Key Points
-
S-layers are two-dimensional (2D) protein arrays that are frequently found on the surface of bacteria and archaea.
-
Genetic analysis reveals a wide diversity of genes that encode S-layer proteins (SLPs) in some species, and several mechanisms are found to facilitate gene switching and regulation.
-
Secretion of S-layer proteins often involves a dedicated secretion system, such as accessory Sec systems in Bacillus anthracis and Clostridium difficile, and a wide range of mechanisms for anchoring S-layers to the underlying cell envelope have been identified.
-
Gram-positive species, including B. anthracis and C. difficile, possess large families of genes encoding proteins that are related to the S-layer protein and that share a common anchoring mechanism.
-
In many species, the SLPs are glycosylated. Dedicated glycosylation loci are found that specify all the genes that are necessary for the synthesis of glycan, its secretion across the membrane and ligation to the SLP via N- or O- linkages.
-
S-layers have been the subject of intensive structural analysis since their identification in the 1950s. Recent efforts are gradually improving our high-resolution structural knowledge of various S-layer proteins and finally enabling reasonable quality models of an entire S-layer to be made.
-
SLPs have evolved to mediate a broad range of functions, including biogenesis of the cell wall, control of cell division and specialized activities, such as swimming. In pathogens, SLPs can interfere with the immune system and can aid survival via adhesion to host cells. In Gram-positive bacteria, functions are often associated with an effector domain that can confer properties that are distinct from the ability to form a 2D array.
Abstract
The outer surface of many archaea and bacteria is coated with a proteinaceous surface layer (known as an S-layer), which is formed by the self-assembly of monomeric proteins into a regularly spaced, two-dimensional array. Bacteria possess dedicated pathways for the secretion and anchoring of the S-layer to the cell wall, and some Gram-positive species have large S-layer-associated gene families. S-layers have important roles in growth and survival, and their many functions include the maintenance of cell integrity, enzyme display and, in pathogens and commensals, interaction with the host and its immune system. In this Review, we discuss our current knowledge of S-layer and related proteins, including their structures, mechanisms of secretion and anchoring and their diverse functions.
Similar content being viewed by others
Main
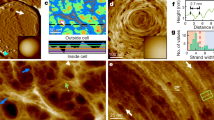
S-layers are found on both Gram-positive and Gram-negative bacteria and are highly prevalent in archaea1,2,3. They are defined as two-dimensional (2D) crystalline arrays that coat the entire cell, and they are thought to provide important functional properties. S-layers consist of one or more (glyco)proteins, known as S-layer proteins (SLPs), that undergo self-assembly to form a regularly spaced array on the surface of the cell. As SLPs are some of the most abundant proteins in the cell2, their biogenesis consumes considerable metabolic resources, which reflects their importance to the organism. S-layers were first recognized in the 1950s and were studied in several species during the following decades, which revealed considerable details of their structures using techniques such as freeze-etch electron microscopy (Box 1). Such studies1,2,4 showed structurally diverse S-layers, with oblique (p1, p2), square (p4) or hexagonal (p6) lattice symmetries. Some Gram-positive species harbour protein families containing SLPs and SLP-related proteins that share a common cell wall-anchoring mechanism. However, it is not yet clear if all family members contribute productively to S-layer self-assembly. These proteins — for example, the Bacillus anthracis S-layer-associated proteins (BSLs) and the Clostridium difficile cell wall proteins (CWPs) (see below) — are included in this Review, as many are functionally relevant. Table 1 outlines all the SLPs and related proteins that are described in this Review.
The lack of S-layers in the model organisms Escherichia coli and Bacillus subtilis hindered their molecular analysis during the 'molecular microbiology' era of the 1980s. Recent advances in genomics and structural biology, together with the development of new molecular cloning tools for many species, have facilitated structural and functional studies of SLPs. Comprehensive reviews about S-layers were written over a decade ago1,2, and others have emphasized the exploitation of SLPs in nanotechnology1,2,5,6. An excellent review of the archaeal cell envelope, which includes the properties of S-layers in this domain of life, was recently published3. In this Review, we discuss the biology of bacterial SLPs and highlight recent discoveries that have shaped our understanding of these important proteins.
Diversity of S-layer genes and proteins
S-layers are usually composed of a single protein, and their structural genes can be linked to genes that encode components of modification or secretory pathways. Despite the apparently conserved function of providing a 2D array surrounding the cell, genetic and functional studies show that there is a wide diversity in both the sequences and the roles of S-layer proteins.
Genetic variation. Several bacterial species show genetic variation in SLP expression; perhaps the best example of this is Campylobacter fetus, in which S-layer variation is very well characterized7. In serotype A strains of C. fetus, the genome contains up to eight surface array A gene (sapA) homologues and one promoter element within a ∼65 kb sap island. Usually only one sap homologue is expressed in culture, although bacterial subpopulations that express additional Sap proteins can be identified. Sap homologues are expressed from a single sap promoter8 and have extensive sequence homology. Broad, high-frequency chromosomal rearrangements that involve DNA inversion and recombination lead to phenotypic switching and expression of alternate sap homologues on the cell surface, which results in an antigenically distinct S-layer8,9,10.
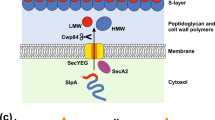
The C. difficile S-layer is composed of two proteins: the high-molecular weight (HMW) SLP and the low-molecular weight (LMW) SLP, both of which are produced by proteolytic cleavage of the precursor S-layer protein SlpA. The LMW SLP, which probably faces the environment, shows considerable sequence variability among strains11,12. This variation affects recognition by antibodies, which presumably reflects pressure from the host immune response. The genetic basis of S-layer diversity in C. difficile was analysed using high-throughput genome sequencing of a panel of clinically diverse strains, which revealed the presence of a ∼10 kb cassette that encodes SlpA, a protein translocase subunit (SecA2) and two CWPs (see below)13. The authors identified 12 divergent cassettes among the strains and proposed that recombinational switching occurs in C. difficile populations to generate antigenic diversity. Interestingly, one of the cassettes is substantially larger than the others (24 kb compared with 10 kb) and contains 19 extra genes that encode a putative glycosylation island (see below). Another C. difficile S-layer protein, cell wall protein V (CwpV), undergoes phase-variable expression14, which is mediated by DNA inversion of an element that is situated between the promoter and the structural gene15.
S-layer gene families. Some firmicutes contain multiple S-layer gene homologues that exhibit varying degrees of sequence identity, which suggests that gene duplication has led to a family of genes that have functional diversity.
The best studied examples of SLP and related protein gene families are found in B. anthracis and C. difficile. On the basis of the presence of three tandem surface layer homology (SLH) motifs (see below) within predicted surface proteins, 24 putative BSLs were identified in B. anthracis Sterne16, including the two major S-layer proteins, Sap and EA1 (Fig. 1). The B. anthracis SLPs are incorporated into the S-layer at different stages of growth; the Sap S-layer is produced during the exponential growth phase and is replaced in the stationary phase by the EA1 S-layer17. Three BSLs are encoded by plasmids: two of these are encoded by pOX1 and one is encoded by pOX216. In each BSL, the three tandemly arranged SLH motifs are located at either the amino terminus or carboxyl terminus of the protein. In some cases, the proteins are considerably larger than the ∼220 residues that are required for the SLH domains; for example, the Sap1 and EA1 proteins are more than 800 residues in length. In some cases, functional effector domains can be identified in these larger proteins, including domains that encode leucine rich repeats (LRRs) and β-lactamase and several domains that are involved in peptidoglycan synthesis and hydrolysis. Bacillus cereus, which is a close relative of B. anthracis, lacks the bslG, bslK and amiA genes of B. anthracis but harbours three unique bsl genes, bslV, bslW, and bslX, which are not found in B. anthracis18. It should be emphasized that these BSLs have not been shown to form 2D arrays; this property is seen only in Sap and EA1 in B. anthracis.
Domain identification and organization among all members of the Clostridium difficile CW_binding_2 (CWB2) family and the Bacillus anthracis surface layer homology (SLH) family were determined using the Pfam protein families database120 and outlined in Supplementary information S1 (table). The amino-terminal signal peptide, which is removed upon translocation through the Sec membrane channel, is shown as a black box (see also Fig. 2a). In C. difficile, the secretion of at least surface layer protein A (SlpA) and cell wall protein V (CwpV) is dependent on the accessory Sec secretion system28. In B. anthracis, the two major S-layer proteins, surface array protein (Sap) and EA1, also require the accessory Sec system for secretion104. Although limited data are available, it is possible that secretion via the accessory Sec system is a common feature of these two protein families.
