dependent PGE2 secretion34,35,36. Moreover, PGE2 secretion from MSCs in response to inflammatory cytokines or ligands is critical for the immunomodulatory function of MSCs in various immune disease models, including arthritis and colitis2,37. The secretion of PGE2 from MSCs has been reported to regulate dendritic cells, CD4+ helper T cells, B cells, NK cells, monocytes and macrophages, exerting anti-inflammatory effects1. Interestingly, the regulation of COX2 expression through promoter methylation has been reported in other cell types. Fang et al. reported that the enhancement of COX2 expression in antiviral defense is regulated through DNA hypomethylation. These authors showed that COX2 expression is regulated through DNMT3a and DNMT3b in both A549 cells and peripheral blood mononuclear cells infected with influenza A virus38. Akhtar et al. reported that Helicobacter pylori (H. pylori)-stimulated COX2 expression in gastric epithelial cells was dependent on the methylation status of the individual COX2 promoter. Based on these findings, the methylation status of the COX2 promoter is different among different donor-derived gastric epithelial cells, suggesting that the increased COX2 expression in response to H. pylori was dependent on the methylation status of COX2 promoters. Moreover, after treatment with 5-aza, gastric epithelial cells containing methylated COX2 promoters showed a significant enhancement of COX2 expression and PGE2 secretion in response to H. pylori39.
In the present study, we also observed that 5-aza treatment increased hMSC migration through the CXCR2/IL-8, CXCR2/CXCL1 and CXCR4/SDF-1 axes. IL-8 and CXCL1 are proinflammatory cytokines that play a role in the recruitment of neutrophils in inflamed tissues and the expression of these cytokines is elevated in inflammatory diseases, including IBD40,41,42,43,44,45. Xu et al. reported that the over-expression of CXCR1/CXCR2 in mesenchymal stromal cells could increase recruitment to the sites of degenerated myocardium46. On the other hand, SDF-1 plays a role in the recruitment of lymphocytes, hematopoietic stem cells (HSCs) and MSCs5,47,48. It has been reported that the SDF-1/CXCR4 axis regulates the migration of transplanted bone marrow MSCs towards the inflamed tissues in an acute pancreatitis rat model33. A number of studies have reported that the SDF-1/CXCR4 axis is important for the migration of MSCs and there have been several attempts to improve the migration of MSCs through the up-regulation of CXCR45,49,50,51. In the present study, we observed that the expression of CXCR2 and CXCR4 was up-regulated and that their associated promoters were demethylated in hMSCs after 5-aza treatment. The epigenetic regulation of CXCR2 or CXCR4 expression has been addressed in several studies. CXCR2 has been reported to be epigenetically regulated through histone H3 acetylation52,53. Considering the association between DNA methylation and histone acetylation, these reports propose the regulation of CXCR2 through DNA methylation54. Sato et al. reported that CXCR4 is regulated through DNA methylation in pancreatic cancer. These authors observed a significant inverse correlation between methylation and the mRNA expression of CXCR4 in a large panel of pancreatic cancer cell lines55.
In the DSS-induced colitis mouse model, we observed that 5-aza-treated hMSCs showed augmented therapeutic effects and migration capacities compared with naïve hMSCs. In the present study, naïve hMSCs showed a limited protective effect on body weight loss and lethality in colitic mice and slightly reduced the shortening of colon length. Because hMSCs are derived from different donor tissues, there might be variations in DNA methylation among the hMSCs and these variations could reflect the inconsistent therapeutic efficacy of MSCs against IBD56,57. Moreover, the difference in the promoter methylation status could account for the different responses to DNA demethylating agents, including 5-aza. The hMSCs with hypomethylated immunomodulatory gene promoters would have more significant effects than naïve cells and show weak responses to DNA demethylating agents. In contrast, hMSCs carrying hypermethylated gene promoters would only exhibit slight immunomodulatory effects compared with naïve cells and have a strong response to DNA demethylating agents. Further studies regarding the relationships between variations in DNA methylation and the therapeutic effects of hMSCs are needed for the clinical application of hMSCs. However, regardless of the variations among hMSCs derived from different donor tissues, DNMT inhibitor treatment consistently augmented the physiological effect of hMSCs through the activation of COX-2 signaling.
