Abstract
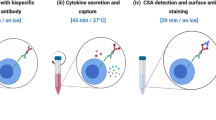
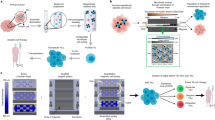
Cellular immunity has an inherent high level of functional heterogeneity. Capturing the full spectrum of these functions requires analysis of large numbers of effector molecules from single cells. We report a microfluidic platform designed for highly multiplexed (more than ten proteins), reliable, sample-efficient (∼1 × 104 cells) and quantitative measurements of secreted proteins from single cells. We validated the platform by assessment of multiple inflammatory cytokines from lipopolysaccharide (LPS)-stimulated human macrophages and comparison to standard immunotechnologies. We applied the platform toward the ex vivo quantification of T cell polyfunctional diversity via the simultaneous measurement of a dozen effector molecules secreted from tumor antigen–specific cytotoxic T lymphocytes (CTLs) that were actively responding to tumor and compared against a cohort of healthy donor controls. We observed profound, yet focused, functional heterogeneity in active tumor antigen–specific CTLs, with the major functional phenotypes quantitatively identified. The platform represents a new and informative tool for immune monitoring and clinical assessment.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Gordon, S. & Taylor, P.R. Monocyte and macrophage heterogeneity. Nat. Rev. Immunol. 5, 953–964 (2005).
O'Shea, J.J., Hunter, C.A. & Germain, R.N. T cell heterogeneity: firmly fixed, predominantly plastic or merely malleable? Nat. Immunol. 9, 450–453 (2008).
Kaech, S.M. & Wherry, E.J. Heterogeneity and cell-fate decisions in effector and memory CD8+ T cell differentiation during viral infection. Immunity 27, 393–405 (2007).
Darrah, P.A. et al. Multifunctional TH1 cells define a correlate of vaccine-mediated protection against Leishmania major. Nat. Med. 13, 843–850 (2007).
Lee, P.P. et al. Characterization of circulating T cells specific for tumor-associated antigens in melanoma patients. Nat. Med. 5, 677–685 (1999).
Seder, R.A., Darrah, P.A. & Roederer, M. T-cell quality in memory and protection: implications for vaccine design. Nat. Rev. Immunol. 8, 247–258 (2008).
Re, F. & Strominger, J.L. Heterogeneity of TLR-induced responses in dendritic cells: from innate to adaptive immunity. Immunobiology 209, 191–198 (2004).
Chattopadhyay, P.K., Yu, J. & Roederer, M. A live-cell assay to detect antigen-specific CD4+ T cells with diverse cytokine profiles. Nat. Med. 11, 1113–1117 (2005).
Zak, D.E. & Aderem, A. Systems biology of innate immunity. Immunol. Rev. 227, 264–282 (2009).
Precopio, M.L. et al. Immunization with vaccinia virus induces polyfunctional and phenotypically distinctive CD8+ T cell responses. J. Exp. Med. 204, 1405–1416 (2007).
Mantovani, A., Allavena, P., Sica, A. & Balkwill, F. Cancer-related inflammation. Nature 454, 436–444 (2008).
Coussens, L.M. & Werb, Z. Inflammation and cancer. Nature 420, 860–867 (2002).
Medzhitov, R. Origin and physiological roles of inflammation. Nature 454, 428–435 (2008).
Hagemann, T., Balkwill, F. & Lawrence, T. Inflammation and cancer: a double-edged sword. Cancer Cell 12, 300–301 (2007).
Appay, V., van Lier, R.A.W., Sallusto, F. & Roederer, M. Phenotype and function of human T lymphocyte subsets: consensus and issues. Cytometry A 73, 975–983 (2008).
Song, K. et al. Characterization of subsets of CD4+ memory T cells reveals early branched pathways of T cell differentiation in humans. Proc. Natl. Acad. Sci. USA 102, 7916–7921 (2005).
Kotecha, N. et al. Single-cell profiling identifies aberrant STAT5 activation in myeloid malignancies with specific clinical and biologic correlates. Cancer Cell 14, 335–343 (2008).
Bailey, R.C., Kwong, G.A., Radu, C.G., Witte, O.N. & Heath, J.R. DNA-encoded antibody libraries: A unified platform for multiplexed cell sorting and detection of genes and proteins. J. Am. Chem. Soc. 129, 1959–1967 (2007).
Fan, R. et al. Integrated barcode chips for rapid, multiplexed analysis of proteins in microliter quantities of blood. Nat. Biotechnol. 26, 1373–1378 (2008).
Shin, Y.S. et al. Chemistries for patterning robust DNA microbarcodes enable multiplex assays of cytoplasm proteins from single cancer cells. ChemPhysChem 11, 3063–3069 (2010).
Wang, J. et al. A self-powered, one-step chip for rapid, quantitative and multiplexed detection of proteins from pinpricks of whole blood. Lab Chip 10, 3157–3162 (2010).
Aderem, A. & Ulevitch, R.J. Toll-like receptors in the induction of the innate immune response. Nature 406, 782–787 (2000).
Fan, J. & Malik, A.B. Toll-like receptor 4 (TLR4) signaling augments chemokine-induced neutrophil migration by modulating cell surface expression of chemokine receptors. Nat. Med. 9, 315–321 (2003).
