Abstract
Na+/Ca2+ exchangers use the Na+ electrochemical gradient across the plasma membrane to extrude intracellular Ca2+ and play a central role in Ca2+ homeostasis. Here, we elucidate their mechanisms of extracellular ion recognition and exchange through a structural analysis of the exchanger from Methanococcus jannaschii (NCX_Mj) bound to Na+, Ca2+ or Sr2+ in various occupancies and in an apo state. This analysis defines the binding mode and relative affinity of these ions, establishes the structural basis for the anticipated 3:1 Na+/Ca2+-exchange stoichiometry and reveals the conformational changes at the onset of the alternating-access transport mechanism. An independent analysis of the dynamics and conformational free-energy landscape of NCX_Mj in different ion-occupancy states, based on enhanced-sampling molecular dynamics simulations, demonstrates that the crystal structures reflect mechanistically relevant, interconverting conformations. These calculations also reveal the mechanism by which the outward-to-inward transition is controlled by the ion occupancy, thereby explaining the emergence of strictly coupled Na+/Ca2+ antiport.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$189.00 per year
only $15.75 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Blaustein, M.P. & Lederer, W.J. Sodium/calcium exchange: its physiological implications. Physiol. Rev. 79, 763–854 (1999).
DiPolo, R. & Beaugé, L. Sodium/calcium exchanger: influence of metabolic regulation on ion carrier interactions. Physiol. Rev. 86, 155–203 (2006).
Clapham, D.E. Calcium signaling. Cell 131, 1047–1058 (2007).
Berridge, M.J., Bootman, M.D. & Roderick, H.L. Calcium signalling: dynamics, homeostasis and remodelling. Nat. Rev. Mol. Cell Biol. 4, 517–529 (2003).
Hilgemann, D.W. Unitary cardiac Na+, Ca2+ exchange current magnitudes determined from channel-like noise and charge movements of ion transport. Biophys. J. 71, 759–768 (1996).
Hilgemann, D.W., Nicoll, D.A. & Philipson, K.D. Charge movement during Na+ translocation by native and cloned cardiac Na+/Ca2+ exchanger. Nature 352, 715–718 (1991).
Reeves, J.P. & Hale, C.C. The stoichiometry of the cardiac sodium-calcium exchange system. J. Biol. Chem. 259, 7733–7739 (1984).
Blaustein, M.P. & Russell, J.M. Sodium-calcium exchange and calcium-calcium exchange in internally dialyzed squid giant axons. J. Membr. Biol. 22, 285–312 (1975).
Rasgado-Flores, H. & Blaustein, M.P. Na/Ca exchange in barnacle muscle cells has a stoichiometry of 3 Na+/1 Ca2+ . Am. J. Physiol. 252, C499–C504 (1987).
Kimura, J., Noma, A. & Irisawa, H. Na-Ca exchange current in mammalian heart cells. Nature 319, 596–597 (1986).
Kang, T.M. & Hilgemann, D.W. Multiple transport modes of the cardiac Na+/Ca2+ exchanger. Nature 427, 544–548 (2004).
Jardetzky, O. Simple allosteric model for membrane pumps. Nature 211, 969–970 (1966).
Hilgemann, D.W. Regulation and deregulation of cardiac Na+-Ca2+ exchange in giant excised sarcolemmal membrane patches. Nature 344, 242–245 (1990).
Matsuoka, S., Nicoll, D.A., Reilly, R.F., Hilgemann, D.W. & Philipson, K.D. Initial localization of regulatory regions of the cardiac sarcolemmal Na+-Ca2+ exchanger. Proc. Natl. Acad. Sci. USA 90, 3870–3874 (1993).
Philipson, K.D. & Nicoll, D.A. Sodium-calcium exchange: a molecular perspective. Annu. Rev. Physiol. 62, 111–133 (2000).
Nicoll, D.A., Hryshko, L.V., Matsuoka, S., Frank, J.S. & Philipson, K.D. Mutation of amino acid residues in the putative transmembrane segments of the cardiac sarcolemmal Na+-Ca2+ exchanger. J. Biol. Chem. 271, 13385–13391 (1996).
Nicoll, D.A., Ottolia, M. & Philipson, K.D. Toward a topological model of the NCX1 exchanger. Ann. NY Acad. Sci. 976, 11–18 (2002).