Taken together, treatment with a demethylating agent (5-aza) increased the immunomodulation and migration of hMSCs through the demethylation of target gene promoters. These findings provide new insight to enhance the immune-modulatory effects of hMSCs via epigenetic modifications. Moreover, the different responses to 5-aza of hMSCs derived from different donors suggest that variations in DNA methylation influence immune-modulatory effects of hMSCs. Further studies concerning the epigenetic regulation of the immune-modulatory gene expression in hMSCs would be beneficial for the controlled use of effective hMSCs as therapeutics and the development of novel methodologies to increase the therapeutic effects of hMSCs in human diseases.
Methods
Isolation and Culture of hUCB-Derived MNCs and MSCs
hUCB-MSCs and MNCs were isolated and maintained as described in the Supplementary Methods section.
Mixed Leukocyte Reaction (MLR)
hMSCs were plated onto 6-well plates at 3 × 105 cells/well. After 24 hr, the cells were treated with 2 μM 5-aza for 24 hr. For direct co-culture, hMSCs were treated with 25 mg/ml mitomycin C for 1 hr and the cells were plated at 1 × 104 cells/well onto 96-well plates. MNCs were activated using Concanavalin A (ConA) and seeded onto plates at 5 × 104 cells/well. For indirect co-culture, 0.4-μm-poresize transwells (24-well plate, Corning, Corning, NY) were used. MNCs were activated with ConA and plated at 1 × 106 cells/well in the lower chamber and 5-aza-treated hMSCs were seeded at 1 × 105 cells/well in the upper chamber. After 3 days, the proliferation rate was analyzed using a BrdU assay (Bromodeoxyuridine kit, Roche, Upper Bavaria, Germany).
For the MLR assay using CM, MSCs were seeded at 3 × 105 cells/well onto 6-well plates and treated with 5-aza for 24 hr. After washing 3 times with PBS, the media were changed to RPMI 1640 (Gibco BRL) without FBS and enriched for 4~5 days. The CM were collected and used for MNC culture. MNCs were plated at 1 × 105 cells/well onto 96-well plates and treated with CM containing 10% FBS with or without ConA. After 3 days, the BrdU assay was performed.
RNA Interference
The cells were transfected with siRNA at 60% confluency. The siRNAs for PTGS2 (siCOX-2, L-004557-00) and the nontargeting control (siControl 1, D-001810-01) were purchased from Dharmacon (Chicago, IL). The experiments were conducted using DharmaFECT1 (Dharmacon) as a transfection agent and siRNA at a concentration of 50 nmol/L. After 48 hr, the medium was changed and the cells were used for the following studies.
Animal study and Ethical Statement
C57BL/6J mice (male; age, 8–10 wk) were obtained from Jackson Laboratory (Bar Harbor, ME). The mice were housed in groups under conventional conditions in the animal facility of Seoul National University. Acute colitis was induced in mice after the addition of 3% (w/v) dextran sulfate sodium (DSS, MP Biochemicals, Solon, OH) in drinking water for 7 days. hMSCs were treated with siRNA or 2 μM 5-aza before administration and washed with PBS to remove residual 5-aza. The body weight and morbidity were monitored over 11 days. At day 7, colitis severity was measured by evaluating the disease activity index through the scoring of weight loss (0~4), stool consistency (0~4), bleeding (0~4) and mouse activity (0~2). At the peak of disease (day 10), the mice were sacrificed and colon length was measured. Mice were monitored until they had reached criteria for humane endpoints. Mice losing above 40% of their original body weight or showing signs of distress (unphysiological bodily posture, shaggy fur and breathlessness/panting etc.) were sacrificed using CO2 asphyxiation. At the designated time points for tissue harvesting, mice were sacrificed by CO2-asphyxiation. Every effort was made to minimize suffering. All experimental protocols were approved by the Seoul National University Institutional Animal Care and Use Committee (Approval #; SNU-100125-8, SNU-111223-1 and SNU-130130-2) and carried out in accordance with the approved guidelines.
