Abstract
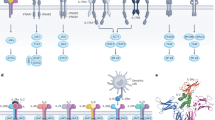
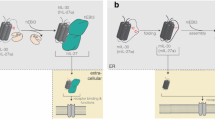
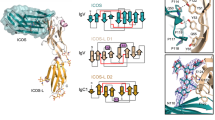
Interleukin 15 (IL-15) and IL-2 have distinct immunological functions even though both signal through the receptor subunit IL-2Rβ and the common γ-chain (γc). Here we found that in the structure of the IL-15–IL-15Rα–IL-2Rβ–γc quaternary complex, IL-15 binds to IL-2Rβ and γc in a heterodimer nearly indistinguishable from that of the IL-2–IL-2Rα–IL-2Rβ–γc complex, despite their different receptor-binding chemistries. IL-15Rα substantially increased the affinity of IL-15 for IL-2Rβ, and this allostery was required for IL-15 trans signaling. Consistent with their identical IL-2Rβ–γc dimer geometries, IL-2 and IL-15 showed similar signaling properties in lymphocytes, with any differences resulting from disparate receptor affinities. Thus, IL-15 and IL-2 induced similar signals, and the cytokine specificity of IL-2Rα versus IL-15Rα determined cellular responsiveness. Our results provide new insights for the development of specific immunotherapeutics based on IL-15 or IL-2.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout








Similar content being viewed by others
References
Waldmann, T.A. The biology of interleukin-2 and interleukin-15: implications for cancer therapy and vaccine design. Nat. Rev. Immunol. 6, 595–601 (2006).
Rochman, Y., Spolski, R. & Leonard, W.J. New insights into the regulation of T cells by gamma(c) family cytokines. Nat. Rev. Immunol. 9, 480–490 (2009).
Sneller, M.C. et al. IL-15 administered by continuous infusion to rhesus macaques induces massive expansion of CD8+ T effector memory population in peripheral blood. Blood 118, 6845–6848 (2011).
Liao, W., Lin, J.X. & Leonard, W.J. IL-2 family cytokines: new insights into the complex roles of IL-2 as a broad regulator of T helper cell differentiation. Curr. Opin. Immunol. 23, 598–604 (2011).
Wuest, S.C. et al. A role for interleukin-2 trans-presentation in dendritic cell-mediated T cell activation in humans, as revealed by daclizumab therapy. Nat. Med. 17, 604–609 (2011).
Stonier, S.W. & Schluns, K.S. Trans. -presentation: a novel mechanism regulating IL-15 delivery and responses. Immunol. Lett. 127, 85–92 (2010).
Marks-Konczalik, J. et al. IL-2-induced activation-induced cell death is inhibited in IL-15 transgenic mice. Proc. Natl. Acad. Sci. USA 97, 11445–11450 (2000).
Syed, R.S. et al. Efficiency of signalling through cytokine receptors depends critically on receptor orientation. Nature 395, 511–516 (1998).
Wang, X., Rickert, M. & Garcia, K.C. Structure of the quaternary complex of interleukin-2 with its alpha, beta, and gammac receptors. Science 310, 1159–1163 (2005).
Chirifu, M. et al. Crystal structure of the IL-15-IL-15Ralpha complex, a cytokine-receptor unit presented in trans. Nat. Immunol. 8, 1001–1007 (2007).
Walter, T.S. et al. Lysine methylation as a routine rescue strategy for protein crystallization. Structure 14, 1617–1622 (2006).
LaPorte, S.L. et al. Molecular and structural basis of cytokine receptor pleiotropy in the interleukin-4/13 system. Cell 132, 259–272 (2008).
Collins, L. et al. Identification of specific residues of human interleukin 2 that affect binding to the 70-kDa subunit (p70) of the interleukin 2 receptor. Proc. Natl. Acad. Sci. USA 85, 7709–7713 (1988).
Eisenman, J. et al. Interleukin-15 interactions with interleukin-15 receptor complexes: characterization and species specificity. Cytokine 20, 121–129 (2002).
Pettit, D.K. et al. Structure-function studies of interleukin 15 using site-specific mutagenesis, polyethylene glycol conjugation, and homology modeling. J. Biol. Chem. 272, 2312–2318 (1997).
Zurawski, S.M. et al. Definition and spatial location of mouse interleukin-2 residues that interact with its heterotrimeric receptor. EMBO J. 12, 5113–5119 (1993).
Boulanger, M.J., Bankovich, A.J., Kortemme, T., Baker, D. & Garcia, K.C. Convergent mechanisms for recognition of divergent cytokines by the shared signaling receptor gp130. Mol. Cell 12, 577–589 (2003).
McFarland, B.J. & Strong, R.K. Thermodynamic analysis of degenerate recognition by the NKG2D immunoreceptor: not induced fit but rigid adaptation. Immunity 19, 803–812 (2003).