A remarkably similar situation is found in C. difficile. A number of Clostridia spp. use the cell wall binding 2 (CWB2) domain in an analogous manner to the SLH motifs to anchor the S-layer to the underlying cell wall (see below). In C. difficile, there are 29 CWPs, each of which has three tandem CWB2 domains, including the major SLP, SlpA19. Comparison of the effector domains that are associated with the B. anthracis BSL proteins and the C. difficile CWPs show many similar functions, including peptidoglycan hydrolases, putative adhesins and LRR proteins (Fig. 1). Families of CWB2-containing surface proteins, including some that have effector domains, are also found in Clostridium botulinum and Clostridium tetani20,21.
The large number of SLP paralogues in the Clostridia and Bacilli that carry an effector domain — many of which are predicted to be exposed to the environment — inspires the idea that the S-layer functions as a scaffold to display proteins or glycoproteins to the external environment. In this way, the S-layer can impart various functions to its host (see below), depending on the properties of the protein or glycoprotein that is displayed.
From proteins to functional S-layers
SLPs are transported to the cell surface, where they assemble into the ordered structures of the S-layers. In addition, to build a fully functional S-layer, SLPs are anchored to the cell wall and, in some organisms, are highly glycosylated.
Secretion of S-layer proteins. Translocation of proteins across the cell envelope is an essential process in all bacteria. Secretion of S-layer proteins presents a particular problem for bacteria, owing to the large quantity of protein that is required to form a contiguous paracrystalline array; for example, we estimate that the C. difficile S-layer contains up to 500,000 subunits, which requires the secretion of approximately 140 subunits per second per cell during exponential growth. Several distinct mechanisms have evolved to cope with this high protein flux, but, as S-layer secretion is now being studied in several bacterial species, a number of trends are emerging (Fig. 2).
So far, S-layer secretion has been studied in a small number of species, in which several dedicated secretion systems have been identified. a | In the Gram-positive bacteria Clostridium difficile and Bacillus anthracis, secretion of the S-layer precursors is mediated by the accessory Sec secretion system30. The proteins contain an amino-terminal signal peptide (white box, SP) which directs the nascent polypeptide to the secretion apparatus and is cleaved upon membrane translocation (indicated by the black arrow). In both C. difficile and B. anthracis, translocation requires the accessory ATPase, SecA2 (Refs 27,28). Following recognition by SecA2, the nascent polypeptide is translocated across the membrane through a pore that consists of SecY, SecE and SecG (in C. difficile) or SecY2, SecE and SecG (in B. anthracis). b | Secretion of the S-layer proteins (SLPs) in Aeromonas salmonicida and Aeromonas hydrophila requires a dedicated type II secretion system25,26. Type II secretion is a two-step process: the unfolded precursor is first translocated across the cytoplasmic membrane by the canonical Sec secretion system, the protein then folds and is transported across the outer membrane by a complex multiprotein secretion apparatus that is closely related to type IV pili121. A complete type II secretion system is encoded alongside tetragonal surface virulence array A (vapA) in A. salmonicida, but only one component of this system has been directly linked to SLP secretion; A. salmonicida ApsE is homologous to the secretion ATPase, pullulanase secretion protein PulE. A PulD homologue, S-protein secretion component D (SpsD), which probably forms an outer-membrane pore, has also been identified in A. hydrophila. Further analysis is required to confirm whether or not SpsD and ApsE are from the same conserved secretion system. In Campylobacter fetus, the SLPs are secreted in a single step by a type I secretion system that is encoded by sapDEF24. Type I secretion involves an inner membrane ATP-binding cassette (ABC) transporter, a membrane fusion protein and an outer membrane pore168. Where known, the anchoring domains of the SLPs are highlighted as hatched boxes (see Fig. 1). ApsE, A-protein secretion protein E; CWP, cell wall protein; HMW, high molecular weight; LMW, low molecular weight; Sap, surface array protein.
In many Gram-negative species, including Caulobacter crescentus, Serratia marcescens and C. fetus, S-layer secretion relies on a specific type I secretion system22,23,24 that comprises an inner membrane ATP-binding cassette (ABC) transporter and an outer membrane pore (Fig. 2b). In C. fetus, a four-gene operon that is adjacent to the phase-variable S-layer cassette, encodes three proteins, SapD, SapE and SapF, which have homology to type I secretion systems, and an additional unique protein, SapC. Mutagenesis of this operon blocks S-layer secretion and the four-gene operon is sufficient for the secretion of the S-layer protein SapA in a heterologous host24.
In the fish pathogens Aeromonas hydrophila and Aeromonas salmonicida, S-layer secretion seems to be dependent on specific type II secretion systems (Fig. 2b). The SLPs of both species possess an amino-terminal secretion signal and, in each species, additional secretion proteins that have homology to components of the prototypic pullulanase secretion system — a type II secretion system that is found in Klebsiella spp. — have been identified: A. hydrophila S-protein secretion component D (SpsD) is homologous to pullulanase D (PulD) and A. salmonicida secretion protein ApsE is homologous to PulE. Mutagenesis of either spsD or apsE results in periplasmic accumulation of the SLPs25,26. A. salmonicida ApsE is encoded within a complete type II secretion cluster that is adjacent to the S-layer gene vapA. The other genes in this cluster have not been studied in depth but it seems probable that the encoded type II secretion system is responsible for the secretion of VapA.
S-layer secretion has been studied in detail in two Gram-positive bacteria, B. anthracis and C. difficile (Fig. 2a). In both cases, the secretion of the S-layer precursor is dependent on the accessory Sec secretion system27,28. The accessory Sec secretion system was first identified in Mycobacterium tuberculosis29 and has since been characterized in a small number of Gram-positive species30. Organisms that possess an accessory Sec system have two copies of the ATPase SecA (SecA1 and SecA2); some bacteria, such as B. anthracis, also have an accessory SecY (SecY2). The accessory ATPase, SecA2, is responsible for the secretion of a small subset of proteins. In B. anthracis, efficient secretion of both major S-layer proteins, EA1 and Sap, requires SecA2 and the S-layer assembly protein SlaP27. In C. difficile, the accessory Sec system is responsible for the secretion of the S-layer precursor SlpA and the major phase-variable cell wall protein CwpV28.
Interestingly, there is a striking degree of genetic linkage between the genes that encode S-layer proteins and their dedicated secretion systems in many of the organisms that are described above, including C. difficile, B. anthracis, A. salmonicida, C. fetus and C. crescentus. There is also evidence for horizontal transfer of the gene cassette that contains slpA and secA2 between different lineages of C. difficile13. This emphasizes the importance of the S-layer and its secretion in the life cycle of these organisms. As molecular characterization of S-layers extends to new species, it will be fascinating to see if dedicated secretion systems and tight genetic linkage are common features of S-layer biogenesis.
Anchoring of S-layer proteins to the cell surface. The S-layer is anchored to the cell surface via non-covalent interactions with cell surface structures, usually with lipopolysaccharides (LPSs) in Gram-negative bacteria and with cell wall polysaccharides in Gram-positive bacteria. In general, S-layer anchoring in Gram-negative bacteria is less well characterized than in Gram-positive bacteria. However, the S-layers of C. crescentus and C. fetus have been studied in some detail. In C. crescentus, the amino-terminal ∼225 amino acids of the 98 kDa RsaA SLP is required for binding to LPS on the cell surface31,32. The exact mechanism of this interaction has yet to be characterized, partly because the exact structure of the C. crescentus LPS is unknown; however, the interaction does require an intact O-antigen31. The highly variable C. fetus S-layer also binds non-covalently to LPS. However, C. fetus strains possess one of two distinct LPS serotypes and, consequently, two distinct S-layer-anchoring modules: serotype A is exclusively associated with a SapA-type S-layer and serotype B with a SapB-type S-layer. Each C. fetus genome contains multiple copies of either sapA or sapB, enabling high-frequency antigenic variation (see above). All SapA-type homologues have a highly conserved amino-terminal domain that is responsible for anchoring to serotype A LPS. SapB-type SLPs are similar to SapA in general but have an entirely unrelated amino-terminal domain that anchors the proteins to serotype B LPS10,33,34.