Histopathological Evaluation
Colon samples were collected, fixed in 10% formalin, subjected to consecutive steps of alcohol–xylene changes and embedded in paraffin. Sections that were 5-μm thick were prepared and stained with H&E. Leukocyte infiltration (0~4) and intestinal damage (0~4) were graded blindly.
Cell Tracking
To track the injected cells, hUCB-MSCs were labeled with 10 μmol/L CFSE (Molecular Probes) according to the manufacturer's protocol. CFSE-labeled cells were injected intraperitoneally. At 3 days after injection, 10-μm frozen colon sections were cut and examined for green fluorescence with a confocal microscope. To analyze hMSC concentration using real-time qPCR with Alu primers, DNAs were isolated from colon segments. Standard samples of mouse colon DNA which containing 16%, 4%, 1%, 0.25%, 0.063%, 0.031%, 0.015%, 0.006% and 0%(NC) of hMSCs DNA were prepared, respectively and obtained standard curve. We fixed the detection limit as 0.031% hMSCs of which Cp value has no difference with that of negative control. Based on standard curve, we obtained the concentration of hMSCs in each sample. The sequences of Alu primers which were used for real-time qPCR are as follows;
GTCAGGAGATCGAGACCATCCC (forward) and TCCTGCCTCAGCCTCCCAAG (reverse)58.
Microarray analysis
Total RNA was extracted from human cells using the TRI Reagent (Molecular Research Center, Inc., Cincinnati, OH) according to the manufacturer's instructions. Following homogenization, 1 ml of solution was transferred to a 1.5-ml Eppendorf tube and centrifuged at 12,000 g for 10 min at 4°C to remove insoluble material. The supernatant containing RNA was collected, mixed with 0.2 ml of chloroform and centrifuged at 12,000 g for 15 min at 4°C. Subsequently, the RNA in the aqueous phase was transferred to a new tube, precipitated with 0.5 ml of isopropyl alcohol and recovered through centrifugation at 12,000 g for 10 min at 4°C. The RNA pellet was briefly washed in 1 ml of 75% ethanol and centrifuged at 7,500 g for 5 min at 4°C. The RNA pellet was dissolved in nuclease-free water and the quality and quantity of the RNA was assessed through Agilent Bioanalyzer2100 analysis. Gene expression was analyzed using a GeneChip® Affymetrix Primeview array (Affymetrix, Santa Clara, CA), comprising more than 530,000 probes representing approximately 20,000 well-characterized human genes. For each gene, eleven pairs of oligonucleotide probes were synthesized in situ on the arrays.
Biotinylated cRNA was prepared according to the standard Affymetrix protocol from 250 ng of total RNA (Expression Analysis Technical Manual, 2001, Affymetrix). Following fragmentation, 12 μg of RNA was hybridized for 16 hr at 45°C onto the GeneChip Human Genome Array. The GeneChips were washed, stained in the Affymetrix Fluidics Station 450 and scanned using the Affymetrix Gene Chip Scanner 3000 7G. The data were analyzed using Microarray Suite version 5.0 (MAS 5.0) with Affymetrix default analysis settings and global scaling as a normalization method. The trimmed mean target intensity of each array was arbitrarily set to 100. The normalized and log-transformed intensity values were analyzed using GeneSpring GX 12.5 (Agilent Technologies, Santa Clara, CA).The fold-change filters required that the genes be present in at least 200% of controls for up-regulated genes and less than 50% of controls for down-regulated genes. Hierarchical clustering data were clustered into groups that behave similarly across experiments using GeneSpring GX 12.5 (Agilent Technologies). The Euclidean distance, with average linkage, was used as the clustering algorithm.
Promoter array
After treatment with 2μM 5-aza for 24 hr, the samples for promoter array analysis were prepared using the MethylMiner™ Methylated DNA Enrichment Kit (Applied Biosystems) according to the manufacturer's instructions. We used arrays comprising 0.4 million 50-mer to 60-mer oligonucleotides covering chromosome 21 at a median probe density of 1 per 60 bp (Agilent Technologies). Microarray protocols, including labeling, hybridization and post-hybridization washing procedures, were conducted according to the manufacturer's instructions and are available at http://www.agilent.com.
