Key Points
-
The rationale for using microRNAs (miRNAs) as anticancer drugs is based on two major findings: miRNA expression is deregulated in cancer compared with normal tissues and the cancer phenotype can be changed by targeting miRNA expression.
-
One of the most appealing properties of miRNAs as therapeutic agents is their ability to target multiple genes, frequently in the context of a network, making them highly efficient in regulating distinct biological cell processes relevant to normal and malignant cell homeostasis
-
There are two main strategies to target miRNA expression in cancer. Direct strategies involve the use of oligonucleotides or virus-based constructs to either block the expression of an oncogenic miRNA or to substitute for the loss of expression of a tumour suppressor miRNA. The indirect strategy involves the use of drugs to modulate miRNA expression by targeting their transcription and their processing.
-
Several in vitro and in vivo studies using locked nucleic acid (LNA) antimiR have shown feasibility and high efficiency of this approach. A Phase I trial in humans using LNA anti-miR-122 is ongoing. This study will provide valuable information about pharmacokinetics and safety profiles.
-
The challenges for developing miRNA-based therapeutics are the same as for siRNA therapeutics and comprise delivery, potential off-target effects and safety concerns.
-
Reprogramming aberrant miRNA networks in cancer could be achieved by modulating several of the key miRNAs in a network using known drugs, including chemotherapy agents or biocompounds.
-
miRNA effects are currently largely interpreted as the result of miRNA–mRNA 3′untranslated region (UTR) interactions that cause target post-translational inhibition or degradation. However, focusing on this mechanism to design miRNA therapeutics is likely to prove too simplistic, owing to the emerging miRNA mechanisms, which include decoy activity and 5′ UTR and direct DNA regulatory activities.
Abstract
MicroRNAs (miRNAs) are evolutionarily conserved small non-coding RNAs that regulate gene expression. Early studies have shown that miRNA expression is deregulated in cancer and experimental data indicate that cancer phenotypes can be modified by targeting miRNA expression. Based on these observations, miRNA-based anticancer therapies are being developed, either alone or in combination with current targeted therapies, with the goal to improve disease response and increase cure rates. The advantage of using miRNA approaches is based on its ability to concurrently target multiple effectors of pathways involved in cell differentiation, proliferation and survival. In this Review, we describe the role of miRNAs in tumorigenesis and critically discuss the rationale, the strategies and the challenges for the therapeutic targeting of miRNAs in cancer.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Ambros, V. The functions of animal microRNAs. Nature 431, 350–355 (2004).
Rodriguez, A., Griffiths-Jones, S., Ashurst, J. L. & Bradley, A. Identification of mammalian microRNA host genes and transcription units. Genome Res. 14, 1902–1910 (2004).
Lee, Y. et al. MicroRNA genes are transcribed by RNA polymerase II. EMBO J. 23, 4051–4060 (2004).
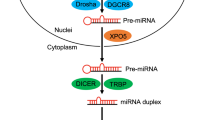
Lee, Y., Ahn, C., Han, J., Choi, H. & Kim, J. The nuclear RNase III Drosha initiates microRNA processing. Nature 425, 415–419 (2003).
Landthaler, M., Yalcin, A. & Tuschl, T. The human DiGeorge syndrome critical region gene 8 and its D. melanogaster homolog are required for miRNA biogenesis. Curr. Biol. 14, 2162–2167 (2004).
Bohnsack, M. T., Czaplinski, K. & Gorlich, D. Exportin 5 is a RanGTP-dependent dsRNA-binding protein that mediates nuclear export of pre-miRNAs. RNA 10, 185–191 (2004).
Hammond, S., Bernstein, E., Beach, D. & Hannon, G. An RNA-directed nuclease mediates post-transcriptional gene silencing in Drosophila cells. Nature 404, 293–229 (2000).
Thimmaiah, P. et al. TRBP recruits the Dicer complex to Ago2 for microRNA processing and gene silencing. Nature 436, 740–744 (2005).
Hutvanger, G. & Zamore, P. D. A microRNA in a multiple-turnover RNAi enxyme complex. Science 297, 2056–2060 (2002).
Bartel, D. P. MicroRNAs: target recognition and regulatory functions. Cell 136, 215–233 (2009).
