Abstract
Several attributes intuitively considered to be typical mammalian features, such as complex behavior, live birth and malignant disease such as cancer, also appeared several times independently in lower vertebrates. The genetic mechanisms underlying the evolution of these elaborate traits are poorly understood. The platyfish, X. maculatus, offers a unique model to better understand the molecular biology of such traits. We report here the sequencing of the platyfish genome. Integrating genome assembly with extensive genetic maps identified an unexpected evolutionary stability of chromosomes in fish, in contrast to in mammals. Genes associated with viviparity show signatures of positive selection, identifying new putative functional domains and rare cases of parallel evolution. We also find that genes implicated in cognition show an unexpectedly high rate of duplicate gene retention after the teleost genome duplication event, suggesting a hypothesis for the evolution of the behavioral complexity in fish, which exceeds that found in amphibians and reptiles.
Similar content being viewed by others
Main
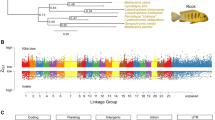
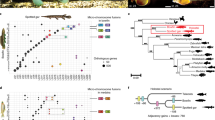
We sequenced the whole genome of a single platyfish female (XX, 2n = 46 chromosomes, Jp163A strain; Fig. 1) from generation 104 of continuous brother-sister matings. Total sequence coverage of 19.6-fold (Supplementary Note) produced an assembly with N50 contig and supercontig lengths of 22 kb and 1.1 Mb, respecively (Supplementary Table 1). Assembly errors, mostly single-nucleotide insertions or deletions, were corrected with Illumina paired-end reads. A total of 669 Mb of the estimated genome length of 750–950 Mb was assembled in contigs. Gene predictions identified 20,366 coding genes, 348 noncoding genes and 28 pseudogenes (Supplementary Note).
(a) Female (top) and male (bottom) platyfish, of strain Jp163A with black pigment spots on the dorsal fin that develop when the activity of an X-chromosomal oncogene is appropriately controlled. In hybrid genotypes, this control is compromised, and malignant melanoma develops from the spots. (b) Phylogenetic position of the platyfish relative to other fish species.
As in other teleosts, transposable elements (TEs) in platyfish were highly diverse, including many families absent in mammals1 and birds (Supplementary Figs. 1–3, Supplementary Tables 2 and 3 and Supplementary Note). We found that 4.8% of the transcriptome was derived from TE sequences representing about 40 different families, indicating that many of the platyfish TEs are most likely still active. The most active TEs were Tc1 DNA transposons (>16,000 copies), followed by the RTE family (>9,000 copies). Notably, we identified several almost-intact envelope-encoding copies of a foamy retrovirus (Spumaviridae) integrated into the platyfish genome (Fig. 2). Foamy viruses are known as exogenous infectious agents in mammals2. Only recently have endogenous foamy virus sequences that may be used to represent a fossil record of infections been described in the genomes of the sloth3 and aye-aye4 in mammals and in the coelacanth5. A foamy virus–like sequence in zebrafish6, a sequence in cod discovered during this work and the platyfish genome sequence reported here show an even broader spectrum of hosts. The molecular phylogeny of foamy viruses is consistent with host phylogeny (Fig. 2). This result supports the notion of an ancient marine evolutionary origin of this type of virus, with possible host-virus coevolution5. The nearly intact copies of foamy virus found in the genomes of some divergent fish species, absent from other sequenced fish genomes, might indicate independent germline introductions through infection. Exogenous foamy virus had not been described in fish; however, our results suggest that exogenous foamy viruses have been and might still be infectious in the fish lineage.
Foamy virus (FV) sequences (light-blue shading) form two distinct phylogenetic groups, one tetrapod specific and one teleost specific. Both groups contain endogenous foamy virus (EFV) sequences (the ewly identified platyfish and cod sequences are highlighted by dark-blue shading). Alignment was carried out with ClustalW (223 amino acids), and the phylogenetic tree was constructed with the PhyML package using maximum-likelihood methods38 with default bootstrap (shown at the beginning of branches) and optimized calculation options. FV, foamy virus; MuERV-L, Mus musculus endogenous retrovirus-L; BAEV, baboon endogenous virus; FENV1, feline endogenous virus 1; EFV, endogenous foamy virus, MLV, murine leukemia virus; HERV-K, human endogenous retrovirus-K; MMTV, mouse mammary tumor virus; HIV-1, human immunodeficiency virus-1. The scale bar represents the number of substitutions per site.