Dubois, S., Mariner, J., Waldmann, T.A. & Tagaya, Y. IL-15Ralpha recycles and presents IL-15 In trans to neighboring cells. Immunity 17, 537–547 (2002).
Hanick, N.A. et al. Elucidation of the interleukin-15 binding site on its alpha receptor by NMR. Biochemistry 46, 9453–9461 (2007).
Mortier, E. et al. Soluble interleukin-15 receptor alpha (IL-15R alpha)-sushi as a selective and potent agonist of IL-15 action through IL-15R beta/gamma. Hyperagonist IL-15 x IL-15R alpha fusion proteins. J. Biol. Chem. 281, 1612–1619 (2006).
Rubinstein, M.P. et al. Converting IL-15 to a superagonist by binding to soluble IL-15R{alpha}. Proc. Natl. Acad. Sci. USA 103, 9166–9171 (2006).
Rickert, M., Wang, X., Boulanger, M.J., Goriatcheva, N. & Garcia, K.C. The structure of interleukin-2 complexed with its alpha receptor. Science 308, 1477–1480 (2005).
Levin, A.M. et al. Exploiting a natural conformational switch to engineer an interleukin-2 'superkine'. Nature 484, 529–533 (2012).
Balasubramanian, S. et al. Ligand binding kinetics of IL-2 and IL-15 to heteromers formed by extracellular domains of the three IL-2 receptor subunits. Int. Immunol. 7, 1839–1849 (1995).
Thanos, C.D., Randal, M. & Wells, J.A. Potent small-molecule binding to a dynamic hot spot on IL-2. J. Am. Chem. Soc. 125, 15280–15281 (2003).
Bowman, G.R., Huang, X. & Pande, V.S. Using generalized ensemble simulations and Markov state models to identify conformational states. Methods 49, 197–201 (2009).
Zambricki, E. et al. Signaling T-cell survival and death by IL-2 and IL-15. Am. J. Transplant. 5, 2623–2631 (2005).
Castro, I., Yu, A., Dee, M.J. & Malek, T.R. The basis of distinctive IL-2- and IL-15-dependent signaling: weak CD122-dependent signaling favors CD8+ T central-memory cell survival but not T effector-memory cell development. J. Immunol. 187, 5170–5182 (2011).
Cornish, G.H., Sinclair, L.V. & Cantrell, D.A. Differential regulation of T-cell growth by IL-2 and IL-15. Blood 108, 600–608 (2006).
Lindemann, M.J., Hu, Z., Benczik, M., Liu, K.D. & Gaffen, S.L. Differential regulation of the IL-17 receptor by gammac cytokines: inhibitory signaling by the phosphatidylinositol 3-kinase pathway. J. Biol. Chem. 283, 14100–14108 (2008).
Demirci, G. & Li, X.C. IL-2 and IL-15 exhibit opposing effects on Fas mediated apoptosis. Cell Mol. Immunol. 1, 123–128 (2004).
Krieg, C., Letourneau, S., Pantaleo, G. & Boyman, O. Improved IL-2 immunotherapy by selective stimulation of IL-2 receptors on lymphocytes and endothelial cells. Proc. Natl. Acad. Sci. USA 107, 11906–11911 (2010).
Ahmadzadeh, M. & Rosenberg, S.A. IL-2 administration increases CD4+ CD25(hi) Foxp3+ regulatory T cells in cancer patients. Blood 107, 2409–2414 (2006).
Bessard, A., Sole, V., Bouchaud, G., Quemener, A. & Jacques, Y. High antitumor activity of RLI, an interleukin-15 (IL-15)-IL-15 receptor alpha fusion protein, in metastatic melanoma and colorectal cancer. Mol. Cancer Ther. 8, 2736–2745 (2009).
Minor, W., Cymborowski, M., Otwinowski, Z. & Chruszcz, M. HKL-3000: the integration of data reduction and structure solution–from diffraction images to an initial model in minutes. Acta Crystallogr. D Biol. Crystallogr. 62, 859–866 (2006).
Adams, P.D. et al. PHENIX: building new software for automated crystallographic structure determination. Acta Crystallogr. D Biol. Crystallogr. 58, 1948–1954 (2002).
Emsley, P. & Cowtan, K. Coot: model-building tools for molecular graphics. Acta Crystallogr. D Biol. Crystallogr. 60, 2126–2132 (2004).
Krissinel, E. & Henrick, K. Inference of macromolecular assemblies from crystalline state. J. Mol. Biol. 372, 774–797 (2007).
Hess, B., Kutzner, C., van der Spoel, D. & Lindahl, E. GROMACS 4: Algorithms for highly efficient, load-balanced, and scalable molecular simulation. J. Chem. Theory Comput. 4, 435–447 (2008).