Lewis, B., Shih, I., Jones-Rhoades, M., Bartel, D. & Burge, C. Prediction of mammalian microRNA targets. Cell 115, 787–798 (2003).
Krek, D. et al. Combinatorial microRNA target predictions. Nature Genet. 37, 495–500 (2005).
Betel, D., Wilson, M., Gabow, A., Marks, D. S. & Sander, C. The microRNA.org resource: targets and expression. Nucleic Acids Res. 36, D149–D153 (2008).
Friedman, R. C., Farh, K. K., Burge, C. B. & Bartel, D. P. Most mammalian mRNAs are conserved targets of microRNAs. Genome Res. 19, 92–105 (2009).
Ivanovska, I. & Cleary, M. A. Combinatorial microRNAs: working together to make a difference. Cell Cycle 20, 3137–3142 (2008).
Stark, A. et al. Discovery of functional elements in 12 Drosophila genomes using evolutionary signatures. Nature 450, 219–232 (2007).
Ørom, U. A., Nielsen, F. N. & Lund, A. H. MicroRNA-10a binds the 5′UTR of ribosomal protein mRNAs and enhances their translation. Mol. Cell 4, 460–471 (2008).
Eiring, A. et al. miR-328 functions as an RNA decoy to modulate hnRNP E2 regulation of mRNA translation in leukemic blasts. Cell 140, 652–665 (2010). First evidence of miRNA decoy activity in gene regulation.
Beitzinger, M. & Meister, G. MicroRNAs: from decay to decoy. Cell 140, 612–614 (2010).
Khraiwesh, B. et al. Transcriptional control of gene expression by microRNAs. Cell 140, 112–122 (2010). First report indicating transcriptional control of gene expression by miRNAs.
Gonzalez, S., Pisano, D. G. & Serrano, M. Mechanistic principles of chromatin remodeling guided by siRNAs and miRNAs. Cell Cycle 7, 2601–2608 (2008).
Kim, D. H., Saetrom, P., Snøve, O. Jr & Rossi, J. J. MicroRNA directed transcriptional gene silencing in mammalian cells. Proc. Natl Acad. Sci. USA 105, 16230–16235 (2008).
Calin, G. A. et al. Frequent deletions and downregulation of micro-RNA genes miR15 and miR16 at 13q14 in chronic lymphocytic leukemia. Proc. Natl Acad. Sci. USA 99, 15524–15529 (2002). First report of miRNA deregulation in cancer.
Lu, J. et al. MicroRNA expression profiles classify human cancers. Nature 435, 834–838 (2005).
Volinia, S. et al. A microRNA expression signature in human solid tumors defines cancer targets. Proc. Natl Acad. Sci. USA 103, 2257–2261 (2005). Comprehensive analysis of miRNA deregulation in solid tumours.
Ota, A. et al. Identification and characterization of a novel gene, C13orf25, as a target for 13q31–q32 amplification in malignant lymphoma. Cancer Res. 64, 3087–3095 (2004).
Petrocca, F. et al. E2F1-regulated microRNAs impair TGFβ-dependent cell-cycle arrest and apoptosis in gastric cancer. Cancer Cell 13, 272–286 (2008).
Yanaihara, N. et al. Unique microRNA molecular profiles in lung cancer diagnosis and prognosis. Cancer Cell 9, 189–198 (2006).
Diosdado, B. et al. MiR-17-92 cluster is associated with 13q gain and c-myc expression during colorectal adenoma to adenocarcinoma progression. Br. J. Cancer 101, 707–714 (2009).
Calin, G. A. et al. A microRNA signature associated with prognosis and progression in chronic lymphocytic leukemia. N. Engl. J. Med. 353, 1793–1801 (2005).
Iorio, M. V. et al. MicroRNA gene expression deregulation in human breast cancer. Cancer Res. 65, 7065–7070 (2005).
Garzon, R. et al. MicroRNA signatures associated with cytogenetics and prognosis in acute myeloid leukemia. Blood 111, 3183–3189 (2008).
Schetter, A. J. et al. MicroRNA expression profiles associated with prognosis and therapeutic outcome in colon adenocarcinoma. JAMA 299, 425–436 (2008).
Ciafre, S. A. et al. Extensive modulation of a set of microRNAs in primary glioblastoma. Biochem. Biophys. Res. Commun. 334, 1351–1358 (2005).