The MeDIP (cyanine 5-dUTP) and reference DNA (cyanine 3-dUTP) were labeled (Genomic DNA Enzymatic Labeling Kit, Agilent Technologies), cleaned (Amicon filters, Millipore) and quantified. Competitive hybridization onto the microarray was subsequently performed (Methylation Hybridization Kit, Agilent Technologies) according to the manufacturer's instructions in a rotating SureHyb chamber at 67°C for 40 hr. The washed slides were scanned (High-Resolution C Scanner, Agilent Technologies) and fluorescence was measured using Feature Extraction software.
These two signal values were normalized using background subtraction and the signal ratio (MeDIP/input), signal log ratio [log2 (MeDIP/input)], P[X] and P were obtained using Agilent Genomic Workbench software (Agilent Technologies). The log2 value is the Cy5:Cy3 fluorescence ratio (methylated DNA recovered by IP: total input DNA) for each probe, converted to a log2 scale, representing a relative measure of the amount of methylated DNA at each locus. We applied Median and the Lowess normalization to the raw data and filtered outlier probes to remove low-quality data points. The P[X] and P values, used in the Methylation IP (MeDIP) array analysis to obtain a binding call, were defined as the probability of how the X ( ) value deviates from the Gaussian distribution of X ( ) values of the entire genome of a sample. Here, the X value for a probe was obtained as the difference between the MeDIP and the input signals after adjusting the symmetry of its distribution. The value for a probe was calculated as an average X, considering the signals for neighboring probes (within 1 kb of the probe).
We defined hypermethylation as a normalized log2 ratio > 1.0 (equivalent to ~2.0-fold MeDIP/input signal intensity) based on previous validation experiments.
Methylation-specific PCR
For methylation-specific PCR, genomic DNA was extracted from cells using Accuprep® (Bioneer) according to the manufacturer's instructions. The bisulfite conversion of genomic DNA was performed using the MethyCode™ (Invitrogen) according to the manufacturer's instructions. The sodium bisulfite-modified DNA was amplified using the Accupower PCR premix (Bioneer). The primers used for each promoter were designed through an online web site (www.urogene.org/methprimer/) and the primer sequences are listed in Table S4.
Migration assay
The migration assay was performed based on the previous report59. Briefly, the migration of hMSCs was assessed using transwell chambers with polycarbonate membrane filters of 6.5 mm diameter and 8 μm pore size. The cells (4 × 105) in 400 μl of media (RPMI1640, 1% BSA) were added to the upper chamber and the lower chamber was filled with 800 μl of media (RPMI1640, 1% BSA) containing CD3/CD28 activated MNCs (8 × 106) with 100ng/ml IL-8, 100ng/ml CXCL1 or 100ng/ml SDF-1. T cell activation was performed using Dynabeads Human T-Activator CD3/CD28 (Invitrogen) according to the manufacturer's instructions. The transwell chambers were incubated for 24 hr at 37°C and 5% CO2 to facilitate cell migration. The cells on the top surface of the filters were wiped off with cotton swabs and the cells that had migrated into the lower chamber and attached to the lower surface of the filter were stained with crystal violet. To count the migrated hMSCs, at least three fields per sample were randomly chosen and photographed at × 40 magnification. The measurements were performed at least three times using different samples.
Statistical analysis
All experiments were conducted at least three times (n = 3) and the results are expressed as the mean ± SD. The statistical analysis was conducted via Student's t-test or one-way ANOVA. A value of p < 0.05 was considered significant.
References
Murphy, M. B., Moncivais, K. & Caplan, A. I. Mesenchymal stem cells: environmentally responsive therapeutics for regenerative medicine. Exp Mol Med 45, e54, 10.1038/emm.2013.94 (2013).
Kim, H. S. et al. Human umbilical cord blood mesenchymal stem cells reduce colitis in mice by activating NOD2 signaling to COX2. Gastroenterology 145, 1392–1403 e1391–1398 10.1053/j.gastro.2013.08.033 (2013).