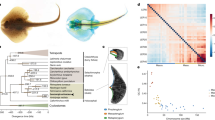
Mammalian chromosome homology maps show a patchwork arrangement of about 35 large conserved synteny blocks on average (but about 80 in dog and 200 in mouse) and numerous small blocks assembled in different combinations among the varied species and spanning over 90 million years of evolution7. We constructed the most extensive meiotic genetic map for any vertebrate yet published, which allowed the ordering of X. maculatus scaffolds and precise conserved synteny analysis comparing fish genomes (Supplementary Note). We used the innovative restriction site–associated DNA (RAD)-tag approach8 to construct a meiotic map consisting of 16,245 polymorphic markers that define 24 linkage groups equivalent to the haploid chromosome number of the platyfish9. Thus, 90.17% of the total sequences in contigs could be assigned a chromosomal position. Long-range comparisons of the order of genes across species10 identified novel evolutionary relationships between platyfish and other teleost chromosomes. Medaka, the closest relative with a sequenced genome, also has 24 chromosomes, and 19 of these showed a strict one-to-one relationship with the platyfish chromosomes (Fig. 3a,b). The remaining five platyfish chromosomes were also each orthologous to a single medaka chromosome, with the exception of one or two short segments (∼1 Mb in length) that were located on another medaka chromosome (Fig. 3c and Supplementary Fig. 4). Thus, quite a few translocations, all very short, have disrupted karyotypes since the divergence of medaka and platyfish 120 million years ago11,12. A similar picture emerged from comparisons of platyfish chromosomes to those of stickleback (divergence 180 million years ago)11,12. These findings detail the previously unknown broad extent to which the genetic content of chromosomes in these teleosts has been conserved over nearly 200 million years of evolution, a conservation much greater than that found in mammals over about half that time7,11,12. This is somewhat unexpected, given the teleost genome duplication (TGD) event, because one might have thought that the illegitimate pairing of paralogous chromosomes (arising from TGD) might have facilitated translocations. The mechanisms that may have mitigated such translocations remain unknown.
(a) The medaka orthologs of genes on X. maculatus chromosome 9 (Xma9) tend to lie on Oryzias latipes chromosome 4 (Ola4), showing that the genic content of these chromosomes has remained intact with no translocations in the 120 million years since the lineages of these species diverged. Each gray dot along the horizontal axis labeled Xma9 represents the position of a platyfish gene whose medaka ortholog (as judged by reciprocal best-BLAST hit analysis) lies directly vertical to the Xma9 gene, plotted on the appropriate medaka chromosome10. (b) Reciprocally, nearly all of the platyfish orthologs of genes on medaka chromosome Ola4 lie on Xma9. (c) Nearly all of the medaka orthologs of Xma19 lie on Ola22, except for a segment about 1 Mb long at position 20 Mb on Ola22 that appears on Ola24 (dashed box).
The platyfish is a well-known model in cancer research13. Its genome contains a tumor control region (TCR), including the oncogene xmrk14 that triggers melanoma development. The TCR also contains the tumor modifier mdl15,16. mdl allelic variants control the body compartment, time of onset and severity of tumors17. In addition, mdl alleles manifest in platyfish as a high diversity of genetically defined pigment patterns. The mapped genome allowed us to rule out many pigment genes as the responsible factors for these sex-associated pigment variants and melanoma modifiers. All known pigment genes18 were present in the XX female platyfish genome; thus, none is Y chromosome specific. Only 6 of the 174 known pigment genes (asip2a, egfrb, muted, myca, rps20 and tfap2a) were located on the X chromosome (Xma21). Of these six, only the proto-oncogene egfrb resided close enough to the melanoma oncogene xmrk (Supplementary Table 4) to be considered a candidate gene for mdl. Indeed, biochemical studies have shown that Egfrb can cooperate with Xmrk19, but the expression levels of these genes are inversely regulated in melanoma20. Further studies are needed to evaluate egfrb function and to find other non-classical pigmentation gene candidates in this genomic region that may control both pigment pattern and melanoma phenotype.