Bloomston, M. et al. MicroRNA expression patterns to differentiate pancreatic adenocarcinoma from normal pancreas and chronic pancreatitis. JAMA 297, 1901–1908 (2007).
Pichiorri, F. et al. MicroRNAs regulate critical genes associated with multiple myeloma pathogenesis. Proc. Natl Acad. Sci. USA 105, 12885–12890 (2008).
Metzler, M. et al. High expression of precursor microRNA-155/BIC RNA in children with Burkitt lymphoma. Genes Chromosomes Cancer 39, 167–169 (2004).
Kluiver, J. et al. BIC and miR-155 are highly expressed in Hodgkin, primary mediastinal and diffuse large B cell lymphomas. J. Pathol. 207, 243–249 (2005).
Garzon, R. et al. Distinctive miRNA signature of acute myeloid leukemia bearing cytoplasmic mutated nucleophosmin. Proc. Natl Acad. Sci. USA 105, 3945–3950 (2008).
Wang, H. et al. NF-κB–YY1–miR-29 regulatory circuitry in skeletal myogenesis and rhabdomyosarcoma. Cancer Cell 14, 369–381 (2008).
Mott, J. L., Kobayashi, S., Bronk, S. F. & Gores, G. J. Mir-29 regulates Mcl-1 protein expression and apoptosis. Oncogene 26, 6133–6140 (2007).
Xiong, Y. et al. Effects of microRNA-29 on apoptosis, tumorigenicity, and prognosis of hepatocellular carcinoma. Hepatology 51, 836–845 (2010).
Zhao, J. J. et al. MicroRNA expression profile and identification of miR-29 as a prognostic marker and pathogenetic factor by targeting CDK6 in mantle cell lymphoma. Blood 115, 2630–2639 (2010).
Bonci, D. et al. The miR-15a–miR-16-1 cluster controls prostate cancer by targeting multiple oncogenic activities. Nature Med. 14, 1271–1277 (2008).
Bottoni, A. et al. MiR-15a and miR-16-1 down-regulation in pituitary adenomas. J. Cell Physiol. 204, 280–285 (2005).
Johnson, S. M. et al. RAS is regulated by the let-7 microRNA family. Cell 120, 635–647 (2005).
Takamizawa, J. et al. Reduced expression of the let-7 microRNAs in human lung cancers in association with shortened postoperative survival. Cancer Res. 64, 3753–3756 (2004).
Yu, F. et al. let-7 regulates self renewal and tumorigenicity of breast cancer cells. Cell 131, 1109–1123 (2007).
Akao, Y., Nakagawa, Y. & Naoe, T. let-7 microRNA functions as a potential growth suppressor in human colon cancer cells. Biol. Pharm. Bull. 29, 903–906 (2006).
Yang, N. et al. MicroRNA microarray identifies let-7i as a novel biomarker and therapeutic target in human epithelial ovarian cancer. Cancer Res. 68, 10307–10314 (2008).
Motoyama, K. et al. Clinical significance of high mobility group A2 in human gastric cancer and its relationship to let-7 microRNA family. Clin. Cancer Res. 14, 2334–2340 (2008).
Calin, G. A. et al. Human microRNA genes are frequently located at fragile sites and genomic regions involved in cancers. Proc. Natl Acad. Sci. USA 101, 2999–3004 (2004). A systematic analysis of miRNA localization and the fragile sites associated with cancer.
Saito, Y. et al. Specific activation of microRNAs-127 with downregulation of the proto-oncogene BCL6 by chromatin-modifying drugs in human cancer cells. Cancer Cell 9, 435–443 (2006). First evidence that miRNAs are regulated by epigenetic mechanisms in cancer.
Lujambio, A. et al. Genetic unmasking of an epigenetically silenced microRNA in human cancer cells. Cancer Res. 67, 1424–1429 (2007).
Hackanson, B. et al. Epigenetic modification of CCAAT/enhancer binding protein α expression in acute myeloid leukemia. Cancer Res. 68, 3142–3151 (2008).
Kumar, M. S. et al. Dicer1 functions as a haploinsufficient tumor suppressor. Genes Dev. 23, 2700–2704 (2009).
Merritt, WA. et al. Dicer, drosha, and outcomes in patients with ovarian cancer. N. Engl. J. Med. 359, 2641–2650 (2008).
