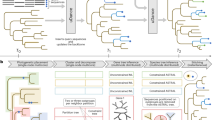
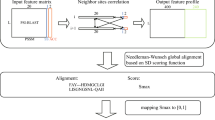
Abstract
Software for visualizing sequence alignments and trees are essential tools for life scientists. In this review, we describe the major features and capabilities of a selection of stand-alone and web-based applications useful when investigating the function and evolution of a gene family. These range from simple viewers, to systems that provide sophisticated editing and analysis functions. We conclude with a discussion of the challenges that these tools now face due to the flood of next generation sequence data and the increasingly complex network of bioinformatics information sources.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Altschul, S.F. et al. Gapped BLAST and PSI-BLAST: a new generation of protein database search programs. Nucleic Acids Res. 25, 3389–3402 (1997).
Johnson, M. et al. NCBI BLAST: a better web interface. Nucleic Acids Res. 36, W5–W9 (2008).
Lu, G. & Moriyama, E.N. Vector NTI, a balanced all-in-one sequence analysis suite. Brief. Bioinform. 5, 378–388 (2004).
Thompson, J.D., Gibson, T.J. & Higgins, D.G. Multiple sequence alignment using ClustalW and ClustalX. Curr. Protoc. Bioinformatics 2, 2.3.1–2.3.22 (2002).
Notredame, C., Higgins, D.G. & Heringa, J. T-Coffee: A novel method for fast and accurate multiple sequence alignment. J. Mol. Biol. 302, 205–217 (2000).
Edgar, R.C. & Batzoglou, S. Multiple sequence alignment. Curr. Opin. Struct. Biol. 16, 368–373 (2006). A comprehensive review of the approaches available for the alignment of many sequences.
Raghava, G.P., Searle, S.M., Audley, P.C., Barber, J.D. & Barton, G.J. OXBench: a benchmark for evaluation of protein multiple sequence alignment accuracy. BMC Bioinformatics 4, 47 (2003).
Gouet, P., Robert, X. & Courcelle, E. ESPript/ENDscript: extracting and rendering sequence and 3D information from atomic structures of proteins. Nucleic Acids Res. 31, 3320–3323 (2003).
Barton, G.J. ALSCRIPT: a tool to format multiple sequence alignments. Protein Eng. 6, 37–40 (1993).
Goodstadt, L. & Ponting, C.P. CHROMA: consensus-based colouring of multiple alignments for publication. Bioinformatics 17, 845–846 (2001).
Barrio, A.M., Lagercrantz, E., Sperber, G.O., Blomberg, J. & Bongcam-Rudloff, E. Annotation and visualization of endogenous retroviral sequences using the Distributed Annotation System (DAS) and eBioX. BMC Bioinformatics 10 (suppl. 6), S18 (2009).
Finn, R.D. et al. The Pfam protein families database. Nucleic Acids Res. 36, D281–D288 (2008).
Lin, K., May, A.C. & Taylor, W.R. Amino acid encoding schemes from protein structure alignments: multi-dimensional vectors to describe residue types. J. Theor. Biol. 216, 361–365 (2002). The empirical analysis underlying the 'Taylor' amino acid color scheme; this builds on Taylor's earlier work (1986) concerning approaches for the classification of amino acids.
Valdar, W.S. Scoring residue conservation. Proteins 48, 227–241 (2002).
Chakrabarti, S. & Lanczycki, C.J. Analysis and prediction of functionally important sites in proteins. Protein Sci. 16, 4–13 (2007).
Schneider, T.D. & Stephens, R.M. Sequence logos: a new way to display consensus sequences. Nucleic Acids Res. 18, 6097–6100 (1990).
Schneider, T.D. Twenty years of Delila and molecular information theory: the Altenberg-Austin Workshop in Theoretical Biology biological information, beyond metaphor: causality, explanation, and unification Altenberg, Austria, 11–14 July 2002. Biol. Theory 1, 250–260 (2006).
Caffrey, D.R. et al. PFAAT version 2.0: a tool for editing, annotating, and analyzing multiple sequence alignments. BMC Bioinformatics 8, 381 (2007).
Rastogi, P.A. MacVector. Integrated sequence analysis for the Macintosh. Methods Mol. Biol. 132, 47–69 (2000).
Gille, C. & Robinson, P.N. HotSwap for bioinformatics: a STRAP tutorial. BMC Bioinformatics 7, 64 (2006).