Kang, J. W. et al. Soluble factors-mediated immunomodulatory effects of canine adipose tissue-derived mesenchymal stem cells. Stem Cells Dev 17, 681–693, 10.1089/scd.2007.0153 (2008).
Bassi, E. J., Aita, C. A. & Camara, N. O. Immune regulatory properties of multipotent mesenchymal stromal cells: Where do we stand? World J Stem Cells 3, 1–8, 10.4252/wjsc.v3.i1.1 (2011).
Marquez-Curtis, L. A. & Janowska-Wieczorek, A. Enhancing the migration ability of mesenchymal stromal cells by targeting the SDF-1/CXCR4 axis. Biomed Res Int 2013, 561098, 10.1155/2013/561098 (2013).
Zhang, X. et al. Isolation and characterization of mesenchymal stem cells from human umbilical cord blood: reevaluation of critical factors for successful isolation and high ability to proliferate and differentiate to chondrocytes as compared to mesenchymal stem cells from bone marrow and adipose tissue. J Cell Biochem 112, 1206–1218, 10.1002/jcb.23042 (2011).
Sioud, M., Mobergslien, A., Boudabous, A. & Floisand, Y. Mesenchymal stem cell-mediated T cell suppression occurs through secreted galectins. Int J Oncol 38, 385–390, 10.3892/ijo.2010.869 (2011).
Liu, G. Y. et al. Secreted galectin-3 as a possible biomarker for the immunomodulatory potential of human umbilical cord mesenchymal stromal cells. Cytotherapy 15, 1208–1217, 10.1016/j.jcyt.2013.05.011 (2013).
Sivanathan, K. N., Gronthos, S., Rojas-Canales, D., Thierry, B. & Coates, P. T. Interferon-gamma modification of mesenchymal stem cells: implications of autologous and allogeneic mesenchymal stem cell therapy in allotransplantation. Stem Cell Rev 10, 351–375, 10.1007/s12015-014-9495-2 (2014).
Pistoia, V. & Raffaghello, L. Unveiling the role of TNF-alpha in mesenchymal stromal cell-mediated immunosuppression. Eur J Immunol 44, 352–356, 10.1002/eji.201344372 (2014).
Fan, H. et al. Pre-treatment with IL-1beta enhances the efficacy of MSC transplantation in DSS-induced colitis. Cell Mol Immunol 9, 473–481, 10.1038/cmi.2012.40 (2012).
Duijvestein, M. et al. Pretreatment with interferon-gamma enhances the therapeutic activity of mesenchymal stromal cells in animal models of colitis. Stem Cells 29, 1549–1558, 10.1002/stem.698 (2011).
Li, H. et al. CCR7 guides migration of mesenchymal stem cell to secondary lymphoid organs: A novel approach to separate GvHD from GvL effect. Stem Cells, 10.1002/stem.1656 (2014).
Shen, W. et al. Intra-articular injection of human meniscus stem/progenitor cells promotes meniscus regeneration and ameliorates osteoarthritis through stromal cell-derived factor-1/CXCR4-mediated homing. Stem Cells Transl Med 3, 387–394, 10.5966/sctm.2012-0170 (2014).
Chen, W. et al. Overexpression of the mesenchymal stem cell Cxcr4 gene in irradiated mice increases the homing capacity of these cells. Cell Biochem Biophys 67, 1181–1191, 10.1007/s12013-013-9632-6 (2013).
Lee, S. et al. Histone deacetylase inhibitors decrease proliferation potential and multilineage differentiation capability of human mesenchymal stem cells. Cell Prolif 42, 711–720, 10.1111/j.1365-2184.2009.00633.x (2009).
Jung, J. W. et al. Histone deacetylase controls adult stem cell aging by balancing the expression of polycomb genes and jumonji domain containing 3. Cell Mol Life Sci 67, 1165–1176, 10.1007/s00018-009-0242-9 (2010).
Song, S. H., Han, S. W. & Bang, Y. J. Epigenetic-based therapies in cancer: progress to date. Drugs 71, 2391–2403, 10.2165/11596690-000000000-00000 (2011).
Kristensen, L. S., Nielsen, H. M. & Hansen, L. L. Epigenetics and cancer treatment. Eur J Pharmacol 625, 131–142, 10.1016/j.ejphar.2009.10.011 (2009).