Another so-far-unidentified genetic component of the Xiphophorus melanoma model is the R/Diff gene. R/Diff suppresses melanoma formation in wild platyfish, and the elimination of its expression by interspecies hybridization allows tumor growth. R/Diff was mapped to a 10-cM interval on Xma5 near the cdkn2a/b locus21. Despite the orthologous human CDKN2A gene being a well-described tumor suppressor gene in certain human melanomas22, cdkn2a/b was excluded from being R/Diff because it is not mutated but is instead overexpressed in the Xiphophorus melanoma model23. The Xma5 sequence now defines a number of R/Diff candidate genes for further exploration. For example, scaffold 182 (1,085,500 bp), which harbors cdkn2a/b, contains several genes with high potential of having a role as the R/Diff tumor suppressor (for example, tet2, cxxc4, mtap, topo-rs, mdx4 and pdcd4a). Alternatively, the region may represent a complex locus comprising several genes that act in a synergistic or compensatory manner to regulate the xmrk oncogene, consistent with previous reports of spontaneous and induced carcinogenesis in the many Xiphophorus interspecies hybrid tumor models24,25,26.
Viviparity is an elaborate reproductive mode involving diverse levels of maternal investment in offspring, ranging from fully provisioning eggs before fertilization and retaining them through development to minimally provisioning eggs before fertilization and provisioning them after fertilization via a placenta, as in mammals. The fish family Poeciliidae, a monophyletic clade of more than 260 species27, is unusual in including species that span the spectrum from negligible to extensive post-fertilization provisioning28,29. The platyfish genome is the first from a non-mammalian viviparous vertebrate. We performed analysis in platyfish as well as in a second livebearing fish, the swordtail Xiphophorus hellerii, both of which have well-provisioned eggs before fertilization30,31, of 3 groups of viviparity genes (yolk, placenta and egg coat genes; n = 34) for gene loss and positive selection compared to 4 species of egg-laying teleosts (medaka, tetraodon, stickleback and zebrafish).
In mammals, the rise of viviparity has been proposed to involve the progressive loss of vitellogenins (yolk precursors)32. In platyfish and swordtail, all yolk-related genes (vitellogenins and their transporters/receptors; Supplementary Table 5) were present and evolved under purifying selection, consistent with both species fully provisioning eggs before fertilization, with the exception of one gene that evolved under positive selection, vitellogenin1 (Supplementary Fig. 5a).
Three of 13 platyfish genes, whose mammalian orthologs are related to placenta development, evolved under positive selection (Fig. 4a, Supplementary Fig. 5b–d and Supplementary Table 5). Igf2, which in mouse regulates placenta permeability33, evolved under strong positive selection in platyfish (Fig. 4a), which particularly affected the region distal to the proteolysis site. The igf2 sequence33 was also available from another poeciliid, the desert topminnow Poeciliopsis lucida, which shares a livebearing ancestor with Xiphophorus species but differs in having evolved placentation recently. In the desert topminnow, the same region as in platyfish evolved under positive selection, but the selection was even stronger (Supplementary Fig. 5b), suggesting ongoing molecular adaptive evolution since the two genera containing these fish diverged several million years ago. The two other placental genes, pparg and ncoa6, had multiple regions with signals for positive selection outside known functional domains, suggesting novel regions important for viviparity. The same genes under selection in livebearing fish, however, did not show positive selection signatures when orthologous genes from the egg-laying platypus and from marsupials and placental mammals were analyzed (Supplementary Table 6). This result is in line with the fact that the placentas of mammals and fish are convergent but not homologous structures.
Class 1 sites are under purifying selection (Ka/Ks ratio of ∼0), class 2 sites are under neutral selection (Ka/Ks ratio of ∼1), and class 3 sites are under positive selection in Xiphophorus species. (a) Insulin-like growth factor 2 (IGF2). Colored bars below the plot show known functional domains, and the arrow shows the proteolysis site (between residues 118 and 119). (b) ChoriogeninH minor. Top, comparison of egg-laying versus livebearing fish. Bottom, comparison of placental versus non-placental mammals. The same regions are under positive selection in fishes and mammals.
