Abstract
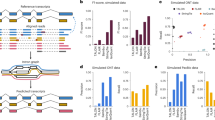
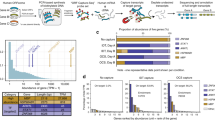
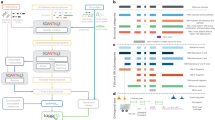
Hundreds of transcript isoforms with varying boundaries and alternative regulatory signals are transcribed from the genome, even in a genetically homogeneous population of cells. To study this transcriptional heterogeneity, we developed transcript isoform sequencing (TIF-seq), a method that allows the genome-wide profiling of full-length transcript isoforms defined by their exact 5′ and 3′ boundaries. TIF-seq entails the generation of full-length cDNA libraries, followed by their circularization and the sequencing of the junction fragments spanning the 5′ and 3′ transcript ends. By determining the respective co-occurrence of start and end sites of individual transcript molecules, TIF-seq can distinguish variations that conventional approaches for mapping single ends cannot, such as short abortive transcripts, bicistronic messages and overlapping transcripts that differ in lengths. The TIF-seq protocol we describe here can be applied to any eukaryotic organism (e.g., yeast, human), and it requires 6–10 d for generating TIF-seq libraries, 10 d for sequencing and 2–3 d for analysis.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout



Similar content being viewed by others
References
Djebali, S. et al. Landscape of transcription in human cells. Nature 489, 101–108 (2012).
Di Giammartino, D.C., Nishida, K. & Manley, J.L. Mechanisms and consequences of alternative polyadenylation. Mol. Cell 43, 853–866 (2011).
Gupta, I. et al. Alternative polyadenylation diversifies post-transcriptional regulation by selective RNA-protein interactions. Mol. Syst. Biol. 10, 719 (2014).
Kelemen, O. et al. Function of alternative splicing. Gene 514, 1–30 (2013).
Xu, Z. et al. Bidirectional promoters generate pervasive transcription in yeast. Nature 457, 1033–1037 (2009).
Nagalakshmi, U. et al. The transcriptional landscape of the yeast genome defined by RNA sequencing. Science 320, 1344–1349 (2008).
Wei, W., Pelechano, V., Jarvelin, A.I. & Steinmetz, L.M. Functional consequences of bidirectional promoters. Trends Genet. 27, 267–276 (2011).
Jacquier, A. The complex eukaryotic transcriptome: unexpected pervasive transcription and novel small RNAs. Nat. Rev. Genet. 10, 833–844 (2009).
Carninci, P. et al. Genome-wide analysis of mammalian promoter architecture and evolution. Nat. Genet. 38, 626–635 (2006).
Zhang, Z. & Dietrich, F.S. Mapping of transcription start sites in Saccharomyces cerevisiae using 5′ SAGE. Nucleic Acids Res. 33, 2838–2851 (2005).
Ozsolak, F. et al. Comprehensive polyadenylation site maps in yeast and human reveal pervasive alternative polyadenylation. Cell 143, 1018–1029 (2010).
Moqtaderi, Z., Geisberg, J.V., Jin, Y., Fan, X. & Struhl, K. Species-specific factors mediate extensive heterogeneity of mRNA 3′ ends in yeasts. Proc. Natl. Acad. Sci. USA 110, 11073–11078 (2013).
Wilkening, S. et al. An efficient method for genome-wide polyadenylation site mapping and RNA quantification. Nucleic Acids Res. 41, e65 (2012).
Pelechano, V., Wilkening, S., Jarvelin, A.I., Tekkedil, M.M. & Steinmetz, L.M. Genome-wide polyadenylation site mapping. Methods in Enzymology 513, 271–296 (2012).
Pelechano, V., Wei, W. & Steinmetz, L.M. Extensive transcriptional heterogeneity revealed by isoform profiling. Nature 497, 127–131 (2013).
Ng, P. et al. Multiplex sequencing of paired-end ditags (MS-PET): a strategy for the ultra-high-throughput analysis of transcriptomes and genomes. Nucleic Acids Res. 34, e84 (2006).
Ng, P. et al. Gene identification signature (GIS) analysis for transcriptome characterization and genome annotation. Nat. Methods 2, 105–111 (2005).
Fullwood, M.J. et al. An oestrogen receptor-α–bound human chromatin interactome. Nature 462, 58–64 (2009).
Ruan, X. & Ruan, Y. Genome wide full-length transcript analysis using 5′ and 3′ paired-end-tag next generation sequencing (RNA-PET). Methods Mol. Biol. 809, 535–562 (2012).