Ren, X., Nicoll, D.A., Xu, L., Qu, Z. & Philipson, K.D. Transmembrane segment packing of the Na+/Ca2+ exchanger investigated with chemical cross-linkers. Biochemistry 49, 8585–8591 (2010).
Cai, X. & Lytton, J. The cation/Ca2+ exchanger superfamily: phylogenetic analysis and structural implications. Mol. Biol. Evol. 21, 1692–1703 (2004).
Liao, J. et al. Structural insight into the ion-exchange mechanism of the sodium/calcium exchanger. Science 335, 686–690 (2012).
Almagor, L. et al. Functional asymmetry of bidirectional Ca2+-movements in an archaeal sodium-calcium exchanger (NCX_Mj). Cell Calcium 56, 276–284 (2014).
Marinelli, F. et al. Sodium recognition by the Na+/Ca2+ exchanger in the outward-facing conformation. Proc. Natl. Acad. Sci. USA 111, E5354–E5362 (2014).
Blaustein, M.P. & Santiago, E.M. Effects of internal and external cations and of ATP on sodium-calcium and calcium-calcium exchange in squid axons. Biophys. J. 20, 79–111 (1977).
Trosper, T.L. & Philipson, K.D. Effects of divalent and trivalent cations on Na+-Ca2+ exchange in cardiac sarcolemmal vesicles. Biochim. Biophys. Acta 731, 63–68 (1983).
DiPolo, R. & Beaugé, L. Asymmetrical properties of the Na-Ca exchanger in voltage-clamped, internally dialyzed squid axons under symmetrical ionic conditions. J. Gen. Physiol. 95, 819–835 (1990).
Matsuoka, S. & Hilgemann, D.W. Steady-state and dynamic properties of cardiac sodium-calcium exchange: ion and voltage dependencies of the transport cycle. J. Gen. Physiol. 100, 963–1001 (1992).
Miura, Y. & Kimura, J. Sodium-calcium exchange current: dependence on internal Ca and Na and competitive binding of external Na and Ca. J. Gen. Physiol. 93, 1129–1145 (1989).
Waight, A.B. et al. Structural basis for alternating access of a eukaryotic calcium/proton exchanger. Nature 499, 107–110 (2013).
Nishizawa, T. et al. Structural basis for the counter-transport mechanism of a H+/Ca2+ exchanger. Science 341, 168–172 (2013).
Wu, M. et al. Crystal structure of Ca2+/H+ antiporter protein YfkE reveals the mechanisms of Ca2+ efflux and its pH regulation. Proc. Natl. Acad. Sci. USA 110, 11367–11372 (2013).
Giladi, M. et al. Asymmetric preorganization of inverted pair residues in the sodium-calcium exchanger. Sci. Rep. 6, 20753 (2016).
Iwamoto, T., Uehara, A., Imanaga, I. & Shigekawa, M. The Na+/Ca2+ exchanger NCX1 has oppositely oriented reentrant loop domains that contain conserved aspartic acids whose mutation alters its apparent Ca2+ affinity. J. Biol. Chem. 275, 38571–38580 (2000).
John, S.A., Liao, J., Jiang, Y. & Ottolia, M. The cardiac Na+-Ca2+ exchanger has two cytoplasmic ion permeation pathways. Proc. Natl. Acad. Sci. USA 110, 7500–7505 (2013).
Ottolia, M., Nicoll, D.A. & Philipson, K.D. Mutational analysis of the alpha-1 repeat of the cardiac Na+-Ca2+ exchanger. J. Biol. Chem. 280, 1061–1069 (2005).
Reeves, J.P. & Sutko, J.L. Competitive interactions of sodium and calcium with the sodium-calcium exchange system of cardiac sarcolemmal vesicles. J. Biol. Chem. 258, 3178–3182 (1983).
Caffrey, M. & Cherezov, V. Crystallizing membrane proteins using lipidic mesophases. Nat. Protoc. 4, 706–731 (2009).
Otwinowski, Z. & Minor, W. Processing of X-ray diffraction data collected in oscillation mode. Methods Enzymol. 276, 307–326 (1997).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Petrek, M. et al. CAVER: a new tool to explore routes from protein clefts, pockets and cavities. BMC Bioinformatics 7, 316 (2006).
Phillips, J.C. et al. Scalable molecular dynamics with NAMD. J. Comput. Chem. 26, 1781–1802 (2005).