Bailey, T.L. et al. MEME SUITE: tools for motif discovery and searching. Nucleic Acids Res. 37, W202–W208 (2009).
Landan, G. & Graur, D. Characterization of pairwise and multiple sequence alignment errors. Gene 441, 141–147 (2009). To our knowledge, this is the first detailed analysis of the errors that may be introduced by tree based sequence alignment algorithms.
Galtier, N., Gouy, M. & Gautier, C. SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny. Comput. Appl. Biosci. 12, 543–548 (1996).
Lord, P.W., Selley, J.N. & Attwood, T.K. CINEMA-MX: a modular multiple alignment editor. Bioinformatics 18, 1402–1403 (2002).
Waterhouse, A.M., Procter, J.B., Martin, D.M., Clamp, M. & Barton, G.J. Jalview Version 2–a multiple sequence alignment editor and analysis workbench. Bioinformatics 25, 1189–1191 (2009).
Margulies, E.H. & Birney, E. Approaches to comparative sequence analysis: towards a functional view of vertebrate genomes. Nat. Rev. Genet. 9, 303–313 (2008).
Hulo, N. et al. The 20 years of PROSITE. Nucleic Acids Res. 36, D245–D249 (2008).
Wingender, E. The TRANSFAC project as an example of framework technology that supports the analysis of genomic regulation. Brief. Bioinform. 9, 326–332 (2008).
Matys, V. et al. TRANSFAC and its module TRANSCompel: transcriptional gene regulation in eukaryotes. Nucleic Acids Res. 34, D108–D110 (2006).
Zvelebil, M.J., Barton, G.J., Taylor, W.R. & Sternberg, M.J. Prediction of protein secondary structure and active sites using the alignment of homologous sequences. J. Mol. Biol. 195, 957–961 (1987).
Chakrabarti, S. & Panchenko, A.R. Ensemble approach to predict specificity determinants: benchmarking and validation. BMC Bioinformatics 10, 207 (2009).
Horner, D.S., Pirovano, W. & Pesole, G. Correlated substitution analysis and the prediction of amino acid structural contacts. Brief. Bioinform. 9, 46–56 (2008).
Casari, G., Sander, C. & Valencia, A. A method to predict functional residues in proteins. Nat. Struct. Biol. 2, 171–178 (1995).
Schwarz, R. et al. Detecting species-site dependencies in large multiple sequence alignments. Nucleic Acids Res. 37, 5959–5968 (2009).
Joachimiak, M.P. & Cohen, F.E. JEvTrace: refinement and variations of the evolutionary trace in JAVA. Genome Biol. 3, RESEARCH0077 (2002).
Goldenberg, O., Erez, E., Nimrod, G. & Ben-Tal, N. The ConSurf-DB: pre-calculated evolutionary conservation profiles of protein structures. Nucleic Acids Res. 37, D323–D327 (2009).
Li, W. & Godzik, A. VISSA: a program to visualize structural features from structure sequence alignment. Bioinformatics 22, 887–888 (2006).
Brown, J.W. et al. The RNA structure alignment ontology. RNA 15, 1623–1631 (2009).
Chen, K., Durand, D. & Farach-Colton, M. NOTUNG: a program for dating gene duplications and optimizing gene family trees. J. Comput. Biol. 7, 429–447 (2000).
Vernot, B., Stolzer, M., Goldman, A. & Durand, D. Reconciliation with non-binary species trees. J. Comput. Biol. 15, 981–1006 (2008).
Bingham, J. & Sudarsanam, S. Visualizing large hierarchical clusters in hyperbolic space. Bioinformatics 16, 660–661 (2000).
Hughes, T., Hyun, Y. & Liberles, D.A. Visualising very large phylogenetic trees in three dimensional hyperbolic space. BMC Bioinformatics 5, 48 (2004).
Livingstone, C.D. & Barton, G.J. Protein sequence alignments: a strategy for the hierarchical analysis of residue conservation. Comput. Appl. Biosci. 9, 745–756 (1993).
Sankararaman, S. & Sjolander, K. INTREPID–INformation-theoretic TREe traversal for Protein functional site IDentification. Bioinformatics 24, 2445–2452 (2008).
Engelen, S., Trojan, L.A., Sacquin-Mora, S., Lavery, R. & Carbone, A. Joint evolutionary trees: a large-scale method to predict protein interfaces based on sequence sampling. PLoS Comput. Biol. 5, e1000267 (2009).