The anchoring of S-layers has been studied in many Gram-positive species. So far, two conserved Gram-positive S-layer-anchoring modules have been identified, which use either the SLH domain or the CWB2 domain. Both modules use three domains, which are located either at the amino-terminal or carboxyl-terminal region of the protein. The SLH domain is the most widely distributed and is found in the SLPs of many Bacillus species, Thermus thermophillus, Deinococcus radiodurans and at least one Clostridia species, Clostridium thermocellum35,36,37,38,39,40. The two major SLPs that are produced by B. anthracis, Sap and EA1, each have three tandem copies of the SLH motif41,42. These motifs fold as a pseudo-trimer43 (Box 1) and cooperatively function to bind to a pyruvylated secondary cell wall polymer (SCWP). Pyruvylation of the SCWP relies on an enzyme, CsaB, which is encoded adjacent to genes that encode Sap and EA1 in the B. anthracis genome40. The thermophilic bacterium Geobacillus stearothermophilus possesses one of the most intensively studied S-layers. G. stearothermophilus strains can produce five different S-layer types, which are encoded by sbsA–sbsD and sgsE44,45,46,47, with two distinct anchoring mechanisms. SbsB has three amino-terminal copies of the SLH domain that anchor the protein to a pyruvylated SCWP48. However, no SLH domains can be identified in the remaining four Geobacillus SLPs. Instead, SbsC has been shown to interact with an N-acetylmannosaminuronic acid-containing SCWP49 via the first 240 residues of the mature SLP50 (Box 1). As these residues are highly conserved in SbsA, SbsD and SgsE, it is probable that these SLPs are anchored by the same mechanism.
The second conserved mechanism involves the CWB2 motif, which was first identified in CwlB — an autolysin that cleaves peptidoglycan in the cell wall of B. subtilis51,52. B. subtilis does not produce an S-layer, but the CWB2 motif is necessary for the retention of CwlB in the cell wall. The CWB2 motif is found in many Clostridia species, including the important human pathogens C. difficile, C. tetani and C. botulinum53. In C. difficile, a family of 29 CWPs, including CwpV and the S-layer precursor SlpA, all use the CWB2 domain for anchoring to the cell wall19. The cell wall ligand for this domain is currently unknown but is likely to be a cell surface polysaccharide that is either free or linked to peptidoglycan. We know little about the specificity of CWB2–polysaccharide interactions. However, surface polysaccharides in the Firmicutes show considerable diversity54, and it is possible that the CWB2 motif recognizes more than one chemical entity or that, in different species, the motif has evolved to recognize a specific polysaccharide. Each cell wall protein has three tandem copies of the CWB2 motif; this is analogous to the arrangement of SLH domains that is seen in other S-layer proteins. As more structural information becomes available, it will be interesting to see whether the pseudo-trimer binding arrangement is also shared between SLH and CWB2 domains, which would suggest that they have a common or convergent evolutionary origin.
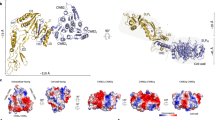
Formation of an ordered array on the cell surface. An S-layer is, by definition, a 2D array of a single protein, but how exactly is the array formed? It is clear that SLPs that form arrays have at least two functional domains: an anchoring domain, such as the tandem SLH or CWB2 motifs, which attaches the protein to the underlying cell wall; and a crystallization domain that mediates the SLP–SLP interaction. Crystallization domains, which may contain several structural domains, have been identified in G. stearothermophilus SbsB55 and SbsC56 and are present in SLPs of other species, including those of B. anthracis57. In a landmark publication58, the 3D crystal structure of SbsB was described, showing the atomic contacts between adjacent 718 residue-long SLP crystallization domains, as well as the way in which individual structural domains within molecules are coordinated by Ca2+ — an anion that is known to be essential for S-layer formation in G. stearothermophilus58 (Box 1). The structure also shows pores of approximately 30 Å in diameter that are formed at the interface between three adjacent subunits, which is consistent with a role in permeability (see below).
It should be noted that not all SLPs have been shown to form 2D arrays, and little is known about the potential for self-assembly of the associated proteins, such as the B. anthracis BSL proteins and C. difficile CWPs. However, these proteins are found within the S-layer and, although they are probably held in place by interactions with cell wall ligands, we cannot rule out that they form lateral interactions with the rest of the S-layer.
Glycosylation of bacterial S-layers. The first description of bacterial protein glycosylation was in the S-layer of Halobacterium salinarium59, and since then, many glycosylated S-layer proteins have been identified in numerous bacterial and archaeal species. S-layer glycosylation has been reviewed in excellent detail elsewhere60,61,62, and only the salient points will be discussed here. S-layer glycan modifications involve sugars that are often found in glycosylated eukaryotic proteins, together with some unusual sugars60 (see below). Although N- and O-linkages have been described in archaeal SLPs3, so far, only O-linkages have been found in bacterial SLPs, despite other bacterial surface proteins having N-linked glycans63. O-linkage in SLPs of the Bacillaceae family can involve serine, threonine or tyrosine. The overall structure and architecture of the S-layer glycan resembles that of Gram-negative LPS, containing a linkage unit and up to 50 repeating units, each of which consists of 2–6 sugars. This resemblance suggests a common evolutionary origin of LPS biosynthesis and S-layer glycosylation64, which is an idea that is further strengthened by recent descriptions of the S-layer glycan (slg) gene clusters in G. stearothermophilus, Paenibacillus alvei, Geobacillus tepidamans, Aneurinibacillus thermoaerophilus and Tannerella forsythia encoding the glycosyltransferases, glycan processing enzymes and membrane transport machinery that are sufficient for glycan biosynthetic pathways (reviewed in Ref. 60). S-layer glycosylation pathways have been described in several species, including G. stearothermophilus65, P. alvei66 and T. forsythia67, which has led to the proposal of a biosynthetic route that involves the transfer of galactose from the nucleotide-activated sugar uridine diphosphate (UDP)-α-D-galactose to a lipid carrier, formation of the linkage unit by the addition of glycans and assembly of the growing repetitive glycan chain onto the linkage unit. These reactions occur in the cytoplasm before transport of the completed glycan chain via an ABC-transporter (in the case of G. stearothermophilus) to the distal side of the membrane, where the glycan is ligated to the S-layer protein substrate60. The glycan chains that decorate S-layer proteins are fairly diverse; for example, in G. stearothermophilus, the glycan chain is a simple polymer of L-rhamnose, in P. alvei, L-rhamnose, N-acetylmannosamine, D-glucose and D-galactose are found, and in T. forsythia the unusual sugars N-acetylmannosaminuronic acid, 5-acetimidol-7-N-glycolylpseudaminic acid and digtoxose are present67. Whether the glycan chain is co-transported with the S-layer protein substrate remains to be determined, but protein transport (that is, secretion) is not dependent on glycosylation.
Strains of C. difficile that contain a putative slg locus adjacent to slpA were recently described13. This bacterium does not normally elaborate a glycosylated S-layer68, but these variant strains contain a distinct S-layer cassette (see above) and produce S-layer proteins of reduced polypeptide length12,13. Whether these strains do indeed have a glycosylated S-layer and what, if any, phenotype that might confer is currently unknown.
Functional heterogeneity of S-layers
It is perhaps not surprising that, as the major proteinaceous surface component of the cell, various functions have been described and proposed for the S-layer1,69 (Fig. 3). However, after decades of research, no single function can be ascribed to the S-layer, and, in many species, the S-layer has no known function. The ability to form a 2D array seems to be the result of convergent evolution and is observed in proteins of quite distinct sequence. SLPs and associated proteins have further evolved to carry out a multitude of functions, some of which are essential to the physiology of the cell and some of which facilite survival in specific niches. Although we do not definitively know the function of many S-layers, it is clear from their wide occurrence in the bacterial domain, apparent convergent evolution and the enormous metabolic load that is required to produce and maintain these structures, that they have an important role in the bacteria that produce them.
