Abstract
The major-histocompatibility-complex-(MHC)-class-I-related molecule MR1 can present activating and non-activating vitamin-B-based ligands to mucosal-associated invariant T cells (MAIT cells). Whether MR1 binds other ligands is unknown. Here we identified a range of small organic molecules, drugs, drug metabolites and drug-like molecules, including salicylates and diclofenac, as MR1-binding ligands. Some of these ligands inhibited MAIT cells ex vivo and in vivo, while others, including diclofenac metabolites, were agonists. Crystal structures of a T cell antigen receptor (TCR) from a MAIT cell in complex with MR1 bound to the non-stimulatory and stimulatory compounds showed distinct ligand orientations and contacts within MR1, which highlighted the versatility of the MR1 binding pocket. The findings demonstrated that MR1 was able to capture chemically diverse structures, spanning mono- and bicyclic compounds, that either inhibited or activated MAIT cells. This indicated that drugs and drug-like molecules can modulate MAIT cell function in mammals.
This is a preview of subscription content, access via your institution
Access options
Access Nature and 54 other Nature Portfolio journals
Get Nature+, our best-value online-access subscription
$29.99 / 30 days
cancel any time
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Jones, E.Y., Fugger, L., Strominger, J.L. & Siebold, C. MHC class II proteins and disease: a structural perspective. Nat. Rev. Immunol. 6, 271–282 (2006).
Koning, F., Thomas, R., Rossjohn, J. & Toes, R.E. Coeliac disease and rheumatoid arthritis: similar mechanisms, different antigens. Nat. Rev. Rheumatol. 11, 450–461 (2015).
Illing, P.T. et al. Immune self-reactivity triggered by drug-modified HLA-peptide repertoire. Nature 486, 554–558 (2012).
Godfrey, D.I., Uldrich, A.P., McCluskey, J., Rossjohn, J. & Moody, D.B. The burgeoning family of unconventional T cells. Nat. Immunol. 16, 1114–1123 (2015).
Van Rhijn, I., Godfrey, D.I., Rossjohn, J. & Moody, D.B. Lipid and small-molecule display by CD1 and MR1. Nat. Rev. Immunol. 15, 643–654 (2015).
Eckle, S.B. et al. Recognition of vitamin B precursors and byproducts by mucosal associated invariant T cells. J. Biol. Chem. 290, 30204–30211 (2015).
Treiner, E. et al. Selection of evolutionarily conserved mucosal-associated invariant T cells by MR1. Nature 422, 164–169 (2003).
Tilloy, F. et al. An invariant T cell receptor α chain defines a novel TAP-independent major histocompatibility complex class Ib-restricted α/β T cell subpopulation in mammals. J. Exp. Med. 189, 1907–1921 (1999).
Salio, M., Silk, J.D., Jones, E.Y. & Cerundolo, V. Biology of CD1- and MR1-restricted T cells. Annu. Rev. Immunol. 32, 323–366 (2014).
Gold, M.C. et al. Human mucosal associated invariant T cells detect bacterially infected cells. PLoS Biol. 8, e1000407 (2010).
Le Bourhis, L. et al. Antimicrobial activity of mucosal-associated invariant T cells. Nat. Immunol. 11, 701–708 (2010).
Dusseaux, M. et al. Human MAIT cells are xenobiotic-resistant, tissue-targeted, CD161hi IL-17-secreting T cells. Blood 117, 1250–1259 (2011).
Reantragoon, R. et al. Antigen-loaded MR1 tetramers define T cell receptor heterogeneity in mucosal-associated invariant T cells. J. Exp. Med. 210, 2305–2320 (2013).
Lepore, M. et al. Parallel T-cell cloning and deep sequencing of human MAIT cells reveal stable oligoclonal TCRβ repertoire. Nat. Commun. 5, 3866 (2014).
Corbett, A.J. et al. T-cell activation by transitory neo-antigens derived from distinct microbial pathways. Nature 509, 361–365 (2014).
Kjer-Nielsen, L. et al. MR1 presents microbial vitamin B metabolites to MAIT cells. Nature 491, 717–723 (2012).
Patel, O. et al. Recognition of vitamin B metabolites by mucosal-associated invariant T cells. Nat. Commun. 4, 2142 (2013).
Ussher, J.E., Klenerman, P. & Willberg, C.B. Mucosal-associated invariant T-cells: new players in anti-bacterial immunity. Front. Immunol. 5, 450 (2014).
Kurioka, A. et al. MAIT cells are licensed through granzyme exchange to kill bacterially sensitized targets. Mucosal Immunol. 8, 429–440 (2015).
Hashkes, P.J. et al. Methotrexate: new uses for an old drug. J. Pediatr. 164, 231–236 (2014).