Zona pellucida (Zpc) genes, which produce a glycoprotein-rich coat surrounding the oocyte plasma membrane, showed the most pronounced changes. alveolin was lost from the platyfish genome. Conversely, choriogeninH minor, choriolysinL, choriolysinH and zvep evolved under positive selection (Fig. 4b, Supplementary Fig. 5e–g and Supplementary Table 5). In Xenopus laevis, Zpc genes control species-specific sperm binding and help ensure that only conspecific sperm released into the aqueous environment fertilizes eggs34. Viviparous fish, however, have internal fertilization, where species-specific sperm recognition would not be as crucial. Compared to egg-laying fish, the eggshell in these fish is expected to have adapted to development inside the mother, as it is no longer essential for protection but must facilitate gas and material exchange. Hatching enzyme genes zvep and choriolysinH showed fast-evolving sites generally located adjacent to the catalytic domains (Supplementary Fig. 4f,g), indicating that, during the evolution of viviparity, these enzymes might have altered interactions with target or regulatory proteins. Notably, in choriogeninH minor, the same regions, in particular in the zona pellucida domain, evolved under positive selection in both mammals and fish (Fig. 4b). This is a noticeable example of how convergent evolution at the molecular level manifests on the physiological and ultimately morphological levels.
Our analyses of the consequences of TGD uncovered a functional class of genes that raised our interest because Xiphophorus fish in particular and teleosts in general show a pronounced high level of behavioral complexity35 that other groups of 'cold-blooded' vertebrates such as amphibians and reptiles do not achieve. Using the platyfish genome and gene annotations from six other sequenced teleosts, we asked whether duplicate gene retention from the TGD event could produce through subfunctionalization (differential retention of ancestral subfunctions) and/or neofunctionalization (acquisition of new subfunctions)36 the acquisition of more complex behaviors. We compared 190 cognition-related genes (Supplementary Table 7 and Supplementary Note) to those involved in pigmentation (133 genes, for which increased gene repertoires have been connected to the high complexity and diversity of teleost coloration) and liver functions (187 genes)18 as controls. Analysis of cognition-related genes showed a high duplicate retention rate of 45% in platyfish and similar values in other teleosts (Fig. 5 and Supplementary Fig. 6) compared to the rates seen for genes related to pigmentation (30%) and liver function (15%). The average duplicate retention rate over all genes in teleost genomes is estimated at 12–24% (ref. 37). We found no bias in genes from all three functional categories (cognition, pigmentation and liver function) that were retained after TGD owing to dosage sensitivity or protein complex membership (Supplementary Tables 8 and 9 and Supplementary Note), but a bias in the cognition genes (but not liver function and pigmentation genes) for particularly large proteins (>1,000 amino acids in length) was found (Supplementary Fig. 7, Supplementary Table 10 and Supplementary Note). Plotting gene losses on the phylogenetic tree showed that cognition gene retention was already fixed shortly after TGD and before teleost diversification. This finding supports the hypothesis that paralog retention from the TGD event may have supported the high level of behavioral complexity in Xiphophorus and other teleosts.
(a) Retention rates for TGD-derived duplicates of genes related to cognition, pigmentation and liver function in seven teleost genomes. Time points during teleost evolution that involve the lineage leading to Xiphophorus are connected by lines. (b) Phylogenetic mapping of gene losses for 190 pairs of cognition-related gene duplicates after TGD. Losses are indicated with negative values. The number of retained TGD paralog pairs for each individual teleost genome is given in parentheses. TGD paralog losses were mapped onto the teleost phylogeny provided by Setiamarga et al.39 following the parsimony principle. The TGD event was set to 350 million years ago. The retention rate of TGD paralogs is defined by the number of pairs of TGD-derived duplicates present in a specific lineage divided by the number of pairs of TGD-derived duplicates present at the time of TGD18.
The platyfish genome sequence and analysis have provided new perspectives for several prominent features of this fish model, including its livebearing reproductive mode, variation in pigmentation patterns, sex chromosome evolution in action, complex behavior and both spontaneous and induced carcinogenesis17. Teleosts dominate the extant fish fauna, and, within teleosts (Fig. 1b), the Poeciliidae family, including platyfish, swordtails, guppies and mollies, is a paradigm of this wide spectrum of adaptations. Our study of this first genome of a poeciliid fish illuminates some teleost evolutionary adaptations and provides an important resource to advance the study of melanoma and other segregating phenotypes.