Probable roles of S-layer proteins in infection (part a) include adhesin activity, which is found in several species: S-layer- associated protein A (BslA) of Bacillus anthracis binds to HeLa cells and mutants are attenuated in models of infection79, binding of surface layer proteins (SLPs) to enteric cells has been observed in Clostridium difficile75 and binding to defined ligands (such as types I and IV collagen) has been observed in Lactobacillus crispatus70. In C. difficile, interaction of SlpA with host toll-like receptor 4 (TLR4) receptors is linked to innate immunity74 and, in Campylobacter fetus, SLPs prevent binding of complement factor C3b, which protects the bacterium from host-mediated phagocytosis and serum killing86. A role in resistance to predation has been shown in Bdellovibrio bacteriovirus97, and potential roles in resistance to bacteriophage and bacteriocins are possible but remain speculative (part a; lower panel). SLPs have roles in the maintenance of cell envelope integrity (part b) in Deinococcus radiodurans, where inactivation of slpA causes shedding of surface molecules99, and in C. difficile, where loss of Cwp84 results in an abnormal S-layer and the shedding of surface proteins101. B. anthracis BslO and C. difficile Cwp22 have peptidoglycan hydrolase activity that may remodel the peptidoglycan. A role for SLPs as a permeability barrier has been shown in Bacillus coagulans98. A role in cell division has been found in B. anthracis (part c) where inactivation of S-layer associated protein O (BslO), which is a putative N-acetylglucosaminidase, results in increased bacterial chain length103. SLPs have roles in aggregation (such as C. difficile CwpV15), biofilm formation (in Tanerella forsythia94) and swimming (in Synechococcus spp.106). Finally, BslK from B. anthracis mediates iron uptake by scavenging haem and transferring it to iron-regulated surface determinant protein C (IsdC)80.
Roles of the S-layer in pathogenesis and immunity. In Lactobacillus crispatus, which is an indigenous member of the human and chicken gut microflora, the S-layer CsbA (also known as SlpB) has an amino-terminal domain that binds to types I and IV collagen and a carboxyl-terminal domain that interacts with the bacterial cell wall70. This collagen-binding activity is thought to mediate bacterial colonization of the gut (Fig. 3a). Interestingly, two other putative S-layer genes are present in this strain: slpC, which is located downstream of csbA, is transcriptionally active and slpA is silent70,71,72.
As the major surface antigen of C. difficile, the S-layer has been investigated for its ability to be recognized by, and to activate, the immune system. The SlpA proteins (HMW and LMW SLPs) induce the release of pro-inflammatory cytokines from human monocytes and induce the maturation of human monocyte-derived dendritic cells (MDDCs)73. Further work advanced these findings by showing that the SLPs induced the maturation of mouse bone marrow-derived dendritic cells and the production of the cytokines interleukin-12 (IL-12), tumour necrosis factor α (TNFα) and IL-10, but not the production of of IL-1β. Importantly, activation of dendritic cells was dependent on Toll-like receptor 4 (TLR4) and subsequently induced T helper cell responses that are known to be involved in the clearance of bacterial pathogens74. Infection of TLR4-knockout mice resulted in more severe symptoms than in wild-type mice, which suggests that TLR4 has an important role in bacterial clearance74. Both the HMW and LMW SLPs of C. difficile were required for the activation of dendritic cells, which suggests that either the entire complex is recognized by TLR4, or that one of the SLPs is the ligand, but that it requires to be seen in the context of the HMW–LMW complex (Fig. 3a). SlpA has also been shown to bind to fixed gut enterocytes75, which could be related to an immune-stimulatory role and which infers that the S-layer might function as an adhesin in vivo (Fig. 3a). So far, the role of SlpA as an adhesin has not been investigated using mutagenesis, as slpA is an essential gene, but recent advances in genetics76,77,78 should enable the creation of conditional mutant strains that could be used in infection experiments to test this hypothesis. Finally the phase-variable CwpV protein has bacterial auto-aggregation activity that might have a role during infection15.
In B. anthracis, the precise roles of Sap and EA1 are unknown, but activities of some BSLs have been elucidated. The BslA protein has been shown to mediate the adhesion of encapsulated B. anthracis to HeLa cells16. Mutants of bslA show limited dissemination to tissues and decreased virulence in a guinea pig model of infection, which suggests that BslA is a functional adhesin that is required for full virulence of B. anthracis79. Another S-layer protein, BslK, mediates haem uptake by using a near iron transporter (NEAT) domain80 (Fig. 3c). In B. cereus, which has highly similar BSLs to B. anthracis, but a very different capsule composition, a csaB mutant was found to be defective in retaining Sap, EA1 and BslO on the cell wall18. The csaB mutant exhibited reduced virulence in a mouse model of infection, implicating one or more BSLs in the pathogenesis of anthrax.
The Gram-negative pathogen C. fetus is a leading cause of abortions in sheep and cattle and can cause persistent systemic infections in humans. The S-layer, which is composed of the Sap protein, is essential for host colonization81. Sap is antigenically variable (see above) which contributes to the evasion of the host immune response82,83. The S-layer is crucial for pathogenesis, as mutant strains do not cause disease and are sensitive to phagocytosis and killing in serum mediated by complement C3 (Refs 84, 85, 86) (Fig. 3a). Antigenic variation occurs during infection in humans, as shown by strains that were isolated at early and late stages of infection from four individuals with relapsing C. fetus infections87. In three patients, the strain had undergone a switch in the predominant S-layer protein that is expressed.
T. forsythia (which was previously known as Bacteroides forsythus) is a Gram-negative anaerobe that is associated with severe forms of periodontal disease88,89. Strains of T. forsythia produce two HMW proteins, each more than 200 kDa, that are concomitantly expressed and that constitute the S-layer90,91,92. The structural genes tfsA and tfsB encode proteins of 120 kDa and 140 kDa93, which suggests that post-translational modification occurs. This was explored in detail in a study67 that showed a complex pattern of glycosylation on both proteins, including a modified pseudaminic acid Pse5Am7Gc that has not previously been found on bacterial proteins. This sugar, which is a sialic acid derivative, was suggested to participate in bacterium–host interactions67 on the basis of the prevalence of sialic acid-like sugars in Gram-negative structures that are involved in pathogenesis, such as LPS, capsules, pili and flagella. In T. forsythia transposon mutants that exhibited altered biofilm formation (Fig. 3c), an operon that is involved in exopolysaccharide biosynthesis was identified. Mutation of one gene, weeC, which encodes a putative UDP-N-acetyl-D-mannosaminuronic acid dehydrogenase, increased biofilm formation and altered the mobility of the two T. forsythia S-layer proteins on SDS–PAGE — an effect that is consistent with glycosylation94. A proteomics study also found increased quantities of S-layer proteins in Tannerella biofilms compared with planktonic cells95. The T. forsythia S-layer seems to be essential for the virulence of this pathogen as adhesion to, and invasion of, human gingival epithelial cells (Ca9-22 cells) and mouth epidermal carcinoma cells (KB cells) were decreased or abolished in a mutant that was defective in tfsA and tfsB92.
SLPs have also been shown to have a role in protection against predation. The parasitic bacterium Bdellovibrio bacterovirus has a wide host range and infects and replicates in the periplasm of susceptible Gram-negative species96. Strains of Aquaspirillum serpens and C. cresentus possess an S-layer and are not normally parasitized by Bdellovibrio spp.; however, S-layer-negative variants of both species were shown to be sensitive to predation, which indicates that the S-layer can provide a protective coat against infection with Bdellovibrio spp.97 (Fig. 3a). In addition to this, one might speculate that the S-layer would function as a receptor for bacteriophage and bacteriocins but, to our knowledge, there is no evidence of such activities.
Permeability and biogenesis of the cell envelope. S-layers are often proposed to function as a permeability barrier, and a direct role for SLPs as barriers has been investigated in Bacillus coagulans and other species2,98. Experimentally determined exclusion limits of isolated sacculi and associated proteins of 15–34 kDa are consistent with pore diameters of 20–60 Å within the S-layer lattice2,98, and the crystal structure of SbsB reveals pores of approximately 30 Å in diameter58. Thus, a role for the S-layer as a permeability barrier (Fig. 3b) is certainly conceivable, but our knowledge of this activity would be strengthened by genetic analysis and atomic-resolution imaging of alterations in pore size in vivo.