He, L. et al. A microRNA component of the p53 tumour suppressor network. Nature 447, 1130–1134 (2007).
Chang, T. C. et al. Transactivation of miR-34a by p53 broadly influences gene expression and promotes apoptosis. Mol. Cell 26, 745–752 (2007). References 58 and 59 report a link between p53 activity and miRNA expression.
Chang, T. S. et al. Widespread microRNA repression by Myc contributes to tumorigenesis. Nature Genetics 40, 43–50 (2007). Comprehensive analysis of MYC-regulated miRNAs in cancer cells.
Cimmino, A. et al. miR-15 and miR-16 induce apoptosis by targeting BCL2. Proc. Natl Acad. Sci. USA 102, 13944–13949 (2006).
Klein, U. et al. The DLEU2/miR-15a/16-11 cluster controls B cell proliferation and its deletion leads to chronic lymphocytic leukemia. Cancer Cell 17, 28–40 (2010).
Garzon, R. et al. MicroRNA-29 functions in AML. Blood 114, 5331–5341 (2009).
Sampson, V. B. et al. MicroRNA let-7a down-regulates MYC and reverts MYC-induced growth in Burkitt lymphoma cells. Cancer Res. 67, 9762–9770 (2007).
Lujambio, A. et al. A microRNA DNA methylation signature for human cancer metastasis. Proc. Natl Acad. Sci. USA 105, 13556–13561 (2008). Comprehensive analysis of epigentically regulated miRNAs in cancer metastasis.
Chan, J. A., Krichevsky, A. M. & Kosik, K. S. MicroRNA-21 is an antiapoptotic factor in human glioblastoma cells. Cancer Res. 65, 6029–6033 (2005).
Meng, F. et al. MicroRNA-21 regulates expression of the PTEN tumor suppressor gene in human hepatocellular cancer. Gastroenterology 133, 647–658 (2007).
Frankel, LB. et al. Programmed cell death 4 (PDCD4) is an important functional target of the microRNA miR-21 in breast cancer cells. J. Biol. Chem. 283, 1026–1033 (2008).
He, L. et al. A microRNA polycistron as a potential human oncogene. Nature 435, 828–833 (2005).
Xiao, C. et al. Lymphoproliferative disease and autoimmunity in mice with elevated miR-17-92 expression in lymphocytes. Nature Immunol. 9, 405–414 (2008). References 69 and 70 established the role of the miR-17-92 cluster in cancer and immunity.
Ventura, A. et al. Targeted deletion reveals essential and overlapping functions of the miR-17 through 92 family of miRNA clusters. Cell 132, 875–886 (2008). First report analysing the contribution of each miR-17-92 paralogues to the phenotype observed in mice with deletion of the whole miR-17-92 cluster.
Mu, P. et al. Genetic dissection of the miR-17-92 cluster of microRNAs in Myc-induced B-cell lymphomas. Genes Dev. 23, 2806–2811 (2009).
Olive, V. et al. miR-19 is a key oncogenic component of miR-17-92. Genes Dev. 23, 2839–2849 (2009). References 72 and 73 established the importance of miR-19 in the oncogenic role of the miR-17-92 cluster.
Costinean, S. et al. Pre-B cell proliferation and lymphoblastic leukemia/high-grade lymphoma in Eμ-miR155 transgenic mice. Proc. Natl Acad. Sci. USA 103, 7024–7029 (2006). First report indicating that the overexpression of a single miRNA can cause cancer.
O'Connell, R. M. et al. Sustained expression of micro-RNA-155 in hematopoietic stem cells causes a myeloproliferative disorder. J. Exp. Med. 205, 585–594 (2008).
O'Connell, R. M., Chaudhuri, A. A, Rao, D. S. & Baltimore, D. Inositol phosphatase SHIP1 is a primary target of miR-155. Proc. Natl Acad. Sci. USA 106, 7113–7118 (2009).
Costinean, S. et al. Src homology 2 domain-containing inositol-5-phosphatase and CCAAT enhancer-binding protein β are targeted by miR-155 in B cells of Eμ-MiR-155 transgenic mice. Blood 114, 1374–1382 (2009).
Liu, Q. et al. The inositol polyphosphate 5-phosphatase ship is a crucial negative regulator of B cell antigen receptor signaling. J. Exp. Med. 188, 1333–1342 (1998).