Carninci, P. et al. High-efficiency full-length cDNA cloning by biotinylated CAP trapper. Genomics 37, 327–336 (1996).
Zhu, Y.Y., Machleder, E.M., Chenchik, A., Li, R. & Siebert, P.D. Reverse transcriptase template switching: a SMART approach for full-length cDNA library construction. BioTechniques 30, 892–897 (2001).
Maruyama, K. & Sugano, S. Oligo-capping: a simple method to replace the cap structure of eukaryotic mRNAs with oligoribonucleotides. Gene 138, 171–174 (1994).
Scotto-Lavino, E., Du, G. & Frohman, M.A. Amplification of 5′ end cDNA with 'new RACE'. Nat. Protoc. 1, 3056–3061 (2006).
Carninci, P. Constructing the landscape of the mammalian transcriptome. J. Exp. Biol. 210, 1497–1506 (2007).
Kuai, L., Fang, F., Butler, J.S. & Sherman, F. Polyadenylation of rRNA in Saccharomyces cerevisiae. Proc. Natl. Acad. Sci. USA 101, 8581–8586 (2004).
Van Nieuwerburgh, F. et al. Quantitative bias in Illumina TruSeq and a novel post amplification barcoding strategy for multiplexed DNA and small RNA deep sequencing. PLoS ONE 6, e26969 (2011).
Chen, Y. et al. Systematic evaluation of factors influencing ChIP-seq fidelity. Nat. Methods 9, 609–614 (2012).
Miura, F. et al. A large-scale full-length cDNA analysis to explore the budding yeast transcriptome. Proc. Natl. Acad. Sci. USA 103, 17846–17851 (2006).
Sharon, D., Tilgner, H., Grubert, F. & Snyder, M. A single-molecule long-read survey of the human transcriptome. Nat. Biotechnol. 31, 1009–1014 (2013).
McManus, C.J., Duff, M.O., Eipper-Mains, J. & Graveley, B.R. Global analysis of trans-splicing in Drosophila. Proc. Natl. Acad. Sci. USA 107, 12975–12979 (2010).
Anders, S. & Huber, W. Differential expression analysis for sequence count data. Genome Biol. 11, R106 (2010).
Wilkening, S. et al. Genotyping 1000 yeast strains by next-generation sequencing. BMC Genomics 14, 90 (2013).
Thorvaldsdottir, H., Robinson, J.T. & Mesirov, J.P. Integrative Genomics Viewer (IGV): high-performance genomics data visualization and exploration. Brief. Bioinform. 14, 178–192 (2013).
Acknowledgements
We thank R. Aiyar for help in editing and refining the manuscript; A.I. Järvelin, J. Zaugg and S. Clauder-Münster for help in optimizing the TIF-seq protocol; and the members of the Steinmetz laboratory for helpful discussions and critical comments. This study was technically supported by the EMBL Genomics Core Facility. This study was financially supported by the US National Institutes of Health and the EMBL (to L.M.S.).
Author information
Authors and Affiliations
Contributions
W.W., V.P. and L.M.S. conceived the project; V.P. developed the TIF-seq method; P.J. contributed to further method optimization; W.W. and V.P. performed data analysis; and L.M.S. supervised the study. All authors wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Rights and permissions
About this article
Cite this article
Pelechano, V., Wei, W., Jakob, P. et al. Genome-wide identification of transcript start and end sites by transcript isoform sequencing. Nat Protoc 9, 1740–1759 (2014). https://doi.org/10.1038/nprot.2014.121
Published:
Issue Date:
DOI: https://doi.org/10.1038/nprot.2014.121
This article is cited by
-
High-resolution analysis of cell-state transitions in yeast suggests widespread transcriptional tuning by alternative starts
Genome Biology (2021)
-
Transcript isoform sequencing reveals widespread promoter-proximal transcriptional termination in Arabidopsis
Nature Communications (2020)
-
Alternative cleavage and polyadenylation in health and disease
Nature Reviews Genetics (2019)
-
Sensitive high-throughput single-cell RNA-seq reveals within-clonal transcript correlations in yeast populations
Nature Microbiology (2019)
-
Genome-wide quantification of 5′-phosphorylated mRNA degradation intermediates for analysis of ribosome dynamics
Nature Protocols (2016)
Comments
By submitting a comment you agree to abide by our Terms and Community Guidelines. If you find something abusive or that does not comply with our terms or guidelines please flag it as inappropriate.