Methods
Source material.
DNA for genome sequencing was derived from a single female X. maculatus, strain Jp163A (sample XMAC-090115_JP163A) from the Xiphophorus Genetic Stock Center (XGSC) at Texas State University (San Marcos, Texas, USA). The Jp163A line is maintained exclusively by brother-sister matings. The sequenced fish came from generation 104. A female fish was chosen because of its XX sex chromosome constitution. RNA that was sequenced to assemble the Jp163A reference transcriptome was isolated from two stages of pooled embryos (stages 15 and 25), a single 5-d-old individual and a 1-month-old fry, a single male and female at 2 months of age, one 9-month-old female, one 15-month-old male and the testes and ovaries of a single male and a single female 10-month-old fish.
A Jp163A BAC library (average insert size of 160 kb; 10× genome coverage with a total of 43,192 clones available)40 was produced from subline WLC#1247, maintained at the Biocenter Fish Facility (BFF) at the University of Würzburg (Würzburg, Germany). WLC#1247 was separated from the XGSC Jp163A line after approximately generation 50 and then maintained by inbreeding at BFF.
For RAD-tag mapping, one X. maculatus Jp163A male (WLC#1325, BFF) was crossed with an X. hellerii female (strain Rio Lancetilla, Db-, WLC#1337, BFF). Two F1 hybrid females from this cross were then backcrossed to X. hellerii males, and DNA from 267 backcross individuals was used for analysis.
Genome sequencing.
All genomic sequences for de novo assembly were generated on Roche 454 Titanium and Illumina Genome Analyzer IIx instruments, with the exception of the BAC-end sequences, which were generated on an ABI3730.
Physical map.
A physical map indicating tiling paths of X. maculatus contigs was constructed by generating fingerprints from the WLC-1247 BAC library (see URLs)40.
Genome assembly.
Two independent assemblies were built with all sequence data, using the Newbler (Roche) and PCAP41 algorithms from ∼19.6× total sequence coverage in whole-genome shotgun reads, a combination of 12× fragments, 9× 3-kb fragments, 0.38× 20-kb fragments and 0.02× BAC-end read pairs. A merged assembly was achieved by assigning the Newbler assembly as the reference and aligning the PCAP assembly via BLAT, followed by assimilation of all aligned scaffolds using an established graph accordance method42. Assembly consensus base error correction was accomplished by aligning Illumina reads (75-base paired-end reads, insert size of 200 bp), the same DNA source used for the reference, to the reference assembly using the Genomics Workbench v.4.03 software (CLC Bio). A consensus sequence was then created that factored the quality scores of both the reference assembly and the individual Illumina reads (Supplementary Fig. 8 and Supplementary Note). The annotated platyfish genome sequence is available at NCBI (AGAJ00000000).
Transcriptome sequencing and annotation.
Total RNA was isolated from platyfish tissues using the RiboPure Total RNA Isolation kit (Ambion). mRNA was isolated from total RNA using the Micro-PolyA Purist kit (Ambion). mRNA was reverse transcribed with SuperScript III Reverse Transcriptase (Invitrogen) using random hexamer primers (Invitrogen). Second-strand cDNA was synthesized using random primers and 15 U of Klenow DNA polymerase exo-minus (Epicentre). Double-stranded cDNA was sheared in a Bioruptor (Diagenode) for 30 cycles (30 sec on, 60 sec off). Sheared DNA was end repaired with the End-It DNA repair kit (Epicentre), and adenine overhangs were added with Klenow DNA polymerase exo-minus. cDNA was ligated to adapters overnight, and 100 ng was PCR amplified for 12 cycles with Phusion DNA polymerase (New England Biolabs). Each mRNA sample was sequenced on an Illumina Genome Analyzer IIx (60-bp reads). The X. maculatus transcriptome was assembled by combining sequences from several tissues, including heart, liver, brain, ovaries and testes, as well as from embryonic stages 15 and 25. For the X. hellerii transcriptome, RNA from 1-month-old whole fish and from the brain, liver, ovaries and testes of mature fishes was sequenced and assembled. Transcriptome sequences were aligned to the genome assembly contigs using Bowtie43, then assembled using the Velvet/Oases package (see URLs)44, reporting putative transcripts and splice variants using a coverage cutoff of 4, an insert length estimate of 120 bp and other parameters at default values.