In D. radiodurans, the S-layer (which is known as the hexagonally packed intermediate (HPI) layer) is composed of two S-layer proteins, SlpA and Hpi, in addition to lipids and carbohydrates. Deletion of slpA results in major structural alterations, such as loss of the Hpi protein and surface glycans, which can be visualized by electron microscopy as layers of material peeling off the cell wall99. This suggests that SlpA is involved in the attachment of Hpi and other components to the underlying cell wall, which is consistent with the presence of SLH domains within SlpA. Other examples of a role for the S-layer in cell envelope biogenesis include C. difficile, in which inhibition of cleavage of the S-layer precursor SlpA by inactivation of the protease Cwp84 leads to incorrect assembly of the S-layer and shedding of full-length SlpA from the cell wall100,101 (Fig. 3b). Interestingly, virulence of this cwp84 mutant is not diminished in the hamster model of infection, presumably as gut proteases, such as trypsin, can cleave SlpA, resulting in a fully functional S-layer100.
In B. anthracis, the physiological functions of the Sap, EA1 and Bsl proteins have been investigated. Sap and EA1 both have a peptidoglycan hydrolase activity102, although they lack known functional domains that specify such an activity. BslO, which has putative N-acetylglucosaminidase activity and is localized at the cell septa, has a role in catalysing cell division; bslO mutants exhibit increased cell chain length103 (Fig. 3c). Elongated chains of bacteria are also seen in a sap mutant, and this lack of Sap can be complemented by the addition of purified BslO protein to the culture medium, restoring cell division and reducing the chain length. In wild-type cells, Sap was predominantly visualized on the lateral cell wall, away from the septa, which suggests that B. anthracis controls the spatial deposition of the SLPs in order to ensure correct localization of functional entities such as BslO104. In the C. difficile family of CWPs and SLPs, several proteins potentially mediate peptidoglycan synthesis and remodelling, including Cwp22, which has L,D-transpeptidase activity105.
Other functions of the S-layer. Another function that is associated with the S-layer is swimming in marine Synechococcus spp.106 (Fig 3c). This bacterium is highly motile but does not possess any obvious flagellum or other organelles that might mediate swimming, and the mechanical basis for its motility is mostly a mystery. In mutants that were defective for swimming, two surface proteins that are essential for this activity were identified: SwmA and SwmB107. SwmA is a 130 kDa glycosylated S-layer protein and SwmB is a large 1.12 MDa protein. Other genes that are essential for swimming encode an ABC transporter and several glycosyltransferases108. Although these results indicate another function for an S-layer and suggest that glycosylation of SwmA is required for motility, they unfortunately do not lead us much further in elucidating this highly unusual mechanism of swimming.
Perspectives
Following their discovery in the 1950s and after decades of research, our knowledge of bacterial SLPs has increased considerably in the last few years. It is clear that S-layers do not have one single function, rather a diversity of functions is apparent, and we expect to see new functions revealed as more species are studied. In some archaea, such as Sulfolobus spp., the S-layer seems to be the sole non-lipid constituent of the envelope3, which suggests that structural integrity might be an ancestral function of SLPs. In some bacterial species, such as the Clostridia, the S-layer seems to be essential for cell viability, as unconditional deletion mutants cannot be constructed. In these species, the SLP is the main protein component of the cell surface, although a range of other macromolecules are found.
Increasingly sophisticated and high-resolution techniques, such as atomic force microscopy (AFM) and electron crystallography, are being applied to study S-layer morphology and symmetry109. Ultimately, these techniques will be combined with structural information from X-ray crystallography or nuclear magnetic resonance (NMR) spectroscopy to generate atomic-resolution models of the complete S-layer. Progress has been recently been made with atomic-resolution structures of several SLPs, and we look forward to a complete description of an assembled S-layer structure in complex with the ligand that is responsible for anchoring to the cell wall. Only then will we be able to address the crucial outstanding questions in S-layer biology: what is the biochemical basis of paracrystalline array self-assembly? What mechanisms are employed to attach the S-layer to the cell surface? And finally, what is the structural basis for the known functions of S-layers?
It is clear that SLPs and their associated proteins have evolved to have specialized functions, and in some species of Firmicutes, SLPs function as a scaffold to display enzymes on the cell surface. It is probable that many more SLPs will be identified from genome sequencing, and it will be a challenge to assign meaningful functions to this diverse family of proteins without laboratory investigation. Priorities for future research include establishing the functions of the S-layers that are present in bacterial pathogens, investigating their potential as therapeutic targets for antimicrobial or vaccine development and in-depth structural analyses of the interactions between S-layers and other surface components. As increasingly sophisticated structural and imaging tools are becoming available, we are now in a position to push forward bacterial S-layer research and perhaps determine the full contribution of these fascinating structures to the growth and survival of the bacteria that produce them.
Accession codes
References
Sara, M. & Sleytr, U. B. S-layer proteins. J. Bacteriol. 182, 859–868 (2000).
Sleytr, U. B. & Beveridge, T. J. Bacterial S-layers. Trends Microbiol. 7, 253–260 (1999).
Albers, S. V. & Meyer, B. H. The archaeal cell envelope. Nature Rev. Microbiol. 9, 414–426 (2011).
Sleytr, U. B. & Glauert, A. M. Ultrastructure of the cell walls of two closely related Clostridia that possess different regular arrays of surface subunits. J. Bacteriol. 126, 869–882 (1976).
Sleytr, U. B. et al. S-layers as a tool kit for nanobiotechnological applications. FEMS Microbiol. Lett. 267, 131–144 (2007).
Schuster, B. & Sleytr, U. B. Nanotechnology with S-layer proteins. Methods Mol. Biol. 996, 153–175 (2013).
Tu, Z. C., Wassenaar, T. M., Thompson, S. A. & Blaser, M. J. Structure and genotypic plasticity of the Campylobacter fetus sap locus. Mol. Microbiol. 48, 685–698 (2003).
Dworkin, J. & Blaser, M. J. Nested DNA inversion as a paradigm of programmed gene rearrangement. Proc. Natl Acad. Sci. USA 94, 985–990 (1997).
Tummuru, M. K. & Blaser, M. J. Rearrangement of sapA homologs with conserved and variable regions in Campylobacter fetus. Proc. Natl Acad. Sci. USA 90, 7265–7269 (1993). This paper provides the first description of site-specific recombination between SLP genes as a mechanism for antigenic variation.
Dworkin, J., Tummuru, M. K. & Blaser, M. J. Segmental conservation of sapA sequences in type B Campylobacter fetus cells. J. Biol. Chem. 270, 15093–15101 (1995).
Eidhin, D., Ryan, A., Doyle, R., Walsh, J. B. & Kelleher, D. Sequence and phylogenetic analysis of the gene for surface layer protein, slpA, from 14 PCR ribotypes of Clostridium difficile. J. Med. Microbiol. 55, 69–83 (2006).
Calabi, E. & Fairweather, N. Patterns of sequence conservation in the S-layer proteins and related sequences in Clostridium difficile. J. Bacteriol. 184, 3886–3897 (2002).
Dingle, K. E. et al. Recombinational switching of the Clostridium difficile S-layer and a novel glycosylation gene cluster revealed by large-scale whole-genome sequencing. J. Infect. Dis. 207, 675–686 (2013). This paper provides a description of SLP cassettes in C. difficile , with genetic evidence of recombinational switching, which is hypothesized to facilitate antigenic variation.
Emerson, J. et al. A novel genetic switch controls phase variable expression of CwpV, a Clostridium difficile cell wall protein. Mol. Microbiol. 74, 541–556 (2009).
Reynolds, C. B., Emerson, J. E., de la Riva, L., Fagan, R. P. & Fairweather, N. F. The Clostridium difficile cell wall protein CwpV is antigenically variable between strains, but exhibits conserved aggregation-promoting function. PLoS Pathog. 7, e1002024 (2011).
Kern, J. W. & Schneewind, O. BslA, a pXO1-encoded adhesin of Bacillus anthracis. Mol. Microbiol. 68, 504–515 (2008).
Mignot, T., Mesnage, S., Couture-Tosi, E., Mock, M. & Fouet, A. Developmental switch of S-layer protein synthesis in Bacillus anthracis. Mol. Microbiol. 43, 1615–1627 (2002).
