Abstract
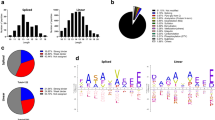
The structural characteristics of the engagement of major histocompatibility complex (MHC) class II–restricted self antigens by autoreactive T cell antigen receptors (TCRs) is established, but how autoimmune TCRs interact with complexes of self peptide and MHC class I has been unclear. Here we examined how CD8+ T cells kill human islet beta cells in type 1 diabetes via recognition of a human leukocyte antigen HLA-A*0201–restricted glucose-sensitive preproinsulin peptide by the autoreactive TCR 1E6. Rigid 'lock-and-key' binding underpinned the 1E6–HLA-A*0201–peptide interaction, whereby 1E6 docked similarly to most MHC class I–restricted TCRs. However, this interaction was extraordinarily weak because of limited contacts with MHC class I. TCR binding was highly peptide centric, dominated by two residues of the complementarity-determining region 3 (CDR3) loops that acted as an 'aromatic-cap' over the complex of peptide and MHC class I (pMHCI). Thus, highly focused peptide-centric interactions associated with suboptimal TCR-pMHCI binding affinities might lead to thymic escape and potential CD8+ T cell–mediated autoreactivity.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
References
Nejentsev, S. et al. Localization of type 1 diabetes susceptibility to the MHC class I genes HLA-B and HLA-A. Nature 450, 887–892 (2007).
Wucherpfennig, K.W. & Sethi, D. T cell receptor recognition of self and foreign antigens in the induction of autoimmunity. Semin. Immunol. 23, 84–91 (2011).
Kappler, J.W., Roehm, N. & Marrack, P. T cell tolerance by clonal elimination in the thymus. Cell 49, 273–280 (1987).
Rudolph, M.G., Stanfield, R.L. & Wilson, I.A. How TCRs bind MHCs, peptides, and coreceptors. Annu. Rev. Immunol. 24, 419–466 (2006).
Colf, L.A. et al. How a single T cell receptor recognizes both self and foreign MHC. Cell 129, 135–146 (2007).
Feng, D., Bond, C.J., Ely, L.K., Maynard, J. & Garcia, K.C. Structural evidence for a germline-encoded T cell receptor-major histocompatibility complex interaction 'codon'. Nat. Immunol. 8, 975–983 (2007).
Sami, M. et al. Crystal structures of high affinity human T-cell receptors bound to peptide major histocompatibility complex reveal native diagonal binding geometry. Protein Eng. Des. Sel. 20, 397–403 (2007).
Borbulevych, O.Y. et al. T cell receptor cross-reactivity directed by antigen-dependent tuning of peptide-MHC molecular flexibility. Immunity 31, 885–896 (2009).
Cole, D.K. et al. Germ line-governed recognition of a cancer epitope by an immunodominant human T-cell receptor. J. Biol. Chem. 284, 27281–27289 (2009).
Gras, S. et al. Structural bases for the affinity-driven selection of a public TCR against a dominant human cytomegalovirus epitope. J. Immunol. 183, 430–437 (2009).
Miles, J.J. et al. Genetic and structural basis for selection of a ubiquitous T cell receptor deployed in epstein-barr virus infection. PLoS Pathog. 6, e1001198 (2010).
Borbulevych, O.Y., Piepenbrink, K.H. & Baker, B.M. Conformational melding permits a conserved binding geometry in TCR recognition of foreign and self molecular mimics. J. Immunol. 186, 2950–2958 (2011).
Sethi, D.K. et al. A highly tilted binding mode by a self-reactive T cell receptor results in altered engagement of peptide and MHC. J. Exp. Med. 208, 91–102 (2011).
Yin, Y., Li, Y., Kerzic, M.C., Martin, R. & Mariuzza, R.A. Structure of a TCR with high affinity for self-antigen reveals basis for escape from negative selection. EMBO J. 30, 1137–1148 (2011).
Godfrey, D.I., Rossjohn, J. & McCluskey, J. The fidelity, occasional promiscuity, and versatility of T cell receptor recognition. Immunity 28, 304–314 (2008).
Hahn, M., Nicholson, M.J., Pyrdol, J. & Wucherpfennig, K.W. Unconventional topology of self peptide-major histocompatibility complex binding by a human autoimmune T cell receptor. Nat. Immunol. 6, 490–496 (2005).