Ma, L, Teruya-Feldstein, J. & Weinberg, R. A. Tumour invasion and metastasis initiated by microRNA-10b in breast cancer. Nature 449, 682–688 (2007). First evidence that miRNAs are involved in tumour invasion and metastasis.
Ma, L. et al. Therapeutic silencing of miR-10b inhibits metastasis in a mouse mammary tumor model. Nature Biotechnol. 28, 341–347 (2010).
Tavazoie, S. F. et al. Endogenous human microRNAs that suppress breast cancer metastasis. Nature 451, 147–152 (2008).
Linsley, P. S. et al. Transcripts targeted by the microRNA-16 family cooperatively regulates cell cycle progression. Mol. Cell Biol. 27, 2240–2252 (2007).
Calin, G. A. et al. MiR-15a and miR-16-1 cluster functions in human leukemia. Proc. Natl Acad. Sci. USA 105, 5166–5171 (2008).
Li, Q. J. et al. miR-181a is an intrinsic modulator of T cell sensitivity and selection. Cell 129, 147–161 (2007). Elegant study showing the multitarget activity of a single mRNA in a pathway.
Croce, C. M. Oncogenes and cancer. N. Engl. J. Med. 358, 502–511 (2008).
Boutla, A., Delidakis, C. & Tabler, M. Developmental defects by antisense-mediated inactivation of micro-RNAs 2 and 13 in Drosophila and the identification of putative targets. Nucleic Acids Res. 31, 4973–4980 (2003).
Hutzvanger, G., Simard, M. J., Mello, C. C. & Zamore, P. D. Sequence specific inhibition of small RNA function. PLoS Biol. 2, E98 (2004).
Weiler, J., Hunziker, J. & Hall, J. Anti-miRNA oligonucleotides (AMOs); amunition to target miRNAs implicated in human disease? Gene Therapy 13, 496–502 (2006).
Krutzfeldt, J. et al. Silencing of microRNAs in vivo with 'antagomirs'. Nature 438, 685–689 (2005). First in vivo report using antisense oligonucleotides against miRNAs with success.
Vester, B. & Wengel, J. LNA (locked nucleic acid): high affinity targeting of complementary RNA and DNA. Biochemistry 43, 13233–13241 (2004).
Elmén, J. et al. Antagonism of microRNA-122 in mice by systemically administered LNA-antimiR leads to up-regulation of a large set of predicted target mRNAs in the liver. Nucleic Acids Res. 36, 1153–1162 (2008).
Jopling, C. L., Yi, M., Lancaster, A. M., Lemon, S. M. & Sarnow, P. Modulation of hepatitis C virus RNA abundance by a liver-specific microRNA. Science 309, 1577–1581 (2005).
Elmén, J. et al. LNA-mediated microRNA silencing in non-human primates. Nature 452, 896–899 (2008).
Lanford, R. E. et al. Therapeutic silencing of microRNA-122 in primates with chronic hepatitis c virus infection. Science 327, 198–201 (2010).
Ebert, M. S., Neilson, J. R. & Sharp, P. A. MicroRNA sponges: competitive inhibitors of small RNAs in mammalian cells. Nature Methods 4, 721–726 (2007).
Xiao, J. et al. Novel approaches for gene-specific interference via manipulating actions of micro-RNAs: examination on the pacemaker channel genes HCN2 and HCN4. J. Cell Physiol. 212, 285–292 (2007).
Loya, C. M., Lu, C. S., Van Vactor, D. & Fulga, T. A. Transgenic microRNA inhibition with spatiotemporal specificity in intact organisms. Nature Methods 6, 897–903 (2009).
Choi, W. Y., Giraldez, A. J. & Schier, A. F. Target protectors reveal dampening and balancing of Nodal agonist and antagonist by miR-430. Science 318, 271–274 (2007).
Gumireddy, K. et al. Small-molecule inhibitors of microrna miR-21 function. Angew. Chem. Int. Ed. Engl. 47, 7482–7484 (2008).
Michelfelder, S. & Trepel, M. Adeno-associated viral vectors and their redirection to cell-type specific receptors. Adv. Genet. 67, 29–60 (2009).
Aagaard, L. & Rossi, J. J. RNAi therapeutics: principles, prospects and challenges. Adv. Drug Deliv. Rev. 59, 75–86 (2007).