Wang, Y. T., Oh, S. Y., Hendrickx, A. P., Lunderberg, J. M. & Schneewind, O. Bacillus cereus G9241 S-layer assembly contributes to the pathogenesis of anthrax-like disease in mice. J. Bacteriol. 195, 596–605 (2012).
Fagan, R. P. et al. A proposed nomenclature for cell wall proteins of Clostridium difficile. J. Med. Microbiol. 60, 1225–1228 (2011).
Bruggemann, H. et al. The genome sequence of Clostridium tetani, the causative agent of tetanus disease. Proc. Natl Acad. Sci. USA 100, 1316–1321 (2003).
Sebaihia, M. et al. Genome sequence of a proteolytic (Group I) Clostridium botulinum strain Hall A and comparative analysis of the clostridial genomes. Genome Res. 17, 1082–1092 (2007).
Awram, P. & Smit, J. The Caulobacter crescentus paracrystalline S-layer protein is secreted by an ABC transporter (type I) secretion apparatus. J. Bacteriol. 180, 3062–3069 (1998).
Kawai, E., Akatsuka, H., Idei, A., Shibatani, T. & Omori, K. Serratia marcescens S-layer protein is secreted extracellularly via an ATP-binding cassette exporter, the Lip system. Mol. Microbiol. 27, 941–952 (1998).
Thompson, S. A. et al. Campylobacter fetus surface layer proteins are transported by a type I secretion system. J. Bacteriol. 180, 6450–6458 (1998).
Noonan, B. & Trust, T. J. Molecular analysis of an A-protein secretion mutant of Aeromonas salmonicida reveals a surface layer-specific protein secretion pathway. J. Mol. Biol. 248, 316–327 (1995).
Thomas, S. R. & Trust, T. J. A specific PulD homolog is required for the secretion of paracrystalline surface array subunits in Aeromonas hydrophila. J. Bacteriol. 177, 3932–3939 (1995).
Nguyen-Mau, S. M., Oh, S. Y., Kern, V. J., Missiakas, D. M. & Schneewind, O. Secretion genes as determinants of Bacillus anthracis chain length. J. Bacteriol. 194, 3841–3850 (2012).
Fagan, R. P. & Fairweather, N. F. Clostridium difficile has two parallel and essential Sec secretion systems. J. Biol. Chem. 286, 27483–27493 (2011).
Braunstein, M., Brown, A. M., Kurtz, S. & Jacobs, W. R. Jr. Two nonredundant SecA homologues function in Mycobacteria. J. Bacteriol. 183, 6979–6990 (2001).
Feltcher, M. E. & Braunstein, M. Emerging themes in SecA2-mediated protein export. Nature Rev. Microbiol. 10, 779–789 (2012).
Awram, P. & Smit, J. Identification of lipopolysaccharide O antigen synthesis genes required for attachment of the S-layer of Caulobacter crescentus. Microbiology 147, 1451–1460 (2001).
Ford, M. J., Nomellini, J. F. & Smit, J. S-layer anchoring and localization of an S-layer-associated protease in Caulobacter crescentus. J. Bacteriol. 189, 2226–2237 (2007).
Yang, L. Y., Pei, Z. H., Fujimoto, S. & Blaser, M. J. Reattachment of surface array proteins to Campylobacter fetus cells. J. Bacteriol. 174, 1258–1267 (1992).
Dworkin, J., Tummuru, M. K. & Blaser, M. J. A lipopolysaccharide-binding domain of the Campylobacter fetus S-layer protein resides within the conserved N terminus of a family of silent and divergent homologs. J. Bacteriol. 177, 1734–1741 (1995).
Ebisu, S. et al. Conserved structures of cell wall protein genes among protein-producing Bacillus brevis strains. J. Bacteriol. 172, 1312–1320 (1990).
Bowditch, R. D., Baumann, P. & Yousten, A. A. Cloning and sequencing of the gene encoding a 125-kilodalton surface-layer protein from Bacillus sphaericus 2362 and of a related cryptic gene. J. Bacteriol. 171, 4178–4188 (1989).
Faraldo, M. M., de Pedro, M. A. & Berenguer, J. Sequence of the S-layer gene of Thermus thermophilus HB8 and functionality of its promoter in Escherichia coli. J. Bacteriol. 174, 7458–7462 (1992).
Kuen, B., Sleytr, U. B. & Lubitz, W. Sequence analysis of the sbsA gene encoding the 130-kDa surface-layer protein of Bacillus stearothermophilus strain PV72. Gene 145, 115–120 (1994).
Lemaire, M., Miras, I., Gounon, P. & Beguin, P. Identification of a region responsible for binding to the cell wall within the S-layer protein of Clostridium thermocellum. Microbiology 144, 211–217 (1998).
Mesnage, S. et al. Bacterial SLH domain proteins are non-covalently anchored to the cell surface via a conserved mechanism involving wall polysaccharide pyruvylation. EMBO J. 19, 4473–4484 (2000). This paper gives the first description of a cell wall ligand for an SLP — a pyruvylated SCWP as the ligand for non-covalent anchoring of B. anthracis SLPs that contain SLH domains.
Etienne-Toumelin, I., Sirard, J. C., Duflot, E., Mock, M. & Fouet, A. Characterization of the Bacillus anthracis S-layer: cloning and sequencing of the structural gene. J. Bacteriol. 177, 614–620 (1995).
Mesnage, S., Tosi-Couture, E., Mock, M., Gounon, P. & Fouet, A. Molecular characterization of the Bacillus anthracis main S-layer component: evidence that it is the major cell-associated antigen. Mol. Microbiol. 23, 1147–1155 (1997).
Kern, J. et al. Structure of the SLH domains from Bacillus anthracis surface array protein. J. Biol. Chem. 286, 26042–26049 (2011).
Sara, M. et al. Dynamics in oxygen-induced changes in S-layer protein synthesis from Bacillus stearothermophilus PV72 and the S-layer-deficient variant T5 in continuous culture and studies of the cell wall composition. J. Bacteriol. 178, 2108–2117 (1996).
Jarosch, M., Egelseer, E. M., Mattanovich, D., Sleytr, U. B. & Sara, M. S-layer gene sbsC of Bacillus stearothermophilus ATCC 12980: molecular characterization and heterologous expression in Escherichia coli. Microbiology 146, 273–281 (2000).
Egelseer, E. M. et al. Characterization of an S-layer glycoprotein produced in the course of S-layer variation of Bacillus stearothermophilus ATCC 12980 and sequencing and cloning of the sbsD gene encoding the protein moiety. Arch. Microbiol. 177, 70–80 (2001).
Schaffer, C. et al. The surface layer (S-layer) glycoprotein of Geobacillus stearothermophilus NRS 2004/3a. Analysis of its glycosylation. J. Biol. Chem. 277, 6230–6239 (2002).
Mader, C., Huber, C., Moll, D., Sleytr, U. B. & Sara, M. Interaction of the crystalline bacterial cell surface layer protein SbsB and the secondary cell wall polymer of Geobacillus stearothermophilus PV72 assessed by real-time surface plasmon resonance biosensor technology. J. Bacteriol. 186, 1758–1768 (2004).
Schaffer, C. et al. The diacetamidodideoxyuronic-acid-containing glycan chain of Bacillus stearothermophilus NRS 2004/3a represents the secondary cell-wall polymer of wild-type B. stearothermophilus strains. Microbiology 145, 1575–1583 (1999).
Ferner-Ortner, J., Mader, C., Ilk, N., Sleytr, U. B. & Egelseer, E. M. High-affinity interaction between the S-layer protein SbsC and the secondary cell wall polymer of Geobacillus stearothermophilus ATCC 12980 determined by surface plasmon resonance technology. J. Bacteriol. 189, 7154–7158 (2007).
Kuroda, A., Rashid, M. H. & Sekiguchi, J. Molecular cloning and sequencing of the upstream region of the major Bacillus subtilis autolysin gene: a modifier protein exhibiting sequence homology to the major autolysin and the spoIID product. J. Gen. Microbiol. 138, 1067–1076 (1992).
Kuroda, A. & Sekiguchi, J. Cloning, sequencing and genetic mapping of a Bacillus subtilis cell wall hydrolase gene. J. Gen. Microbiol. 136, 2209–2216 (1990).
Bruggemann, H. & Gottschalk, G. Comparative genomics of Clostridia: link between the ecological niche and cell surface properties. Ann. NY Acad. Sci. 1125, 73–81 (2008).