Li, Y. et al. Structure of a human autoimmune TCR bound to a myelin basic protein self-peptide and a multiple sclerosis-associated MHC class II molecule. EMBO J. 24, 2968–2979 (2005).
Willcox, A., Richardson, S.J., Bone, A.J., Foulis, A.K. & Morgan, N.G. Analysis of islet inflammation in human type 1 diabetes. Clin. Exp. Immunol. 155, 173–181 (2009).
DiLorenzo, T.P. & Serreze, D.V. The good turned ugly: immunopathogenic basis for diabetogenic CD8+ T cells in NOD mice. Immunol. Rev. 204, 250–263 (2005).
Skowera, A. et al. CTLs are targeted to kill beta cells in patients with type 1 diabetes through recognition of a glucose-regulated preproinsulin epitope. J. Clin. Invest. 118, 3390–3402 (2008).
Foulis, A.K., Farquharson, M.A. & Meager, A. Immunoreactive α-interferon in insulin-secreting beta cells in type 1 diabetes mellitus. Lancet 2, 1423–1427 (1987).
Velthuis, J.H. et al. Simultaneous detection of circulating autoreactive CD8+ T-cells specific for different islet cell-associated epitopes using combinatorial MHC multimers. Diabetes 59, 1721–1730 (2010).
Cole, D.K. et al. Modification of MHC anchor residues generates heteroclitic peptides that alter TCR binding and T cell recognition. J. Immunol. 185, 2600–2610 (2010).
Cole, D.K. et al. Human TCR-binding affinity is governed by MHC class restriction. J. Immunol. 178, 5727–5734 (2007).
Armstrong, K.M., Piepenbrink, K.H. & Baker, B.M. Conformational changes and flexibility in T-cell receptor recognition of peptide-MHC complexes. Biochem. J. 415, 183–196 (2008).
Tynan, F.E. et al. A T cell receptor flattens a bulged antigenic peptide presented by a major histocompatibility complex class I molecule. Nat. Immunol. 8, 268–276 (2007).
Tynan, F.E. et al. T cell receptor recognition of a 'super-bulged' major histocompatibility complex class I-bound peptide. Nat. Immunol. 6, 1114–1122 (2005).
Dai, S. et al. Crossreactive T Cells spotlight the germline rules for αβ T cell-receptor interactions with MHC molecules. Immunity 28, 324–334 (2008).
Wu, L.C., Tuot, D.S., Lyons, D.S., Garcia, K.C. & Davis, M.M. Two-step binding mechanism for T-cell receptor recognition of peptide MHC. Nature 418, 552–556 (2002).
Burrows, S.R. et al. Hard wiring of T cell receptor specificity for the major histocompatibility complex is underpinned by TCR adaptability. Proc. Natl. Acad. Sci. USA 107, 10608–10613 (2010).
Pang, S.S. et al. The structural basis for autonomous dimerization of the pre-T-cell antigen receptor. Nature 467, 844–848 (2010).
Borg, N.A. et al. CD1d-lipid-antigen recognition by the semi-invariant NKT T-cell receptor. Nature 448, 44–49 (2007).
Godfrey, D.I. et al. Antigen recognition by CD1d-restricted NKT T cell receptors. Semin. Immunol. 22, 61–67 (2010).
Borg, N.A. et al. The CDR3 regions of an immunodominant T cell receptor dictate the 'energetic landscape' of peptide-MHC recognition. Nat. Immunol. 6, 171–180 (2005).
Garcia, K.C., Adams, J.J., Feng, D. & Ely, L.K. The molecular basis of TCR germline bias for MHC is surprisingly simple. Nat. Immunol. 10, 143–147 (2009).
Scott-Browne, J.P., White, J., Kappler, J.W., Gapin, L. & Marrack, P. Germline-encoded amino acids in the αβ T-cell receptor control thymic selection. Nature 458, 1043–1046 (2009).
Todd, J.A. et al. Robust associations of four new chromosome regions from genome-wide analyses of type 1 diabetes. Nat. Genet. 39, 857–864 (2007).
Pugliese, A. et al. The insulin gene is transcribed in the human thymus and transcription levels correlated with allelic variation at the INS VNTR-IDDM2 susceptibility locus for type 1 diabetes. Nat. Genet. 15, 293–297 (1997).
Thébault-Baumont, K. et al. Acceleration of type 1 diabetes mellitus in proinsulin 2-deficient NOD mice. J. Clin. Invest. 111, 851–857 (2003).