MacKerell, A.D. et al. All-atom empirical potential for molecular modeling and dynamics studies of proteins. J. Phys. Chem. B 102, 3586–3616 (1998).
Mackerell, A.D. Jr., Feig, M. & Brooks, C.L. III. Extending the treatment of backbone energetics in protein force fields: limitations of gas-phase quantum mechanics in reproducing protein conformational distributions in molecular dynamics simulations. J. Comput. Chem. 25, 1400–1415 (2004).
Staritzbichler, R., Anselmi, C., Forrest, L.R. & Faraldo-Gómez, J.D. GRIFFIN: a versatile methodology for optimization of protein-lipid interfaces for membrane protein simulations. J. Chem. Theory Comput. 7, 1167–1176 (2011).
Piana, S. & Laio, A. A bias-exchange approach to protein folding. J. Phys. Chem. B 111, 4553–4559 (2007).
Branduardi, D., Bussi, G. & Parrinello, M. Metadynamics with adaptive gaussians. J. Chem. Theory Comput. 8, 2247–2254 (2012).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Bonomi, M. et al. PLUMED: a portable plugin for free-energy calculations with molecular dynamics. Comput. Phys. Commun. 180, 1961–1972 (2009).
Branduardi, D., Gervasio, F.L. & Parrinello, M. From A to B in free energy space. J. Chem. Phys. 126, 054103 (2007).
Marinelli, F., Pietrucci, F., Laio, A. & Piana, S. A kinetic model of Trp-Cage folding from multiple biased molecular dynamics simulations. PLoS Comput. Biol. 5, e1000452 (2009).
Biarnes, X., Pietrucci, F., Marinelli, F. & Laio, A. METAGUI: A VMD interface for analyzing metadynamics and molecular dynamics simulations. Comput. Phys. Commun. 183, 203–211 (2012).
Corbi-Verge, C. et al. Two-state dynamics of the SH3-SH2 tandem of Abl kinase and the allosteric role of the N-cap. Proc. Natl. Acad. Sci. USA 110, E3372–E3380 (2013).
Daura, X. et al. Peptide folding: when simulation meets experiment. Angew. Chem. Int. Ed. 38, 236–240 (1999).
Branduardi, D., Marinelli, F. & Faraldo-Gómez, J.D. Atomic-resolution dissection of the energetics and mechanism of isomerization of hydrated ATP-Mg2+ through the SOMA string method. J. Comput. Chem. 37, 575–586 (2016).
Enright, A.J., Van Dongen, S. & Ouzounis, C.A. An efficient algorithm for large-scale detection of protein families. Nucleic Acids Res. 30, 1575–1584 (2002).
Acknowledgements
The experimental results reported in this article derive from measurements made at the Advanced Photon Source of the Argonne National Laboratory, GM/CA (23ID), which is operated by the University of Chicago Argonne, LLC, for the US Department of Energy, Office of Biological and Environmental Research, under contract DE-AC02-06CH11357. We thank the beamline staff for their assistance in data collection. This work was supported in part by the Howard Hughes Medical Institute; by grants from the National Institutes of Health (R01GM079179 to Y.J.) and the Welch Foundation (grant I-1578 to Y.J.); by the General Program of the National Natural Science Foundation of China (project 31470817 to J.L.); and by the Division of Intramural Research of the National Heart, Lung and Blood Institute, National Institutes of Health (to F.M. and J.D.F.-G.).
Author information
Authors and Affiliations
Contributions
J.L. and Y.J. designed the experimental studies and analyzed the resulting data. J.L., C.L. and Y.H. performed the experimental research. F.M. and J.D.F.-G. designed the computational research and analyzed the corresponding data. F.M. performed the computational work. J.L., Y.J., F.M. and J.D.F.-G. wrote the paper.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
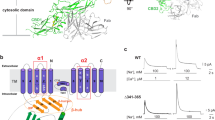
Supplementary Figure 1 Architecture of the Na+/Ca2+ exchanger and atomic structure of the outward-facing Na+-loaded state.