Weidenmaier, C. & Peschel, A. Teichoic acids and related cell-wall glycopolymers in Gram-positive physiology and host interactions. Nature Rev. Microbiol. 6, 276–287 (2008).
Pavkov, T. et al. The structure and binding behavior of the bacterial cell surface layer protein SbsC. Structure 16, 1226–1237 (2008).
Runzler, D., Huber, C., Moll, D., Kohler, G. & Sara, M. Biophysical characterization of the entire bacterial surface layer protein SbsB and its two distinct functional domains. J Biol Chem, 7, 5207–5215 (2003).
Mignot, T. et al. Distribution of S-layers on the surface of Bacillus cereus strains: phylogenetic origin and ecological pressure. Environ. Microbiol. 3, 493–501 (2001).
Baranova, E. et al. SbsB structure and lattice reconstruction unveil Ca2+ triggered S-layer assembly. Nature 487, 119–122 (2012). This paper provides a high-resolution model of the assembled G. stearothermophilus SbsB S-layer, extrapolated from X-ray crystallography. It also gives a plausible explanation for the calcium-dependence of SLP self-assembly.
Mescher, M. F., Strominger, J. L. & Watson, S. W. Protein and carbohydrate composition of the cell envelope of Halobacterium salinarium. J. Bacteriol. 120, 945–954 (1974).
Ristl, R. et al. The S-layer glycome — adding to the sugar coat of bacteria. Int J. Microbiol 2011, 127870 (2011). This study provides the first complete description of a pathway for glycosylation of an S-layer protein.
Messner, P., Steiner, K., Zarschler, K. & Schaffer, C. S-layer nanoglycobiology of bacteria. Carbohydr. Res. 343, 1934–1951 (2008).
Abu-Qarn, M., Eichler, J. & Sharon, N. Not just for eukarya anymore: protein glycosylation in bacteria and archaea. Curr. Opin. Struct. Biol. 18, 544–550 (2008).
Benz, I. & Schmidt, M. A. Never say never again: protein glycosylation in pathogenic bacteria. Mol. Microbiol. 45, 267–276 (2002).
Schaffer, C., Wugeditsch, T., Neuninger, C. & Messner, P. Are S-layer glycoproteins and lipopolysaccharides related? Microb. Drug Resist. 2, 17–23 (1996).
Steiner, K. et al. Molecular basis of S-layer glycoprotein glycan biosynthesis in Geobacillus stearothermophilus. J. Biol. Chem. 283, 21120–21133 (2008).
Zarschler, K., Janesch, B., Zayni, S., Schaffer, C. & Messner, P. Construction of a gene knockout system for application in Paenibacillus alvei CCM 2051T, exemplified by the S-layer glycan biosynthesis initiation enzyme WsfP. Appl. Environ. Microbiol. 75, 3077–3085 (2009).
Posch, G. et al. Characterization and scope of S-layer protein O-glycosylation in Tannerella forsythia. J. Biol. Chem. 286, 38714–38724 (2011).
Qazi, O. et al. Mass spectrometric analysis of the S-layer proteins from Clostridium difficile demonstrates the absence of glycosylation. J. Mass Spectrom. 44, 368–374 (2009).
Beveridge, T. J. et al. Functions of S-layers. FEMS Microbiol. Rev. 20, 99–149 (1997).
Sun, Z. et al. Characterization of a S-layer protein from Lactobacillus crispatus K313 and the domains responsible for binding to cell wall and adherence to collagen. Appl. Microbiol. Biotechnol. 97, 1941–1952 (2013).
Sillanpaa, J. et al. Characterization of the collagen-binding S-layer protein CbsA of Lactobacillus crispatus. J. Bacteriol. 182, 6440–6450 (2000).
Toba, T. et al. A collagen-binding S-layer protein in Lactobacillus crispatus. Appl. Environ. Microbiol. 61, 2467–2471 (1995).
Ausiello, C. M. et al. Surface layer proteins from Clostridium difficile induce inflammatory and regulatory cytokines in human monocytes and dendritic cells. Microbes Infect. 8, 2640–2646 (2006).
Ryan, A. et al. A role for TLR4 in Clostridium difficile infection and the recognition of surface layer proteins. PLoS Pathog 7, e1002076 (2011). This study shows that an SLP can function as a ligand for the innate immune response, activating TLR4 and promoting clearance of C. difficile infection.
Calabi, E., Calabi, F., Phillips, A. D. & Fairweather, N. Binding of Clostridium difficile surface layer proteins to gastrointestinal tissues. Infect. Immun. 70, 5770–5778 (2002).
Faulds-Pain, A. & Wren, B. W. Improved bacterial mutagenesis by high-frequency allele exchange, demonstrated in Clostridium difficile and Streptococcus suis. Appl. Environ. Microbiol. 79, 4768–4771 (2013).
Cartman, S. T., Kelly, M. L., Heeg, D., Heap, J. T. & Minton, N. P. Precise manipulation of the Clostridium difficile chromosome reveals a lack of association between tcdC genotype and toxin production. Appl. Environ. Microbiol. 78, 4683–4690 (2012).
Heap, J. T. et al. Integration of DNA into bacterial chromosomes from plasmids without a counter-selection marker. Nucleic Acids Res. 40, e59 (2012).
Kern, J. & Schneewind, O. BslA, the S-layer adhesin of B. anthracis, is a virulence factor for anthrax pathogenesis. Mol. Microbiol. 75, 324–332 (2010).
Tarlovsky, Y. et al. A Bacillus anthracis S-layer homology protein that binds heme and mediates heme delivery to IsdC. J. Bacteriol. 192, 3503–3511 (2010).
Grogono-Thomas, R., Dworkin, J., Blaser, M. J. & Newell, D. G. Roles of the surface layer proteins of Campylobacter fetus subsp. fetus in ovine abortion. Infect. Immun. 68, 1687–1691 (2000).
Garcia, M. M. et al. Protein shift and antigenic variation in the S-layer of Campylobacter fetus subsp. venerealis during bovine infection accompanied by genomic rearrangement of sapA homologs. J. Bacteriol. 177, 1976–1980 (1995).
Wang, E., Garcia, M. M., Blake, M. S., Pei, Z. & Blaser, M. J. Shift in S-layer protein expression responsible for antigenic variation in Campylobacter fetus. J. Bacteriol. 175, 4979–4984 (1993).
Blaser, M. J. & Pei, Z. Pathogenesis of Campylobacter fetus infections: critical role of high-molecular-weight S-layer proteins in virulence. J. Infect. Dis. 167, 372–377 (1993).
Blaser, M. J. et al. Pathogenesis of Campylobacter fetus infections: serum resistance associated with high-molecular-weight surface proteins. J. Infect. Dis. 155, 696–706 (1987).
Blaser, M. J., Smith, P. F., Repine, J. E. & Joiner, K. A. Pathogenesis of Campylobacter fetus infections. Failure of encapsulated Campylobacter fetus to bind C3b explains serum and phagocytosis resistance. J. Clin. Invest. 81, 1434–1444 (1988).
Tu, Z. C., Gaudreau, C. & Blaser, M. J. Mechanisms underlying Campylobacter fetus pathogenesis in humans: surface-layer protein variation in relapsing infections. J. Infect. Dis. 191, 2082–2089 (2005).
Darveau, R. P. Periodontitis: a polymicrobial disruption of host homeostasis. Nature Rev. Microbiol. 8, 481–490 (2010).
Socransky, S. S., Haffajee, A. D., Cugini, M. A., Smith, C. & Kent, R. L. Jr. Microbial complexes in subgingival plaque. J. Clin. Periodontol 25, 134–144 (1998).
Sabet, M., Lee, S. W., Nauman, R. K., Sims, T. & Um, H. S. The surface (S-) layer is a virulence factor of Bacteroides forsythus. Microbiology 149, 3617–3627 (2003).
Higuchi, N. et al. Localization of major, high molecular weight proteins in Bacteroides forsythus. Microbiol. Immunol. 44, 777–780 (2000).
Sakakibara, J. et al. Loss of adherence ability to human gingival epithelial cells in S-layer protein-deficient mutants of Tannerella forsythensis. Microbiology 153, 866–876 (2007).