Griffiths, E. A. & Gore, S. D. DNA methyltransferase and histone deacetylase inhibitors in the treatment of myelodysplastic syndromes. Semin Hematol 45, 23–30, 10.1053/j.seminhematol.2007.11.007 (2008).
Claus, R. & Lubbert, M. Epigenetic targets in hematopoietic malignancies. Oncogene 22, 6489-6496, 10.1038/sj.onc.1206814 (2003).
Zhu, W. G. & Otterson, G. A. The interaction of histone deacetylase inhibitors and DNA methyltransferase inhibitors in the treatment of human cancer cells. Curr Med Chem Anticancer Agents 3, 187–199 (2003).
Zhang, Z. & Wu, W. S. Sodium butyrate promotes generation of human induced pluripotent stem cells through induction of the miR302/367 cluster. Stem Cells Dev 22, 2268–2277, 10.1089/scd.2012.0650 (2013).
Han, J., Sachdev, P. S. & Sidhu, K. S. A combined epigenetic and non-genetic approach for reprogramming human somatic cells. PLoS One 5, e12297, 10.1371/journal.pone.0012297 (2010).
De Carvalho, D. D., You, J. S. & Jones, P. A. DNA methylation and cellular reprogramming. Trends Cell Biol 20, 609–617, 10.1016/j.tcb.2010.08.003 (2010).
Katz, L. S., Geras-Raaka, E. & Gershengorn, M. C. Reprogramming adult human dermal fibroblasts to islet-like cells by epigenetic modification coupled to transcription factor modulation. Stem Cells Dev 22, 2551–2560, 10.1089/scd.2013.0134 (2013).
Ding, X., Wang, Y., Zhang, D., Guo, Z. & Zhang, Y. Increased pre-implantation development of cloned bovine embryos treated with 5-aza-2'-deoxycytidine and trichostatin A. Theriogenology. 70, 622-630, 10.1016/j.theriogenology.2008.04.042 (2008).
Choi, J. et al. In vivo administration of hypomethylating agents mitigate graft-versus-host disease without sacrificing graft-versus-leukemia. Blood 116, 129–139, 10.1182/blood-2009-12-257253 (2010).
Hu, Y. et al. Decitabine facilitates the generation and immunosuppressive function of regulatory gammadeltaT cells derived from human peripheral blood mononuclear cells. Leukemia 27, 1580–1585, 10.1038/leu.2012.345 (2013).
Chen, K. et al. Human umbilical cord mesenchymal stem cells hUC-MSCs exert immunosuppressive activities through a PGE2-dependent mechanism. Clin Immunol 135, 448–458, 10.1016/j.clim.2010.01.015 (2010).
Yanez, R., Oviedo, A., Aldea, M., Bueren, J. A. & Lamana, M. L. Prostaglandin E2 plays a key role in the immunosuppressive properties of adipose and bone marrow tissue-derived mesenchymal stromal cells. Exp Cell Res 316, 3109–3123, 10.1016/j.yexcr.2010.08.008 (2010).
Ringe, J. et al. Towards in situ tissue repair: human mesenchymal stem cells express chemokine receptors CXCR1, CXCR2 and CCR2 and migrate upon stimulation with CXCL8 but not CCL2. J Cell Biochem 101, 135–146, 10.1002/jcb.21172 (2007).
Gong, J. et al. The SDF-1/CXCR4 axis regulates migration of transplanted bone marrow mesenchymal stem cells towards the pancreas in rats with acute pancreatitis. Mol Med Rep 9, 1575–1582, 10.3892/mmr.2014.2053 (2014).
Duffy, M. M. et al. Mesenchymal stem cell inhibition of T-helper 17 cell- differentiation is triggered by cell-cell contact and mediated by prostaglandin E2 via the EP4 receptor. Eur J Immunol 41, 2840–2851, 10.1002/eji.201141499 (2011).
Boniface, K. et al. Prostaglandin E2 regulates Th17 cell differentiation and function through cyclic AMP and EP2/EP4 receptor signaling. J Exp Med 206, 535–548, 10.1084/jem.20082293 (2009).