(a) Topology of the transmembrane functional unit of NCX. The so-called α-repeats are highlighted with solid colors (orange, blue). Eukaryotic NCX homologs contain a large intracellular regulatory domain between TM5 and TM6, whereas in microbial NCX homologs TM5 and TM6 are linked by a short loop. As shown in Figs. 2 and 4, in NCX_Mj the N-terminal half of TM7 is a straight helix (TM7ab) when the Sext binding site is empty, but is bent into two short helices (TM7a and 7b) when Na+ binds to Sext and this site becomes occluded. It is plausible that similar changes take place in the symmetry-related TM2, in the inward-facing states. (b) Crystal structure of NCX_Mj at high Na+ concentration (PDB code 3V5U) (Liao, J. et al. Structural insight into the ion-exchange mechanism of the sodium/calcium exchanger. Science 335, 686-690, 2012. Green and red spheres represent Na+ and water, respectively. The N- and C-terminal halves are colored yellow and cyan, respectively. (c) Highlight of the two α-repeats flanking the four central binding sites. TM2 and TM3 form α1 (yellow) and TM7 and TM8 form α2 (cyan). (d) Atomic details of the four binding sites, termed Sext, SCa, Smid and Sint. Residues that participate in Na+ chelation are indicated (with carbonyl oxygen atoms in red). Insets show the ion-coordination structure at each site. At Sext, SCa, and Sint, the ion-coordination numbers (5 or 6) and ion-ligand distances (~2.4 Å) are optimal for Na+ binding. The Smid site has a coordination number of 4 and ion-ligand distances of 2.6 Å and 2.9 Å, resulting in a calculated valence of far less than 1.0 for Na+; it is therefore not optimal for Na+, and thus the density at this site was assigned to a water molecule. Asp240 at Smid is suggested to be protonated, based on previous simulation and functional studies (Marinelli, F. et al. Proc. Natl. Acad. Sci. USA 111, E5354-62, 2014).
Supplementary Figure 2 Sr2+ and Ca2+ binding to NCX_Mj in a partially open outward-facing conformation.
(a) Titration of NCX_Mj crystals with decreasing amounts of Sr2+ at low Na+ concentration. The changes in Sr2+ occupancy are shown with Fo-Fc ion-omit maps calculated to 2.8 Å using data scaled to a common reference, and contoured at 7σ (solid-line panels). The map in the inset (dashed line) was calculated to 2.55 Å and contoured at 4σ, to reveal the bound Na+ (green spheres) and water (red sphere) observed at low Sr2+ concentration. (b) Titration with increasing amounts of Na+ at high Sr2+ concentration. The maps in the insets (dashed lines) were recalculated to 2.3 Å for a crystal soaked 10 mM Sr2+ and 10 mM Na+ and to 2.5 Å for a crystal soaked in 10 mM Sr2+ and 100 mM Na+, and contoured at 4σ, to reveal the bound Na+ (green spheres) and water (red sphere). At 10 mM Na+, the density reflects a mixture of partially bound Sr2+ at SCa, partially bound Na+ at Sint and at SCa, and a water molecule bound at Smid. At 100 mM Na+, the NCX_Mj structure reverts to the occluded state. (c) Titrations with Ca2+. At 10 mM, Ca2+ binds either to SCa or Smid, with SCa slightly preferred. At 0.1 mM Ca2+, the density distribution in the ion binding sites is virtually the same as that in the 2.5 mM Na+ only condition (Fig. 1c) indicating no bound Ca2+. Increasing Na+ to 10 mM similarly diminishes Ca2+ occupancy. All Fo-Fc ion omit maps (except those in the inset) were calculated to 2.65 Å using data scaled to a common reference and contoured at 5σ. The map in the inset is calculated at 2.2 Å and contoured at 4σ, and demonstrates that the density detected in the crystal soaked in 10 mM Ca2+ and 10 mM Na+ is a mixture of partially bound Ca2+ and Na+.
Supplementary Figure 3 Statistical survey of Ca2+-binding sites in the Protein Data Bank featuring two concurrent ion-carboxyl interactions.