Gene models and annotation.
Gene annotation using Ensembl genebuild was carried out on assembly Xipmac4.4.2 (GenBank Assembly GCA_000241075.1; see URLs).
Another gene identification analysis was performed by a combination of gene prediction and transcriptome integration. We used ab initio modeling with Augustus45 that had been trained on the medaka gene set and on the alignment of full-length gene models of medaka and zebrafish (both from Ensembl) using BLATX46. Transcriptome sequences were aligned to the assembly scaffolds using Bowtie43, the alignment was adjusted for the most likely exon-intron boundaries using TopHat47, and gene models were created using Cufflinks48. Only those transcripts containing a complete ORF and transcript read coverage of at least 3× were retained, and these were reconciled into a single set of 33,756 unique potential protein-encoding genes. These gene models were further culled to a subset of 17,783 that are amenable by phylogenetic analysis to entry into a whole-genome evolutionary interpretation using PHRINGE (Phylogenetic Resources for the Interpretation of Genomes) system (see URLs) by eliminating any transcripts shorter than 300 nucleotides and retaining only the longest version of any splice variant at each locus (Supplementary Fig. 9, Supplementary Tables 11 and 12 and Supplementary Note).
Estimation of gene number by transcriptome similarity.
We identified known genes by reciprocal BLASTX49 searches of the de novo transcriptome assembly against medaka, stickleback, fugu, tetraodon, zebrafish and human Ensembl gene libraries. To control for the inclusion of alternate transcript forms, we grouped these by the locus number as reported by Oases50.
Estimation of the number of novel genes.
To identify novel genes, we first reduced the redundancy of the platyfish transcriptome by clustering similar (with >95% identity) sequences. Sequences from clusters with no identifiable members were filtered to remove sequences that mapped (by GMAP51) with less than 99% identity to the genome or had predicted coding sequences shorter than 300 bp. Finally, identities for the remaining sequences were sought in the “nr” database (NCBI). Separate clustering by genomic distance (1 kb) produced a very similar gene number estimate (Supplementary Table 13 and Supplementary Note).
Annotation of noncoding RNAs.
To detect small nucleolar RNA (snoRNA), small nuclear RNA (snRNA), microRNA and rRNA, homology-based prediction was carried out using the multispecies RNA database (see URLs) comprised of zebrafish, stickleback, medaka and Takifugu noncoding RNA libraries. tRNAs were annotated using tRNAscan-SE.21 software locally on Linux52. rRNAs, microRNAs, snRNAs and snoRNAs were predicted by BLASTN using other fish noncoding RNA databases as queries, and duplicates were removed from the output files (Supplementary Tables 14 and 15). Fish databases were downloaded from Ensembl on the following genome versions: zv9 (Danio rerio), BROADS1 (Gasterosteus aculeatus), HdrR (O. latipes) and FUGU4.0 (Takifugu rubripes). microRNA sequences were identified with the Vienna RNA package of MiRscan (see URLs).
Annotation of TEs.
Both manual and automatic classification of TEs, on the basis of Wicker's nomenclature53, were performed, and identified elements were combined into a single library. Two TEs were considered to be different if their sequences diverged by more than 20% at the nucleotide level. Manual classification was carried out by searching TE sequence homology using CENSOR54 software, by homology searching specific TE proteins using TBLATN and BLASTP, by identifying terminal repeat features (TIRs, LTRs and TSDs) using BLASTN2 and LTR_FINDER software55, and by reconstructing phylogeny using ClustalW alignment and maximum-likelihood calculation (default aLRT) using the PhyML package38. Phylogenetic reconstructions for the DNA, long interspersed nucleotide element (LINE) and long terminal repeat (LTR) classes (Supplementary Figs. 1–3) were based either on comparisons of transposase or reverse transcriptase proteins. An automatic repeat library was built with RepeatScout software using default parameters on the supercontig assembly corrected for homopolymer errors. The percentage of TEs in the genome was determined from unassembled reads by locally running RepeatMasker software (see URLs) on the UNIX system.