Duan, Y. et al. A point-charge force field for molecular mechanics simulations of proteins based on condensed-phase quantum mechanical calculations. J. Comput. Chem. 24, 1999–2012 (2003).
Hess, B. P-LINCS: A parallel linear constraint solver for molecular simulation. J. Chem. Theory Comput. 4, 116–122 (2008).
Darden, T., York, D. & Pedersen, L. A smooth particle mesh Ewald potential. J. Chem. Phys. 103, 3014–3021 (1995).
Bussi, G., Donadio, D. & Parrinello, M. Canonical sampling through velocity rescaling. J. Chem. Phys. 126, 014101 (2007).
Berendsen, H.J.C., Postma, P.M., van Gunsteren, W.F., DiNola, A. & Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem. Phys. 81, 3684–3690 (1984).
Bowman, G.R., Beauchamp, K.A., Boxer, G. & Pande, V.S. Progress and challenges in the automated construction of Markov state models for full protein systems. J. Chem. Phys. 131, 124101 (2009).
Bowman, G.R., Huang, X. & Pande, V.S. Network models for molecular kinetics and their initial applications to human health. Cell Res. 20, 622–630 (2010).
Noe, F. & Fischer, S. Transition networks for modeling the kinetics of conformational change in macromolecules. Curr. Opin. Struct. Biol. 18, 154–162 (2008).
Krutzik, P.O. & Nolan, G.P. Fluorescent cell barcoding in flow cytometry allows high-throughput drug screening and signaling profiling. Nat. Methods 3, 361–368 (2006).
Tang, F. et al. RNA-Seq analysis to capture the transcriptome landscape of a single cell. Nat. Protoc. 5, 516–535 (2010).
Acknowledgements
We thank E. Long, M. Rubinstein and members of the Leonard and Garcia laboratories for advice and discussions; and N. Goriatcheva, D. Waghray and S. Fischer for technical assistance. Supported by the US National Institutes of Health (R01 AI51321 to K.C.G.; R01 GM062868 to V.S.P.; and National Research Service Award NIH-F30DK094541 to A.M.R.), the American Recovery and Reinvestment Act of 2009 (Public Law 111-5; MRI-R2 to V.S.P.), the Division of Intramural Research of the National Heart, Lung and Blood Institute (US National Institutes of Health; W.J.L., J.-X.L., P.L., S.M. and R.S.), the Stanford Medical Scientist Training Program (NIH-GM07365 to A.M.R.) and the Howard Hughes Medical Institute (K.C.G.).
Author information
Authors and Affiliations
Contributions
A.M.R., D.F. and E.Ö. did crystallographic studies of the IL-15 quaternary complex; A.M.R. and E.Ö. determined and refined that structure; M.R. did SPR measurements; G.R.B. and V.S.P. did and analyzed molecular dynamics simulations; A.M.R. prepared cytokine proteins for signaling and transcriptional studies; A.M.R., S.M., I.M. and R.S. did signaling experiments by flow cytometry with phosphorylation-specific antibodies; A.M.R. did receptor-internalization studies; J.-X.L. and P.L. did and analyzed RNA sequencing transcriptional assays; J.-X.L. confirmed the quantitative PCR; A.M.R., J.-X.L., G.R.B., W.J.L. and K.C.G. designed the experiments; A.M.R., J.-X.L., P.L., S.M. and G.R.B. prepared the figures; A.M.R., W.J.L. and K.C.G. wrote the paper; and W.J.L. and K.C.G. supervised the research.
Corresponding author
Ethics declarations
Competing interests
K.C.G. has filed a patent (US 2011/066911) describing the IL-2 'superkine' H9.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–4 and Tables 1–2 (PDF 1343 kb)
Supplementary Spreadsheet 1
Genes more potently regulated by IL-2. (XLS 188 kb)
Supplementary Spreadsheet 2
Genes more potently regulated by IL-15. (XLS 203 kb)
Rights and permissions
About this article
Cite this article
Ring, A., Lin, JX., Feng, D. et al. Mechanistic and structural insight into the functional dichotomy between IL-2 and IL-15. Nat Immunol 13, 1187–1195 (2012). https://doi.org/10.1038/ni.2449
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ni.2449
This article is cited by
-
Structural insights into IL-11-mediated signalling and human IL6ST variant-associated immunodeficiency
Nature Communications (2024)
-
Human Vγ9Vδ2 T cell expansion and their cytotoxic responses against cholangiocarcinoma
Scientific Reports (2024)
-
New insights into the stemness of adoptively transferred T cells by γc family cytokines
Cell Communication and Signaling (2023)
-
ROGUE: an R Shiny app for RNA sequencing analysis and biomarker discovery
BMC Bioinformatics (2023)
-
Biomimetic nanovaccine-mediated multivalent IL-15 self-transpresentation (MIST) for potent and safe cancer immunotherapy
Nature Communications (2023)