(a) Distributions of observed Ca2+ coordination numbers in binding sites where the carboxyl groups are arranged so as to either two, one or no bi-dentate interactions. The number of Ca2+-binding sites identified in each case are indicated. Note that for the double bi-dentate mode observed in NCX_Mj (E54 and E213), the most probable coordination number, N, is 8 (>90%), but N = 7 is also observed. (b) Orientational preference in observed Ca2+-binding sites with two opposing carboxylate interactions in a bi-dentate mode, for N = 7 or N = 8. The figure shows the most probable angle (Φ) formed by the two opposing carboxyl groups (average ± STD in orange and grey, respectively), as well as the minimum distance between them, on average. The near orthogonal arrangement of E54 and E213 in NCX_Mj (green), and the separation between these groups, are characteristic of binding sites with N = 8. This analysis includes X-ray structures deposited in the PDB with less than 70% of sequence identity and with resolution higher than 2.5 Å. The analysis includes only Ca2+ binding sites that appear to be complete; a binding site is considered to be incomplete if the geometric center of the coordinating atoms deviates from the Ca2+ ion by more than 0.4 Å. (Note this threshold value is two times the standard deviation of this distance for binding sites in which the coordination number is 7 or 8). The coordinating atoms are defined as those within 3.0 Å of the ion, excluding hydrogen or carbon atoms, and other ions.
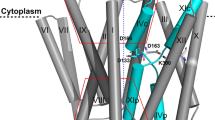
Supplementary Figure 4 Predicted structures of the Na+- and Ca2+-bound states of outward-facing, occluded NCX_Mj, contrasted with mutagenesis data.
(a-b) Close-up view of the predicted configuration of the ion-binding region of NCX_Mj, in the transport-ready Na+- and Ca2+-bound states. Note the striking symmetry of this region, relative to the membrane mid-plane, despite the fact the transporter adopts an outward-facing (i.e. asymmetric) conformation. The predictions derive from MD simulations and from a statistical survey of Ca2+-binding sites in the Protein Data Bank. Only side-chains flanking the four binding sites are highlighted. The dominant hydrogen-bonding patterns observed in the simulations are also indicated. A water-density iso-surface calculated from the simulation data is shown as a grey mesh, to indicate the degree of occlusion of the binding sites from the extracellular solution. (c-d) Residues lining the ion-binding sites in NCX_Mj whose mutation inactivates or significantly inhibits the activity of NCX_Mj or NCX1.1. (75% of the binding-site residues are identical in these two proteins, while the remaining 25% are homologous). Labels “A” and “B” indicate residues in NCX_Mj whose mutation to alanine inactivates the Na+-Ca2+ or Ca2+-Ca2+ exchange mechanism, respectively (<25% of wild-type activity), according to 45Ca2+ uptake assays of overexpressed NCX_Mj in right-side-out E. coli membrane vesicles (Marinelli, F. et al. Proc. Natl. Acad. Sci. USA 111, E5354-62, 2014; Giladi, M. et al. Sci. Rep. 6, 20753, 2016). Labels “C” and “D” indicate residues in NCX_Mj for which alanine mutation reduces the Na+-Ca2+ or Ca2+-Ca2+ exchange activity respectively (<50% of wild-type), according to the same measurements. The label “a” indicates residues in NCX1.1 whose mutation to cysteine inactivates or significantly inhibits the transport mechanism, as reported by patch-clamp recordings of the currents elicited upon Na+/Ca2+ exchange, either in the direct (Na+ inwards, Ca2+ outwards) or reverse mode (John, S. A. et al. Proc. Natl. Acad. Sci. USA 110, 7500-7505, 2013). The label “b” reflects comparable effects according to 45Ca2+ uptake experiments of NCX1.1, upon mutation of Ser to Ala or Cys, Thr to Ala, Glu to Gln, Asp to Asn and Asn to Val (Nicoll, D. A. et al. J Biol Chem 271, 13385-13391, 1996). The label “c” also reflects 45Ca2+ uptake measurements, for cysteine mutations (Iwamoto, T. et al. J. Biol. Chem. 275, 38571-38580, 2000). Lastly, label “d” refers to the inactivating effect of the S139A and N143V mutations in NCX1.1 (homologous to S77 and N81 of NCX_Mj), according to activity assays based on patch-clamp recordings and 45Ca2+ uptake measurements (Ottolia, M. et al. J. Biol. Chem. 280, 1061-1069, 2005). These experiments also showed that N143D and N842D (N81 and D240 in NCX_Mj) are inhibited but regain significant Na+/Ca2+ exchange activity, consistent with our proposal that Smid is not a functional ion-binding site, and that protonation of aspartate residues flanking this site is required for transport (Marinelli, F. et al. Proc. Natl. Acad. Sci. USA 111, E5354-62, 2014; Giladi, M. et al. Sci. Rep. 6, 20753, 2016).
Supplementary Figure 5 Assessment and correction of the CHARMM27 parameters representing the interactions of Ca2+ with water and with carboxyl groups.