Construction of a meiotic map using RAD tags.
Genomic DNA from map cross parents and progeny was digested with the restriction enzyme SbfI (New England Biolabs), and adapters with five-nucleotide barcodes each differing by at least two nucleotides were ligated onto fragments. RAD-tag libraries were made as described8. A 50-ng aliquot of size-selected DNA was PCR amplified for 12 cycles, and fragments 200 to 500 bp long were gel purified and sequenced using 80-nucleotide single-end reads on an Illumina HiSeq2000 sequencer. An equal amount of barcoded DNA from each of 16 progeny was loaded onto each lane. Low-quality reads and ambiguous barcodes were discarded. We used Stacks software56 to sort retained reads into loci and to genotype individuals by implementing the likelihood-based SNP calling algorithm57 to distinguish SNPs from sequencing errors. Stacks exported data into JoinMap 4.0 for linkage analysis using markers that were present in at least 200 of 267 individuals.
Assigning scaffolds to map positions.
To finalize assembly scaffold order and orientation, we used the high-density meiotic map to assign genome contigs to the genetic map. Using 14,391 marker sequences, we could reliably align 1,950 scaffolds to all linkage groups. Of these, 231 scaffolds contained blocks of markers from more than 1 linkage group, suggesting a misassembly event. In these cases, we manually split the scaffolds to maintain order with the genetic map (Supplementary Note).
Genome synteny.
For the analysis of conserved syntenies, the Synteny Database was employed using parameters as described10. In constructing the dot plots, for each gene along a specific platyfish chromosome, the Synteny Database identifies orthologs and paralogs by reciprocal best-BLAST analysis and plots positive results on the chromosomes of the same or other species directly above the index gene on the index chromosome.
Analyses of viviparity-related genes.
Thirty-four protein-coding genes known to function in yolk production, placenta-related characteristics and zona pellucida structures were selected as candidate genes (Supplementary Note) for the evolution of viviparity among Xiphophorus fishes. Eighteen randomly selected genes were used for control. Orthologous sequences for these genes from four fish species (O. latipes, G. aculeatus, Tetraodon nigroviridis and D. rerio) were retrieved from the Ensembl database and aligned using the MAFFT translation alignment. PAML (version 4.4, linux 64 bit) was implemented to test whether genes were under positive selection using a branch site–specific model (see URLs). Genes with P values less than 0.05 in likelihood ratio tests were designated as positively selected in Xiphophorus, and the Bayes empirical Bayes method58 was further used to calculate the selection pressure at each site.
Analysis of post-TGD gene retention.
The orthologs in human, mouse and teleosts of genes involved in cognition, pigmentation and liver function were obtained from Ensembl65, and missing gene annotations were identified with TBLASTN (Supplementary Table 7 and Supplementary Note). EnsemblCompara GeneTrees were checked for teleost duplications, and TGD-based duplications were confirmed using the Synteny Database10. Xiphophorus orthologs were identified from transcriptome v4 and the genome using BLAST searches, and assignment was confirmed with the Synteny Database. Potential bias in TGD-derived duplicate retention due to dosage sensitivity, protein complex membership and gene length was tested (Supplementary Note).
URLs.
Xiphophorus Genetic Stock Center (XGSC), http://www.xiphophorus.txstate.edu/; Platyfish BAC Library, http://bacpac.chori.org/library.php?id=353; Oases software package, http://www.ebi.ac.uk/~zerbino/oases/; PHRINGE resource, http://xiphophorus.genomeprojectsolutions-databases.com/; MiRscan tool, http://genes.mit.edu/mirscan/; RepeatMasker, http://repeatmasker.org/; Geneious software package, http://www.geneious.com/; platyfish transcriptome, http://avogadro.tr.txstate.edu/cgi-bin/gb2/gbrowse/XM_ncbi442/ and http://avogadro.tr.txstate.edu/Xiph_data_link/stable/Xm_transcriptome_v4.0/; platyfish gene models, http://xiphophorus.genomeprojectsolutions-databases.com/ and http://avogadro.tr.txstate.edu/Xiph_data_link/stable/Xm_JB_gene_models/; multispecies RNA database, http://www.ensembl.org/info/data/ftp/index.html; platyfish genome at Ensembl, http://www.ensembl.org/Xiphophorus_maculatus/Info/Index; GenBank assembly GCA_000241075.1, http://www.ncbi.nlm.nih.gov/genome/assembly/?term=GCA_000241075.1; genomic variants database, http://dgvbeta.tcag.ca/dgv/app/home; Human Protein Reference Database, http://www.hprd.org/.