Chevenet, F., Brun, C., Banuls, A.L., Jacq, B. & Christen, R. TreeDyn: towards dynamic graphics and annotations for analyses of trees. BMC Bioinformatics 7, 439 (2006).
Santamaría, R. & Theron, R. Treevolution: visual analysis of phylogenetic trees. Bioinformatics 25, 1970–1971 (2009).
Letunic, I. & Bork, P. Interactive Tree Of Life (iTOL): an online tool for phylogenetic tree display and annotation. Bioinformatics 23, 127–128 (2007).
Müller, J. & Müller, K. TreeGraph: automated drawing of complex tree figures using an extensible tree description format. Mol. Ecol. Notes 4, 786–788 (2004).
Pettifer, S. et al. Visualising biological data: a semantic approach to tool and database integration. BMC Bioinformatics 10 (supp. 6), S19 (2009).
Raphael, B., Zhi, D., Tang, H. & Pevzner, P. A novel method for multiple alignment of sequences with repeated and shuffled elements. Genome Res. 14, 2336–2346 (2004). Introduces the partially ordered alignment algorithm and demonstrates how this graph based alignment visualization provides a more compact view of complex alignments.
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009). Describes the CIRCOS approach for visualization of comparative genomic data, which can provide a more compact view of large multiple sequence alignments.
UniProt Consortium. The Universal Protein Resource (UniProt) 2009. Nucleic Acids Res. 37, D169–D174 (2009).
Berman, H.M. et al. The nucleic acid database. A comprehensive relational database of three-dimensional structures of nucleic acids. Biophys. J. 63, 751–759 (1992).
Taylor, W.R. The classification of amino acid conservation. J. Theor. Biol. 119, 205–218 (1986).
Mirny, L.A. & Shakhnovich, E.I. Universally conserved positions in protein folds: reading evolutionary signals about stability, folding kinetics and function. J. Mol. Biol. 291, 177–196 (1999).
Schuster-Böckler, B. & Bateman, A. Visualizing profile-profile alignment: pairwise HMM logos. Bioinformatics 21, 2912–2913 (2005).
Eddy, S.R. Profile hidden Markov models. Bioinformatics 14, 755–763 (1998).
Seibel, P.N., Muller, T., Dandekar, T. & Wolf, M. Synchronous visual analysis and editing of RNA sequence and secondary structure alignments using 4SALE. BMC Res. Notes 1, 91 (2008).
Wilm, A., Linnenbrink, K. & Steger, G. ConStruct: improved construction of RNA consensus structures. BMC Bioinformatics 9, 219 (2008).
Jossinet, F. & Westhof, E. Sequence to Structure (S2S): display, manipulate and interconnect RNA data from sequence to structure. Bioinformatics 21, 3320–3321 (2005).
Andersen, E.S. et al. Semiautomated improvement of RNA alignments. RNA 13, 1850–1859 (2007).
Gille, C. Structural interpretation of mutations and SNPs using STRAP-NT. Protein Sci. 15, 208–210 (2006).
Mizuguchi, K., Deane, C.M., Blundell, T.L., Johnson, M.S. & Overington, J.P. JOY: protein sequence-structure representation and analysis. Bioinformatics 14, 617–623 (1998).
Zmasek, C.M. & Eddy, S.R. ATV: display and manipulation of annotated phylogenetic trees. Bioinformatics 17, 383–384 (2001).
Archer, J. & Robertson, D.L. CTree: comparison of clusters between phylogenetic trees made easy. Bioinformatics 23, 2952–2953 (2007).
Huson, D.H. et al. Dendroscope: an interactive viewer for large phylogenetic trees. BMC Bioinformatics 8, 460 (2007).
Perrière, G. & Gouy, M. WWW-query: an on-line retrieval system for biological sequence banks. Biochimie 78, 364–369 (1996).
Hillis, D.M., Heath, T.A. & St. John, K. Analysis and visualization of tree space. Syst. Biol. 54, 471–482 (2005). A demonstration of different kinds of tree visualization, and an examination of how spatial techniques such as multidimensional scaling can be used to visualize and compare ensembles of trees.
Page, R.D. TreeView: an application to display phylogenetic trees on personal computers. Comput. Appl. Biosci. 12, 357–358 (1996).
Munzner, T., Guimbretiere, F., Tasiran, S., Zhang, L. & Zhou, Y. TreeJuxtaposer: scalable tree comparison using focus+context with guaranteed visibility. ACM Trans. Graph. 22, 453–462 (2003).