Chatterji, D.C. & Gallelli, J.F. Thermal and photolytic decomposition of methotrexate in aqueous solutions. J. Pharm. Sci. 67, 526–531 (1978).
Gherardin, N.A. et al. Diversity of T cells restricted by the MHC class I-related molecule MR1 facilitates differential antigen recognition. Immunity 44, 32–45 (2016).
Chen, Z. et al. Mucosal-associated invariant T-cell activation and accumulation after in vivo infection depends on microbial riboflavin synthesis and co-stimulatory signals. Mucosal Immunol. http://dx.doi.org/10.1038/mi.2016.39 (4 May 2016).
Chua, B.Y. et al. The use of a TLR2 agonist-based adjuvant for enhancing effector and memory CD8 T-cell responses. Immunol. Cell Biol. 92, 377–383 (2014).
McWilliam, H.E.G. et al. The intracellular pathway for the presentation of vitamin B-related antigens by the antigen-presenting molecule MR1. Nat. Immunol. 17, 531–537 (2016).
Zelcer, S. et al. Methotrexate levels and outcome in osteosarcoma. Pediatr. Blood Cancer 44, 638–642 (2005).
Romão, V.C., Lima, A., Bernardes, M., Canhão, H. & Fonseca, J.E. Three decades of low-dose methotrexate in rheumatoid arthritis: can we predict toxicity? Immunol. Res. 60, 289–310 (2014).
Davies, N.M. & Anderson, K.E. Clinical pharmacokinetics of diclofenac. Therapeutic insights and pitfalls. Clin. Pharmacokinet. 33, 184–213 (1997).
Krasniqi, V., Dimovski, A., Domjanović Iva, K., Bilić, I. & Božina, N. How polymorphisms of the cytochrome P450 genes affect ibuprofen and diclofenac metabolism and toxicity. Arch. Indust. Hyg. Toxico. 67, 1 (2016).
Bharadwaj, M. et al. Drug hypersensitivity and human leukocyte antigens of the major histocompatibility complex. Annu. Rev. Pharmacol. Toxicol. 52, 401–431 (2012).
Skypala, I.J., Williams, M., Reeves, L., Meyer, R. & Venter, C. Sensitivity to food additives, vaso-active amines and salicylates: a review of the evidence. Clin. Transl. Allergy 5, 34 (2015).
Hawkins, P.C.D., Skillman, A.G. & Nicholls, A. Comparison of shape-matching and docking as virtual screening tools. J. Med. Chem. 50, 74–82 (2007).
Fehlner, P.F., Bencsath, A., Lam, T. & King, T.P. The photodecomposition of aminopterin. J. Immunol. Methods 101, 141–145 (1987).
Eckle, S.B. et al. A molecular basis underpinning the T cell receptor heterogeneity of mucosal-associated invariant T cells. J. Exp. Med. 211, 1585–1600 (2014).
Reantragoon, R. et al. Structural insight into MR1-mediated recognition of the mucosal associated invariant T cell receptor. J. Exp. Med. 209, 761–774 (2012).
Gras, S. et al. The shaping of T cell receptor recognition by self-tolerance. Immunity 30, 193–203 (2009).
Heemskerk, M.H.M. et al. Redirection of antileukemic reactivity of peripheral T lymphocytes using gene transfer of minor histocompatibility antigen HA-2-specific T-cell receptor complexes expressing a conserved α joining region. Blood 102, 3530–3540 (2003).
Gras, S. et al. Allelic polymorphism in the T cell receptor and its impact on immune responses. J. Exp. Med. 207, 1555–1567 (2010).
Szymczak, A.L. et al. Correction of multi-gene deficiency in vivo using a single 'self-cleaving' 2A peptide-based retroviral vector. Nat. Biotechnol. 22, 589–594 (2004).
Kawachi, I., Maldonado, J., Strader, C. & Gilfillan, S. MR1-restricted V alpha 19i mucosal-associated invariant T cells are innate T cells in the gut lamina propria that provide a rapid and diverse cytokine response. J. Immunol. 176, 1618–1627 (2006).
Huang, S. et al. Evidence for MR1 antigen presentation to mucosal-associated invariant T cells. J. Biol. Chem. 280, 21183–21193 (2005).
Hoiseth, S.K. & Stocker, B.A.D. Aromatic-dependent Salmonella typhimurium are non-virulent and effective as live vaccines. Nature 291, 238–239 (1981).
Kabsch, W. XDS. Acta Crystallogr. D Biol. Crystallogr. 66, 125–132 (2010).
Winn, M.D. et al. Overview of the CCP4 suite and current developments. Acta Crystallogr. D Biol. Crystallogr. 67, 235–242 (2011).
McCoy, A.J. et al. Phaser crystallographic software. J. Appl. Crystallogr. 40, 658–674 (2007).