Shin, Y.S. et al. Protein signaling networks from single cell fluctuations and information theory profiling. Biophys. J. (in the press).
Attig, S. et al. Simultaneous Infiltration of polyfunctional effector and suppressor T cells into renal cell carcinomas. Cancer Res. 69, 8412–8419 (2009).
De Rosa, S.C. et al. Vaccination in humans generates broad T cell cytokine responses. J. Immunol. 173, 5372–5380 (2004).
Makedonas, G. & Betts, M.R. Polyfunctional analysis of human t cell responses: importance in vaccine immunogenicity and natural infection. Springer Semin. Immunopathol. 28, 209–219 (2006).
Morgan, R.A. et al. Cancer regression in patients after transfer of genetically engineered lymphocytes. Science 314, 126–129 (2006).
Kwong, G.A. et al. Modular nucleic acid assembled p/MHC microarrays for multiplexed sorting of antigen-specific T cells. J. Am. Chem. Soc. 131, 9695–9703 (2009).
Bachmann, M.F., Wolint, P., Schwarz, K., Jager, P. & Oxenius, A. Functional properties and lineage relationship of CD8+ T cell subsets identified by expression of IL-7 receptorα and CD62L. J. Immunol. 175, 4686–4696 (2005).
van Lier, R.A.W., ten Berge, I.J.M. & Gamadia, L.E. Human CD8+ T-cell differentiation in response to viruses. Nat. Rev. Immunol. 3, 931–939 (2003).
Pantaleo, G. & Koup, R.A. Correlates of immune protection in HIV-1 infection: what we know, what we don't know, what we should know. Nat. Med. 10, 806–810 (2004).
Ribas, A. et al. Dendritic cell vaccination combined with CTLA4 blockade in patients with metastatic melanoma. Clin. Cancer Res. 15, 6267–6276 (2009).
Thorsen, T., Maerkl, S.J. & Quake, S.R. Microfluidic large-scale integration. Science 298, 580–584 (2002).
Comin-Anduix, B. et al. Detailed analysis of immunologic effects of the cytotoxic T lymphocyte-associated antigen 4–blocking monoclonal antibody tremelimumab in peripheral blood of patients with melanoma. J. Transl. Med. 6, 22 (2008).
Comin-Anduix, B. et al. Definition of an immunologic response using the major histocompatibility complex tetramer and enzyme-linked immunospot assays. Clin. Cancer Res. 12, 107–116 (2006).
Eisen, M.B., Spellman, P.T., Brown, P.O. & Botstein, D. Cluster analysis and display of genome-wide expression patterns. Proc. Natl. Acad. Sci. USA 95, 14863–14868 (1998).
Acknowledgements
We thank B. Marzolf at the Institute for Systems Biology for printing DNA-spotted arrays and the UCLA nanolab for photomask fabrication. We thank L. Yang, S. Wang, R. Diamond and H. Wu for valuable discussion. C.M. acknowledges the support of the Benjamin M. Rosen Fellowship. R.F. is supported by the US National Institutes of Health K99 Pathway to Independence Award (No. 1 K99 CA136759-01). This work was funded by the US National Cancer Institute Grant No. 5U54 CA119347 (J.R.H.), by the Ivy Foundation and the Jean Perkins Foundation (J.R.H.), by the California Institute for Regenerative Medicine New Faculty Award RN2-00902-1 (A.R.), by the Caltech/UCLA Joint Center for Translational Medicine (A.R. and J.R.H.) and the Melanoma Research Alliance (A.R. and J.R.H.). The UCLA Flow Cytometry Core Facility is supported by the US National Institutes of Health awards CA-16042 and AI-28697.
Author information
Authors and Affiliations
Contributions
C.M. conducted T cell experiments and analyzed data. R.F. conducted macrophage experiments. C.M. and R.F. performed validation experiments and designed the chip. H.A. wrote Excel macros. Q.S., C.-C.L. and G.A.K. helped with experiments. B.C.-A., T.C. and R.C.K. collected T cell samples and conducted flow cytometry phenotyping experiments. C.M., R.F. and J.R.H. conceived of the experiments. C.M., R.F., C.G.R., A.R. and J.R.H. wrote the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–9, Supplementary Table 1–3 and Supplementary Methods (PDF 1568 kb)
Rights and permissions
About this article
Cite this article
Ma, C., Fan, R., Ahmad, H. et al. A clinical microchip for evaluation of single immune cells reveals high functional heterogeneity in phenotypically similar T cells. Nat Med 17, 738–743 (2011). https://doi.org/10.1038/nm.2375
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nm.2375
This article is cited by
-
The progressive trend of modeling and drug screening systems of breast cancer bone metastasis
Journal of Biological Engineering (2024)
-
Identification of druggable regulators of cell secretion via a kinome-wide screen and high-throughput immunomagnetic cell sorting
Nature Biomedical Engineering (2023)
-
High-throughput spatiotemporal monitoring of single-cell secretions via plasmonic microwell arrays
Nature Biomedical Engineering (2023)
-
Single-cell sorting based on secreted products for functionally defined cell therapies
Microsystems & Nanoengineering (2022)