Lee, S. W. et al. Identification and characterization of the genes encoding a unique surface (S-) layer of Tannerella forsythia. Gene 371, 102–111 (2006).
Honma, K., Inagaki, S., Okuda, K., Kuramitsu, H. K. & Sharma, A. Role of a Tannerella forsythia exopolysaccharide synthesis operon in biofilm development. Microb. Pathog. 42, 156–166 (2007).
Pham, T. K. et al. A quantitative proteomic analysis of biofilm adaptation by the periodontal pathogen Tannerella forsythia. Proteomics 10, 3130–3141 (2010).
Sockett, R. E. Predatory lifestyle of Bdellovibrio bacteriovorus. Annu. Rev. Microbiol. 63, 523–539 (2009).
Koval, S. F. & Hynes, S. H. Effect of paracrystalline protein surface layers on predation by Bdellovibrio bacteriovorus. J. Bacteriol. 173, 2244–2249 (1991).
Sara, M. & Sleytr, U. B. Molecular sieving through S layers of Bacillus stearothermophilus strains. J. Bacteriol. 169, 4092–4098 (1987).
Rothfuss, H., Lara, J. C., Schmid, A. K. & Lidstrom, M. E. Involvement of the S-layer proteins Hpi and SlpA in the maintenance of cell envelope integrity in Deinococcus radiodurans R1. Microbiology 152, 2779–2787 (2006).
Kirby, J. M. et al. Cwp84, a surface-associated cysteine protease, plays a role in the maturation of the surface layer of Clostridium difficile. J. Biol. Chem. 284, 34666–34673 (2009).
de la Riva, L., Willing, S. E., Tate, E. W. & Fairweather, N. F. Roles of cysteine proteases Cwp84 and Cwp13 in biogenesis of the cell wall of Clostridium difficile. J. Bacteriol. 193, 3276–3285 (2011).
Ahn, J. S., Chandramohan, L., Liou, L. E. & Bayles, K. W. Characterization of CidR-mediated regulation in Bacillus anthracis reveals a previously undetected role of S-layer proteins as murein hydrolases. Mol. Microbiol. 62, 1158–1169 (2006).
Anderson, V. J., Kern, J. W., McCool, J. W., Schneewind, O. & Missiakas, D. The SLH-domain protein BslO is a determinant of Bacillus anthracis chain length. Mol. Microbiol. 81, 192–205 (2011).
Kern, V. J., Kern, J. W., Theriot, J. A., Schneewind, O. & Missiakas, D. Surface-layer (S-layer) proteins sap and EA1 govern the binding of the S-layer-associated protein BslO at the cell septa of Bacillus anthracis. J. Bacteriol. 194, 3833–3840 (2012).
Peltier, J. et al. Clostridium difficile has an original peptidoglycan structure with a high level of N-acetylglucosamine deacetylation and mainly 3–3 cross-links. J. Biol. Chem. 286, 29053–29062 (2011).
Brahamsha, B. An abundant cell-surface polypeptide is required for swimming by the nonflagellated marine cyanobacterium Synechococcus. Proc. Natl Acad. Sci. USA 93, 6504–6509 (1996).
McCarren, J. & Brahamsha, B. SwmB, a 1.12-megadalton protein that is required for nonflagellar swimming motility in Synechococcus. J. Bacteriol. 189, 1158–1162 (2007).
McCarren, J. & Brahamsha, B. Swimming motility mutants of marine Synechococcus affected in production and localization of the S-layer protein SwmA. J. Bacteriol. 191, 1111–1114 (2009).
Norville, J. E. et al. 7Å projection map of the S-layer protein sbpA obtained with trehalose-embedded monolayer crystals. J. Struct. Biol. 160, 313–323 (2007).
Houwink, A. L. A macromolecular mono-layer in the cell wall of Spirillum spec. Biochim. Biophys. Acta 10, 360–366 (1953). This paper provides the first description of a paracrystalline layer on a bacterial cell.
Severs, N. J. Freeze-fracture electron microscopy. Nature Protoc. 2, 547–576 (2007).
Sleytr, U. B., Messner, P., Pum, D. & Sara, M. Crystalline bacterial cell surface layers (S Layers): from supramolecular cell structure to biomimetics and nanotechnology. Angew. Chem. Int. Ed. 38, 1034–1054 (1999).
Lupas, A. et al. Domain structure of the Acetogenium kivui surface-layer revealed by electron crystallography and sequence-analysis. J. Bacteriol. 176, 1224–1233 (1994).
Dorobantu, L. S., Goss, G. G. & Burrell, R. E. Atomic force microscopy: a nanoscopic view of microbial cell surfaces. Micron 43, 1312–1322 (2012).
Chung, S., Shin, S. H., Bertozzi, C. R. & De Yoreo, J. J. Self-catalyzed growth of S layers via an amorphous-to-crystalline transition limited by folding kinetics. Proc. Natl Acad. Sci. USA 107, 16536–16541 (2010).
Stetefeld, J. et al. Crystal structure of a naturally occurring parallel right-handed coiled coil tetramer. Nature Struct. Biol. 7, 772–776 (2000).
Jing, H. et al. Archaeal surface layer proteins contain β propeller, PKD, and β helix domains and are related to metazoan cell surface proteins. Structure 10, 1453–1464 (2002).
Arbing, M. A. et al. Structure of the surface layer of the methanogenic archaean Methanosarcina acetivorans. Proc. Natl Acad. Sci. USA 109, 11812–11817 (2012).
Fagan, R. P. et al. Structural insights into the molecular organization of the S-layer from Clostridium difficile. Mol. Microbiol. 71, 1308–1322 (2009).
Punta, M. et al. The Pfam protein families database. Nucleic Acids Res. 40, D290–D301 (2012).
Korotkov, K. V., Sandkvist, M. & Hol, W. G. The type II secretion system: biogenesis, molecular architecture and mechanism. Nature Rev. Microbiol. 10, 336–351 (2012).
Acknowledgements
Work in the N.F.F. laboratory is currently supported by the MRC (grants G0800170 and G1001721), the Wellcome Trust (grant 090969Z) and the Leverhulme Trust (grant RF2012-232), and work in the R.P.F. laboratory is supported by the University of Sheffield, UK.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary information S1 (table)
Identification of putative functional domains in the B. anthracis and C. difficile cell surface protein families (PDF 206 kb)
Glossary
- Phase-variable expression
-
The random variation of gene expression in a bacterial population. Expression in individual cells is either on or off, leading to phenotypic heterogeneity in the population.
- Type I secretion system
-
A sec-independent protein secretion system in Gram-negative bacteria. It consists of an inner membrane ATP-binding cassette (ABC) transporter, a periplasmic membrane fusion protein and an outer membrane pore.
- Type II secretion systems
-
sec-dependent multiprotein secretion systems in Gram-negative bacteria; they are closely related to type IV pili.
- Secondary cell wall polymer
-
(SCWP). A carbohydrate-based polymer, other than peptidoglycan and anionic polymers, that is present in the cell wall — for example, the pyruvylated Bacillus anthracis SCWP that anchors the S-layer proteins EA1 and Sap to the cell wall.
- N- and O-linkages
-
Linkage of a sugar to the nitrogen (N) atom of asparagine or to the oxygen (O) atom of serine, threonine or tyrosine.
- Sacculi
-
The sacs of polymerized peptidoglycan that surround bacteria. When they are isolated from the bacterium, they retain the shape of the cell.
Rights and permissions
About this article
Cite this article
Fagan, R., Fairweather, N. Biogenesis and functions of bacterial S-layers. Nat Rev Microbiol 12, 211–222 (2014). https://doi.org/10.1038/nrmicro3213
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrmicro3213
This article is cited by
-
Prokaryotic microvesicles Ortholog of eukaryotic extracellular vesicles in biomedical fields
Cell Communication and Signaling (2024)
-
Cell cycle dependent coordination of surface layer biogenesis in Caulobacter crescentus
Nature Communications (2024)
-
Shape control in 2D molecular nanosheets by tuning anisotropic intermolecular interactions and assembly kinetics
Nature Communications (2023)
-
Previously uncharacterized rectangular bacterial structures in the dolphin mouth
Nature Communications (2023)
-
Role of cell-substrate association during plant biomass solubilization by the extreme thermophile Caldicellulosiruptor bescii
Extremophiles (2023)