(a) Structure of hydrated Ca2+, in terms of the Ca2+-water-oxygen radial distribution function (solid line, black) and the cumulative coordination number (dashed line, red). The data derives from a 20-ns MD simulation at 298 K and 1 bar, using the CHARMM27 force-field and 1,470 water molecules. Experimental ranges from neutron diffraction and EXAFS experiments (Hewish, N.A. et al. Nature 297, 138-139, 1982; Jalilehvand, F. et al. J. Am. Chem. Soc. 123, 431-441, 2001; Fulton, J. L., et al. J. Phys. Chem. A 107, 4688-4696, 2003) are indicated (grey bands) for comparison. (b) Optimization of the CHARMM27 force-field parameters describing the non-bonded interaction between Ca2+ and acetate. The plot shows calculated potential-of-mean-force (PMF) profiles as a function of the distance r between Ca2+ and the central carbon atom in acetate. The corresponding dissociation constants Kd are indicated, along with the experimental values (Daniele, P.G. et al. Coordin. Chem. Revs. 252, 1093-1107, 2008). The PMF was set to zero at a distance of 12 Å. Alternative PMF profiles were computed for different values of the Lennard–Jones Rmin parameter specifically used for the interaction between Ca2+ and the acetate carboxyl–oxygen atoms. The default value in CHARMM27 is 3.067 Å, which results in Kd values that are 3-4 orders of magnitude smaller than the experimental values; the Rmin value used in this study to simulate the Ca2+-bound NCX_Mj is 3.23 Å. To derive the Kd values, the PMF profiles are integrated over the range in r that encompasses both the contact ion-pair (CIP) and the solvent-shared ion-pair (SIP) complexes, i.e., up to Roff = 6.2 Å. (c) Ca2+-water cumulative coordination number in the context of the bi-dentate acetate-Ca2+ CIP, for different values of the Rmin parameter used for the Ca2+-carboxyl–oxygen interaction; the number of water molecules in the first hydration shell is 5, irrespective of the value of Rmin.
Supplementary Figure 6 Assessment of the relative magnitude of ion-charge transfer effects in NCX_Mj and in solution.
Molecular systems considered for (a) Ca2+, (b) Na+ and (c) Sr2+. The value reported, ΔΔq, is the amount of charge (in units of e) that is transferred from each of the cations to the protein, in excess to what is observed for the water clusters (which feature the most probably hydration structure in solution). The results are based on quantum-mechanical calculations of the electronic structure of each system. For Na+, the value indicated in the figure is for the SCa site in the 3 × Na+, occluded state of NCX_Mj. The ΔΔq value for the SCa site in the 2 × Na+ semi-open state is similarly small. The ΔΔq values for the Sext and Sint sites are however ~0.1e.
Supplementary Figure 7 Estimated standard errors in the conformational free-energy landscapes of outward-facing NCX_Mj in different ion-occupancy states.
The two-dimensional maps shown correspond to those in Figs. 5, 6. For each simulation system, the error map was estimated by comparing two free-energy maps calculated from two halves of the simulation data. As shown, the typical statistical error is smaller than 1 kcal/mol, and slightly larger errors are found only in the high-free energy regions.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1 and 2, and Supplementary Notes 1–4 (PDF 3267 kb)
Rights and permissions
About this article
Cite this article
Liao, J., Marinelli, F., Lee, C. et al. Mechanism of extracellular ion exchange and binding-site occlusion in a sodium/calcium exchanger. Nat Struct Mol Biol 23, 590–599 (2016). https://doi.org/10.1038/nsmb.3230
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nsmb.3230
This article is cited by
-
Ca2+ efflux facilitated by co-transport of inorganic phosphate anion in the H+/Ca2+ antiporter YfkE
Communications Biology (2023)
-
Structural insight into the allosteric inhibition of human sodium-calcium exchanger NCX1 by XIP and SEA0400
The EMBO Journal (2023)
-
Calcium dysregulation potentiates wild-type myocilin misfolding: implications for glaucoma pathogenesis
JBIC Journal of Biological Inorganic Chemistry (2022)
-
On the Role of a Conserved Methionine in the Na+-Coupling Mechanism of a Neurotransmitter Transporter Homolog
Neurochemical Research (2022)
-
Salt tolerance involved candidate genes in rice: an integrative meta-analysis approach
BMC Plant Biology (2020)