Tatara, R. et al. Mesenchymal stromal cells inhibit Th17 but not regulatory T-cell differentiation. Cytotherapy 13, 686–694, 10.3109/14653249.2010.542456 (2011).
Bouffi, C., Bony, C., Courties, G., Jorgensen, C. & Noel, D. IL-6-dependent PGE2 secretion by mesenchymal stem cells inhibits local inflammation in experimental arthritis. PLoS One 5, e14247, 10.1371/journal.pone.0014247 (2010).
Fang, J. et al. Epigenetic changes mediated by microRNA miR29 activate cyclooxygenase 2 and lambda-1 interferon production during viral infection. J Virol 86, 1010-1020, 10.1128/JVI.06169-11 (2012).
Akhtar, M. et al. Promoter methylation regulates Helicobacter pylori-stimulated cyclooxygenase-2 expression in gastric epithelial cells. Cancer Res 61, 2399–2403 (2001).
Nahidi, L. et al. Inflammatory bowel disease therapies and gut function in a colitis mouse model. Biomed Res Int 2013, 909613, 10.1155/2013/909613 (2013).
Forster, C. Tight junctions and the modulation of barrier function in disease. Histochem Cell Biol 130, 55–70, 10.1007/s00418-008-0424-9 (2008).
Williams, E. J. et al. Distribution of the interleukin-8 receptors, CXCR1 and CXCR2, in inflamed gut tissue. J Pathol 192, 533–539, 10.1002/1096-9896(2000)9999:9999 <::AID-PATH732 > 3.0.CO;2-X (2000).
Leal, R. F. et al. Identification of inflammatory mediators in patients with Crohn's disease unresponsive to anti-TNFalpha therapy. Gut, 10.1136/gutjnl-2013-306518 (2014).
Moser, B., Clark-Lewis, I., Zwahlen, R. & Baggiolini, M. Neutrophil-activating properties of the melanoma growth-stimulatory activity. J Exp Med 171, 1797–1802 (1990).
Schumacher, C., Clark-Lewis, I., Baggiolini, M. & Moser, B. High- and low-affinity binding of GRO alpha and neutrophil-activating peptide 2 to interleukin 8 receptors on human neutrophils. Proc Natl Acad Sci U S A 89, 10542–10546 (1992).
Xu, J., Chen, Q., Shi, C. & Yin, Z. Overexpression of CXCR1/CXCR2 on mesenchymal stromal cells may be an effective treatment for acute myocardial infarction. Cytotherapy 11, 990–991, 10.3109/14653240903233099 (2009).
Bleul, C. C., Fuhlbrigge, R. C., Casasnovas, J. M., Aiuti, A. & Springer, T. A. A highly efficacious lymphocyte chemoattractant, stromal cell-derived factor 1 (SDF-1). J Exp Med 184, 1101–1109 (1996).
Gul-Uludag, H. et al. Cationic liposome-mediated CXCR4 gene delivery into hematopoietic stem/progenitor cells: implications for clinical transplantation and gene therapy. Stem Cells Dev 21, 1587–1596, 10.1089/scd.2011.0297 (2012).
Marquez-Curtis, L. A., Gul-Uludag, H., Xu, P., Chen, J. & Janowska-Wieczorek, A. CXCR4 transfection of cord blood mesenchymal stromal cells with the use of cationic liposome enhances their migration toward stromal cell-derived factor-1. Cytotherapy 15, 840–849, 10.1016/j.jcyt.2013.02.009 (2013).
Stoicov, C., Li, H., Liu, J. H. & Houghton, J. Mesenchymal stem cells utilize CXCR4-SDF-1 signaling for acute, but not chronic, trafficking to gastric mucosal inflammation. Dig Dis Sci 58, 2466–2477, 10.1007/s10620-013-2782-y (2013).
Tao, Y. H., Ye, L., Wang, Y. M. & Wang, Z. [Mechanism for promoting repair of renal ischemia reperfusion injury by mesenchymal stem cells]. Zhongguo Dang Dai Er Ke Za Zhi 15, 157–161 (2013).