Kota, J. et al. Therapeutic microRNA delivery suppresses tumorigenesis in a murine liver cancer model. Cell 137, 1005–1017 (2009). First report using adenovirus-associated vectors to deliver miRNAS to the liver.
Garzon, R. et al. MicroRNA gene expression during retinoic acid-induced differentiation of human acute promyelocytic leukemia. Oncogene 26, 4148–4157 (2007).
Roccaro, A. et al. MicroRNA expression in the biology, prognosis, and therapy of Waldenström macroglobulinemia. Blood 113, 4391–4402 (2009).
Galm, O., Herman, J. G. & Baylin, S. B. The fundamental role of epigenetics in hematopoietic malignancies. Blood Rev. 20, 1–13 (2006).
Lamb, J. et al. The connectivity map: using gene-expression signatures to connect small molecules, genes, and disease. Science 313, 1929–1935 (2006).
Zhao, X. et al. Controlled delivery of antisense oligonucleotides: a brief review of current strategies. Exp. Opin. Drug Deliv. 6, 673–686 (2009).
Dias, N. & Stein, C. A. Antisense oligonucleotides: basic concepts and mechanisms. Mol. Cancer Ther. 1, 347–355 (2002).
Malik, R. & Roy, L. Design and development of antisense drugs. Exp. Opin. Drug Discov. 3, 1189–1207 (2008).
Chiarantini, L. et al. Comparison of a novel delivery systems for antisense peptide nucleic acids. J. Control. Release 109, 24–36 (2005).
Zuhorn, I. S., Engberts, J. B. & Hoeckstra, D. Gene delivery by cationic lipid vectors: overcoming cellular barriers. Eur. Biophys. J. 36; 349–362 (2007).
Chirila, T. V. et al. The use of synthethic polymers for delivery of therapeutic antisense oligodeoxynucleotides. Biomaterials 23, 321–342 (2002).
Rosi, N. L. et al. Oligonucleotide-modified gold nanoparticles for intracellular gene regulation. Science 312, 1027–1030 (2006).
Li, Z. et al. Biological functions of miR-29b contribute to positive regulation of osteoblast differentiation. J. Biol. Chem. 284, 15676–15684 (2009).
Xu, H., Cheung, I. Y., Guo, H. F. & Cheung, N. K. MicroRNA miR-29 modulates expression of immunoinhibitory molecule B7-H3: potential implications for immune based therapy of human solid tumors. Cancer Res. 69, 6275–6281 (2009).
Hornung, V. et al. Sequence-specific potent induction of IFN-α by short interfering RNA in plasmacytoid dendritic cells through TLR7. Nature Med. 11, 263–270 (2005).
Grimm, D. et al. Fatality in mice due to oversaturation of cellular microRNA/short hairpin RNA pathways. Nature 441, 537–541 (2006).
Garzon, R. et al. MicroRNA-29b induces global DNA hypomethylation and tumor suppressor gene reexpression in acute myeloid leukemia by targeting directly DNMT3A and 3B and indirectly DNMT1. Blood 113, 6411–6418 (2009). This work established miR-29 oligonucleotides as hypomethylating agents in leukaemia.
Fabbri, M. et al. MicroRNA-29 family reverts aberrant methylation in lung cancer by targeting DNA methyltransferases 3A and 3B. Proc. Natl Acad. Sci. USA 104, 15805–15810 (2007). First report that a miRNA can target the DNA methyltransferase machinery.
Thai, T. H. et al. Regulation of the germinal center response by microRNA-155. Science 316, 604–608 (2007).
Rodriguez, A. et al. Requirement of bic/microRNA-155 for normal immune function. Science 316, 608–611 (2007). References 120 and 121 established the role of miR-155 in the regulation of the immune system.
Kotani, Ai. et al. miR-128b is a potent glucocorticoid sensitizer in MLL-AF4 acute lymphocytic leukemia cells and exerts cooperative effects with miR-221. Blood 114, 4169–4178 (2009).
Pekarsky, Y. et al. Tcl1 expression in CLL is regulated by miR-29 and miR-181. Cancer Res. 66, 1590–11593 (2006).
Trang, P. et al. Regression of murine lung tumors by the let-7 microRNA. Oncogene 29, 1580–1587 (2010).