Adams, P.D. et al. PHENIX: a comprehensive Python-based system for macromolecular structure solution. Acta Crystallogr. D Biol. Crystallogr. 66, 213–221 (2010).
Emsley, P., Lohkamp, B., Scott, W.G. & Cowtan, K. Features and development of Coot. Acta Crystallogr. D Biol. Crystallogr. 66, 486–501 (2010).
Chen, V.B. et al. MolProbity: all-atom structure validation for macromolecular crystallography. Acta Crystallogr. D Biol. Crystallogr. 66, 12–21 (2010).
Acknowledgements
We thank the staff at the Australian Synchrotron for assistance with data collection; the staff at the Monash Macromolecular crystallization facility; and T. Hansen (University of Washington) and W.J. Yankelevich (US Food and Drug Administration) for the 26.5 hybridoma. Supported by The University of Melbourne (S.E.), the Australian National Health and Medical Research Council (1020770 and 1027369 to D.I.G and D.P.F.; 1044215 to A.W.P.; 1113293 to J.M.; and 1125493 to J.R.), the Australian Research Council (CE140100011 and DE170100407 to S.E.; FT160100083 to A.J.C.; and FL160100049 to J.R.), the Leukaemia Foundation of Australia (N.A.G.) and Cancer Council Victoria (N.A.G.).
Author information
Authors and Affiliations
Contributions
A.N.K., S.B.G.E. and W.X. designed, performed and analyzed experiments; L.L., V.A.H., J.Y.W.M., B.S.M., T.P., R.W.B., Z.C., H.W., C.D'S., L.K.-N., N.A.G., D.I.G, L.K., A.J.C. and A.W.P. performed experiments, analyzed data and/or provided key reagents for this study; and D.P.F., J.M. and J.R. supervised experiments and wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 In silico screening of MR1 ligands.
a) Superimposition of 9 drugs (sticks) on 5-OP-RU. These drugs are consistent with the shape matching to 6-FP. Colored chemical structures of drugs (e.g. Acyclovir, Lamivudine, Methotrexate) correspond to colored sticks, with other drugs shown in grey. b) Side view of all 20 virtual screening drug hits (shown as surfaces) in complementarity with the MR1 binding site. Carbon (white); oxygen (red); nitrogen (blue); chlorine (green); fluorine (cyan). c) Classification of twenty-two representative structures of active compounds (including 9 drugs) according to their chemical substructures: pyrimidines (black), phenols/anilines (green), enones (red), aromatic aldehydes (orange), aromatic carboxylates (olive), quinones (dark blue), flavones (light blue), isoflavones (pink).
Supplementary Figure 2 Upregulation of MR1 expression and activation of MAIT cells.
a) Graphical display of percentages MR1 upregulating versus agonistic compounds identified as part of the functional screen in Figure 2a. b) gMFI of 26.5 (mean of triplicate samples and SEM) and isotype control (single samples) staining at 7 hours for Nil, vehicle controls (V) and ligands (left panel). Repeat experiment of Figure 2c including 5-F-SA (right panel) showing single samples. c) First 2 rows: Drug/small molecule dose dependent inhibition of Jurkat.MAIT-#6 and Jurkat.MAIT-C-F7 activated by 5-OP-RU in the presence of C1R.MR1 cells and assayed by flow cytometric staining for CD69 as a marker of activation. Displayed is gMFI CD69 fold of background control for one representative of three experiments. 5-OP-RU activation with nil inhibitor/activator was assayed in triplicate displaying mean and SEM (error bars). Third row: gMFI of CD69 (mean of triplicate samples and SEM) for Nil (PBS), vehicle controls with maximum concentration of 5-OP-RU, and ligands co-incubated with PMA/Ionomycin are displayed for Jurkat.MAIT-A-F7, Jurkat.MAIT-#6 and Jurkat.MAIT-C-F7. d) IL-2 production in the presence of PBS, vehicle controls with maximum concentration of 5-OP-RU, and ligands co-incubated with PMA/Ionomycin. Displayed are mean of triplicate samples except for ligands co-incubated with PMA/Ionomycin where single samples are shown. e) Repeat experiment of Figure 2d/f including in addition gMFI of CD69 (mean of triplicate samples and SEM) for vehicle controls with maximum concentration of 5-OP-RU, ligands and ligands co-incubated with PMA/Ionomycin (left panel). In parallel the effect of ligands and vehicles on Jurkat.CD8.LC13 activation by C1R.HLA-B*08:01 in the presence of FLR peptide was tested (right panel).