Sun, Y. et al. Epigenetic regulation of spinal CXCR2 signaling in incisional hypersensitivity in mice. Anesthesiology 119, 1198–1208, 10.1097/ALN.0b013e31829ce340 (2013).
Kiguchi, N. et al. Epigenetic augmentation of the macrophage inflammatory protein 2/C-X-C chemokine receptor type 2 axis through histone H3 acetylation in injured peripheral nerves elicits neuropathic pain. J Pharmacol Exp Ther 340, 577–587, 10.1124/jpet.111.187724 (2012).
Irvine, R. A., Lin, I. G. & Hsieh, C. L. DNA methylation has a local effect on transcription and histone acetylation. Mol Cell Biol 22, 6689–6696 (2002).
Sato, N., Matsubayashi, H., Fukushima, N. & Goggins, M. The chemokine receptor CXCR4 is regulated by DNA methylation in pancreatic cancer. Cancer Biol Ther 4, 70–76 (2005).
van Deen, W. K., Oikonomopoulos, A. & Hommes, D. W. Stem cell therapy in inflammatory bowel disease: which, when and how? Curr Opin Gastroenterol 29, 384–390, 10.1097/MOG.0b013e328361f763 (2013).
Sempere, J. M. et al. Single cell-derived clones from human adipose stem cells present different immunomodulatory properties. Clin Exp Immunol 176, 255–265, 10.1111/cei.12270 (2014).
Nicklas, J. A. & Buel, E. Development of an Alu-based, real-time PCR method for quantitation of human DNA in forensic samples. J Forensic Sci 48, 936–944 (2003).
Hu, X. et al. Hypoxic preconditioning enhances bone marrow mesenchymal stem cell migration via Kv2.1 channel and FAK activation. Am J Physiol Cell Physiol 301, C362–372, 10.1152/ajpcell.00013.2010 (2011).
Acknowledgements
This research was supported by a grant of the Korea Health Technology R&D Project through the Korea Health Industry Development Institute (KHIDI), funded by the Ministry of Health & Welfare, Republic of Korea (grant number : A120176). The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.
Author information
Authors and Affiliations
Contributions
S.L. designed the study, collected and analyzed the data and wrote the manuscript; H.-S.K. designed the study, collected and analyzed the data and wrote the manuscript; K.-H.R. designed the study, collected and analyzed the data and wrote the manuscript; B.-C.L. collected and analyzed the data; T.-H.S. collected and analyzed the data; J.-M.Y. collected and analyzed the data; Y.-L.K. collected and analyzed the data; K.-R.Y. collected and analyzed the data; K.-S.K. and K.-W.S. designed, supervised the study and critically contributed to analyzing the data
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Electronic supplementary material
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-NoDerivs 4.0 International License. The images or other third party material in this article are included in the article's Creative Commons license, unless indicated otherwise in the credit line; if the material is not included under the Creative Commons license, users will need to obtain permission from the license holder in order to reproduce the material. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-nd/4.0/
About this article
Cite this article
Lee, S., Kim, HS., Roh, KH. et al. DNA methyltransferase inhibition accelerates the immunomodulation and migration of human mesenchymal stem cells. Sci Rep 5, 8020 (2015). https://doi.org/10.1038/srep08020
Received:
Accepted:
Published:
DOI: https://doi.org/10.1038/srep08020
This article is cited by
-
DNA methyltransferase 1 deficiency improves macrophage motility and wound healing by ameliorating cholesterol accumulation
npj Regenerative Medicine (2023)
-
Hypomethylating agents (HMA) for the treatment of acute myeloid leukemia and myelodysplastic syndromes: mechanisms of resistance and novel HMA-based therapies
Leukemia (2021)
-
Functional enhancement strategies for immunomodulation of mesenchymal stem cells and their therapeutic application
Stem Cell Research & Therapy (2020)
-
Improving the immunosuppressive potential of articular chondroprogenitors in a three-dimensional culture setting
Scientific Reports (2020)
-
Epigenetic modification of mesenchymal stromal cells enhances their suppressive effects on the Th17 responses of cells from rheumatoid arthritis patients
Stem Cell Research & Therapy (2018)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.