Kumar, S., Nei, M., Dudley, J. & Tamura, K. MEGA: a biologist-centric software for evolutionary analysis of DNA and protein sequences. Brief. Bioinform. 9, 299–306 (2008).
Huson, D.H. & Bryant, D. Application of phylogenetic networks in evolutionary studies. Mol. Biol. Evol. 23, 254–267 (2006). Describes the phylogenetic network visualization approach implemented in SplitsTree4, where evolutionary distance and bootstrap support are represented in one network structure, rather than an annotated tree.
Milne, I. et al. TOPALi v2: a rich graphical interface for evolutionary analyses of multiple alignments on HPC clusters and multi-core desktops. Bioinformatics 25, 126–127 (2009).
Jordan, G.E. & Piel, W.H. PhyloWidget: web-based visualizations for the tree of life. Bioinformatics 24, 1641–1642 (2008).
Prlić, A. et al. Integrating sequence and structural biology with DAS. BMC Bioinformatics 8, 333 (2007).
Berman, H.M. et al. The Protein Data Bank. Nucleic Acids Res. 28, 235–242 (2000).
Thompson, J.D. et al. MACSIMS: multiple alignment of complete sequences information management system. BMC Bioinformatics 7, 318 (2006).
Barrell, D. et al. The GOA database in 2009–an integrated Gene Ontology Annotation resource. Nucleic Acids Res. 37, D396–D403 (2009).
The Gene Ontology's Reference Genome Project. A unified framework for functional annotation across species. PLoS Comput. Biol. 5, e1000431 (2009).
Reeves, G.A. et al. The Protein Feature Ontology: a tool for the unification of protein feature annotations. Bioinformatics 24, 2767–2772 (2008).
Eilbeck, K. et al. The Sequence Ontology: a tool for the unification of genome annotations. Genome Biol. 6, R44 (2005).
Sayers, E.W. et al. Database resources of the National Center for Biotechnology Information. Nucleic Acids Res. 37, D5–D15 (2009).
Holder, M. & Lewis, P.O. Phylogeny estimation: traditional and Bayesian approaches. Nat. Rev. Genet. 4, 275–284 (2003).
Swofford, D.L., Olsen, G.J., Waddell, P.J. & Hillis, D.M. Phylogenetic inference. in Molecular Systematics (eds. Hillis, D.M., Moritz, C. & Mable, B.K.) 407–514 (Sinauer, Sunderland, Massachusetts, USA, 1996).
Felsenstein, J. Inferring Phylogenies (Sinauer, Sunderland, Massachusetts, USA, 2004).
Felsenstein, J. Evolutionary trees from DNA sequences: a maximum likelihood approach. J. Mol. Evol. 17, 368–376 (1981).
Huelsenbeck, J.P., Ronquist, F., Nielsen, R. & Bollback, J.P. Bayesian inference of phylogeny and its impact on evolutionary biology. Science 294, 2310–2314 (2001).
Acknowledgements
J.B.P. acknowledges the support of the ENFIN European Network of Excellence (contract LSHG-CT-2005-518254) awarded to G.J.B. Several tools were made available as prereleases to the authors for evaluation purposes, and we thank the individuals and companies who obliged our requests.
Author information
Authors and Affiliations
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 (PDF 2274 kb)
Rights and permissions
About this article
Cite this article
Procter, J., Thompson, J., Letunic, I. et al. Visualization of multiple alignments, phylogenies and gene family evolution. Nat Methods 7 (Suppl 3), S16–S25 (2010). https://doi.org/10.1038/nmeth.1434
Published:
Issue Date:
DOI: https://doi.org/10.1038/nmeth.1434
This article is cited by
-
Using sound to understand protein sequence data: new sonification algorithms for protein sequences and multiple sequence alignments
BMC Bioinformatics (2021)
-
Replacing the eleven native tryptophans by directed evolution produces an active P-glycoprotein with site-specific, non-conservative substitutions
Scientific Reports (2020)
-
Integrated visual analysis of protein structures, sequences, and feature data
BMC Bioinformatics (2015)
-
Sequence analysis reveals a conserved extension in the capping enzyme of the alphavirus supergroup, and a homologous domain in nodaviruses
Biology Direct (2015)
-
Mu-8: visualizing differences between proteins and their families
BMC Proceedings (2014)