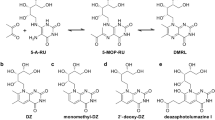
Supplementary Figure 3 Degradation of 2,4-DA-6-FP.
a) Chemical structures of aminopterin/folic acid (I) as they decompose to form respective formyl pterin (II) and aminobenzoylglutamic acid (III). The aldehyde on (II) then further degrades to carboxylic acid (IV). b) Absorbance spectra of aminopterin after exposure to a fluorescent lamp for 0h (green), 18h (orange) and 48h (red). c&d) MR1 surface upregulation by C1R.MR1 cells treated with 20μM or 2μM of photodegraded aminopterin from (b), shown as histogram of 26.5 (c) and as MFI 26.5-fold of PBS vehicle control (mean of triplicate samples with SEM). Representative of two separate experiments. e) Mass spectra and elemental analysis of compound extracted from MR1 refolded in the presence of photodegraded aminopterin compared with theoretical spectra for 2,4-DA-6-FP.
Supplementary Figure 4 Activation of MR1-restricted T cell lines by DCF and DCF metabolites.
Effect of Diclofenac and its metabolites on MR1 restricted T cell lines and Jurkat.CD8.LC13 activation by C1R.HLA-B*08:01 co-incubated with FLR peptide. Displayed are fold of background MFI CD69 (a) or MFI CD69 (b) for one representative of two experiments.
Supplementary Figure 5 Inhibition of the activation of MAIT cells by drugs and drug-related molecules ex vivo.
a) % cytokine production gated on live, CD3+, TRAV1-2+ 5-OP-RU-MR1-tetramer- (representative of non MAIT T cells) or TRAV1-2+ 5-OP-RU-MR1-tetramer+ (MAIT cells) cells. Samples include titrating amounts of 5-OP-RU, vehicle controls in the presence of maximum concentration of 5-OP-RU, Nil (PBS), and maximum concentration of inhibitors in the presence or absence of PMA/Ionomycin stimulus. Displayed is data of one representative donor. b) % CTV dilution gated on live, CD3+, TRAV1-2+ 5-OP-RU-MR1-tetramer- (representative of non MAIT T-cells) or TRAV1-2+ 5-OP-RU-MR1-tetramer+ (MAIT cells) cells. Samples include titrating amounts of 5-OP-RU (triplicate samples, SEM) and Nil (triplicate samples, SEM), vehicles (triplicate samples, SEM) and maximum concentrations of inhibitors in the presence (triplicate samples, SEM) or absence (single samples) of plate bound CD3/CD28. Displayed is data of one representative donor.
Supplementary Figure 6 Inhibition of the activation of MAIT cells by small molecules in vivo.
(a) Inhibitory effect of intranasally administered Ac-6-FP and 3-F-SA on MAIT cell accumulation in the lungs of C57BL/6 mice upon 5-OP-RU and Salm.BRD509ΔribDH stimulus. Matching data in Figure 4c, absolute numbers of MAIT cells and non-MAIT αβ T cells (mean values +/- SEM as error bars of four mice as well as CFU counts in the lungs are shown. (b) Repeat experiment at the maximum inhibitor concentration including in addition IL17 production by non-MAIT αβ T cells in response to Salm.BRD509ΔribDH stimulus in the presence or absence of inhibitors.
Supplementary Figure 7 TCR contacts with MR1.
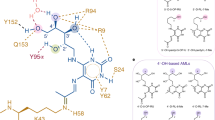
Contact regions of the CDR1α(teal), CDR2α(pink), CDR3α(yellow), CDR1β(cyan), CDR2β(red), CDR3β(orange) and framework residues (slate and deep purple for α- and β-chains, respectively) of A-F7 MAIT TCR on MR1 (white surface), which is presenting 5-OP-RU (A), 6-FP (B), 2,4-DA-6-FP (C), 2-OH-1-NA (D), HMB (E), 3-F-SA (F), DCF (G) or 5-OH-DCF (H).
Supplementary Figure 8 Chemical synthesis of metabolites.
Synthesis of 2,4-diamino-6-formylpteridine (A), 4′,5-dihydroxy diclofenac (B) and 5-hydroxy diclofenac (C).
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–8 and Supplementary Tables 1–8 (PDF 2870 kb)
Rights and permissions
About this article
Cite this article
Keller, A., Eckle, S., Xu, W. et al. Drugs and drug-like molecules can modulate the function of mucosal-associated invariant T cells. Nat Immunol 18, 402–411 (2017). https://doi.org/10.1038/ni.3679
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.3679
This article is cited by
-
MR1 antigen presentation to MAIT cells and other MR1-restricted T cells
Nature Reviews Immunology (2024)
-
Improvement of binding pose prediction of the MR1 covalent ligands by inclusion of simple pharmacophore constraints and structural waters in the docking process
3 Biotech (2023)
-
Deaza-modification of MR1 ligands modulates recognition by MR1-restricted T cells
Scientific Reports (2022)
-
TAPBPR employs a ligand-independent docking mechanism to chaperone MR1 molecules
Nature Chemical Biology (2022)
-
MAIT cells and their implication in human oral diseases
Inflammation Research (2022)