Lee, Y. S. & Dutta, A. The tumor suppressor microRNA let-7 represses the HMGA2 oncogene. Genes Dev. 21, 1025–1030 (2007).
Lodygin, D. et al. Inactivation of miR-34a by aberrant CpG methylation in multiple types of cancer. Cell Cycle 7, 2591–2600 (2008).
Toyota, M. et al. Epigenetic silencing of microRNA-34b/c and B-cell translocation gene 4 is associated with CpG island methylation in colorectal cancer. Cancer Res. 68, 4123–4132 (2008).
Sun, F. et al. Downregulation of CCND1 and CDK6 by miR-34a induces cell cycle arrest. FEBS Lett. 582, 1564–1568 (2008).
Yamakuchi, M., Ferlito, M. & Lowenstein, C. J. miR-34a repression of SIRT1 regulates apoptosis. Proc. Natl Acad. Sci. USA 105, 13421–13426 (2008).
Pigazzi, M., Manara, E., Baron, E. & Basso, G. miR-34b targets cyclic AMP-responsive element binding protein in acute myeloid leukemia. Cancer Res. 69, 2471–2478 (2009).
Cole, K. A. et al. A functional screen identifies miR-34a as a candidate neuroblastoma tumor suppressor gene. Mol. Cancer Res. 6, 735–742 (2008).
Li, Z. et al. Distinct microRNA expression profiles in acute myeloid leukemia with common translocations. Proc. Natl Acad. Sci. USA 105, 15535–15540 (2008).
O'Donnell, K. A. et al. c-Myc-regulated microRNAs modulate E2F1 expression. Nature 435, 839–843 (2005).
Velu, C. S., Baktula, A. M. & Grimes, H. L. Gfi1 regulates miR-21 and miR-196b to control myelopoiesis. Blood 113, 4720–4728 (2009).
Löffler, D. et al. Interleukin-6 dependent survival of multiple myeloma cells involves the Stat3-mediated induction of microRNA-21 through a highly conserved enhancer. Blood 110, 1330–1333 (2007).
Zhu, S., Si., M-L, Wu, H. & Mo, Y. Y. MicroRNA-21 targets the tumor suppressor gene tropomyosin 1 (TPM1). J. Biol. Chem. 282, 14328–14336 (2007).
Voorhoeve, P. M. et al. A genetic screen implicates miRNA-372 and miRNA-373 as oncogenes in testicular germ cell tumors. Cell 124, 1169–1181 (2007).
Author information
Authors and Affiliations
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Glossary
- Antisense
-
The term antisense is generally used for nucleic acid-based approaches that interferes, in a sequence-selective way, with the processing of RNA from its transcription via mRNA to protein or with the effects of other forms of functional RNA.
- Phosphorothioate
-
An oligonucleotide in which the oxygen atom normally linking two consecutive nucleotides has been replaced with sulphur, which resists degradation by cellular enzymes. The phosphorothioate backbone, although reducing affinity to the target RNA, confers considerable stability to nuclease degradation and is essential for in vivo delivery of antisense oligonucleotides to tissues, as the phosphorothioate promotes protein binding and delays plasma clearance.
Rights and permissions
About this article
Cite this article
Garzon, R., Marcucci, G. & Croce, C. Targeting microRNAs in cancer: rationale, strategies and challenges. Nat Rev Drug Discov 9, 775–789 (2010). https://doi.org/10.1038/nrd3179
Published:
Issue Date:
DOI: https://doi.org/10.1038/nrd3179
This article is cited by
-
Mesenchymal stem cell-derived exosomes enriched with miR-218 reduce the epithelial–mesenchymal transition and angiogenesis in triple-negative breast cancer cells
European Journal of Medical Research (2023)
-
Multi-sensitive functionalized niosomal nanocarriers for controllable gene delivery in vitro and in vivo
Cancer Nanotechnology (2023)
-
miR-92a-3p regulates cisplatin-induced cancer cell death
Cell Death & Disease (2023)
-
HSF-1/miR-145-5p transcriptional axis enhances hyperthermic intraperitoneal chemotherapy efficacy on peritoneal ovarian carcinosis
Cell Death & Disease (2023)
-
Small molecule regulators of microRNAs identified by high-throughput screen coupled with high-throughput sequencing
Nature Communications (2023)