Wooldridge, L. et al. A single autoimmune T-cell receptor recognizes over a million different peptides. J. Biol. Chem. 287, 1168–1177 (2012).
Foulis, A.K., Farquharson, M.A. & Hardman, R. Aberrant expression of class II major histocompatibility complex molecules by B cells and hyperexpression of class I major histocompatibility complex molecules by insulin containing islets in type 1 (insulin-dependent) diabetes mellitus. Diabetologia 30, 333–343 (1987).
Harkiolaki, M. et al. T cell-mediated autoimmune disease due to low-affinity crossreactivity to common microbial peptides. Immunity 30, 348–357 (2009).
Huang, G.C. et al. The development of new density gradient media for purifying human islets and islet-quality assessments. Transplantation 77, 143–145 (2004).
Boulter, J.M. et al. Stable, soluble T-cell receptor molecules for crystallization and therapeutics. Protein Eng. 16, 707–711 (2003).
Cole, D.K. et al. T cell receptor engagement of peptide-major histocompatibility complex class I does not modify CD8 binding. Mol. Immunol. 45, 2700–2709 (2008).
Winter, G. xia2: an expert system for macromolecular crystallography data reduction. J. Appl. Crystallogr. 43, 186–190 (2010).
Collaborative Computational Project. N. The CCP4 suite: programs for protein crystallography. Acta Crystallogr. D Biol. Crystallogr. 50, 760–763 (1994).
Trapani, S. & Navaza, J. AMoRe: classical and modern. Acta Crystallogr. D Biol. Crystallogr. 64, 11–16 (2008).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Delano, W.L. The PyMOL Molecular Graphics System (DeLano Scientific, San Carlos, California, 2002).
Acknowledgements
We thank the staff at Diamond Light Source for facilities and support; M. Zhao for help preparing human islets; and J. Todd for INS genotyping. Supported by the Biotechnology and Biological Sciences Research Council (BB/H001085/1), the Wellcome Trust (WT086716 to A.K.S.; WT079848 to L.W.; and WT095767 to D.K.C.), the Juvenile Diabetes Research Foundation (7-2005-877 and 1-2007-1803 to M.P.; and 17-2009-806 to A.K.S., M.P., D.A.P. and A.S.), the European Union Seventh Framework Programme (241447 NAIMIT), the National Institute for Health Research Comprehensive Biomedical Research Center at Guy's & St. Thomas' National Health Service Foundation Trust and King's College London, Research Councils UK (P.J.R.), the National Health and Medical Research Council (J.R.), the Welsh Office of Research and Development (J.J.M.) and the Medical Research Council (D.A.P.).
Author information
Authors and Affiliations
Contributions
D.K.C., P.J.R., M.P. and A.K.S., leadership and project conception; D.K.C. and A.M.B., crystallization and surface plasmon resonance studies; D.K.C., A.M.B., F.M., J.J.M., P.J.R. and A.F., data and crystal collection; D.K.C., S.G. and P.J.R., crystallographic analysis; A.S., J.W.D. and R.R.K., cell experiments; G.C.H., preparation of human islets; G.D., T cell culture; E.G., N.L. and P.E.M., cloning of the 1E6 TCR; D.K.C., A.K.S., M.P., P.J.R., J.J.M., J.R. and S.G., manuscript authorship; L.W. and B.K.J., intellectual input; and A.K.S., M.P., D.A.P. and J.R., study funding.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–5 and Table 1 (PDF 3039 kb)
Rights and permissions
About this article
Cite this article
Bulek, A., Cole, D., Skowera, A. et al. Structural basis for the killing of human beta cells by CD8+ T cells in type 1 diabetes. Nat Immunol 13, 283–289 (2012). https://doi.org/10.1038/ni.2206
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2206
This article is cited by
-
Non-mutational neoantigens in disease
Nature Immunology (2024)
-
Type 1 diabetes mellitus as a disease of the β-cell (do not blame the immune system?)
Nature Reviews Endocrinology (2021)
-
Structural basis for oligoclonal T cell recognition of a shared p53 cancer neoantigen
Nature Communications (2020)
-
eIF5A inhibition influences T cell dynamics in the pancreatic microenvironment of the humanized mouse model of Type 1 Diabetes
Scientific Reports (2019)
-
Targeted suppression of autoreactive CD8+ T-cell activation using blocking anti-CD8 antibodies
Scientific Reports (2016)