Accession codes.
All sequence data have been deposited in the NCBI database under accession AGAJ00000000. All annotated sequences, genes, transcripts and proteins are available from http://www.ensembl.org/Xiphophorus_maculatus/Info/Index and http://xiphophorus.genomeprojectsolutions-databases.com/. Transcriptome data are deposited at http://avogadro.tr.txstate.edu/Xiph_data_link/stable/Xm_transcriptome_v4.0/.
Accession codes
References
Volff, J.N., Bouneau, L., Ozouf-Costaz, C. & Fischer, C. Diversity of retrotransposable elements in compact pufferfish genomes. Trends Genet. 19, 674–678 (2003).
Gifford, R. & Tristem, M. The evolution, distribution and diversity of endogenous retroviruses. Virus Genes 26, 291–315 (2003).
Katzourakis, A., Gifford, R.J., Tristem, M., Gilbert, M.T. & Pybus, O.G. Macroevolution of complex retroviruses. Science 325, 1512 (2009).
Han, G.Z. & Worobey, M. An endogenous foamy virus in the Aye-Aye (Daubentonia madagascariensis). J. Virol. 86, 7696–7698 (2012).
Han, G.Z. & Worobey, M. An endogenous foamy-like viral element in the coelacanth genome. PLoS Pathog. 8, e1002790 (2012).
Llorens, C., Munoz-Pomer, A., Bernad, L., Botella, H. & Moya, A. Network dynamics of eukaryotic LTR retroelements beyond phylogenetic trees. Biol. Direct 4, 41 (2009).
Ferguson-Smith, M.A. & Trifonov, V. Mammalian karyotype evolution. Nat. Rev. Genet. 8, 950–962 (2007).
Amores, A., Catchen, J., Ferrara, A., Fontenot, Q. & Postlethwait, J.H. Genome evolution and meiotic maps by massively parallel DNA sequencing: spotted gar, an outgroup for the teleost genome duplication. Genetics 188, 799–808 (2011).
Walter, R.B. et al. A microsatellite genetic linkage map for Xiphophorus. Genetics 168, 363–372 (2004).
Catchen, J.M., Conery, J.S. & Postlethwait, J.H. Automated identification of conserved synteny after whole-genome duplication. Genome Res. 19, 1497–1505 (2009).
Miya, M. et al. Major patterns of higher teleostean phylogenies: a new perspective based on 100 complete mitochondrial DNA sequences. Mol. Phylogenet. Evol. 26, 121–138 (2003).
Steinke, D., Salzburger, W. & Meyer, A. Novel relationships among ten fish model species revealed based on a phylogenomic analysis using ESTs. J. Mol. Evol. 62, 772–784 (2006).
Patton, E.E., Mitchell, D.L. & Nairn, R.S. Genetic and environmental melanoma models in fish. Pigment Cell and Melanoma Res. 23, 314–337 (2010).
Wittbrodt, J. et al. Novel putative receptor tyrosine kinase encoded by the melanoma-inducing Tu locus in Xiphophorus. Nature 341, 415–421 (1989).
Kallman, K.D. The platyfish Xiphophorus maculatus. in Handbook of Genetics Vol. 4 (ed. King, R.C.) 81–132 (Plenum Press, New York, 1975).
Gutbrod, H. & Schartl, M. Intragenic sex-chromosomal crossovers of Xmrk oncogene alleles affect pigment pattern formation and the severity of melanoma in Xiphophorus. Genetics 151, 773–783 (1999).
Meierjohann, S. & Schartl, M. From Mendelian to molecular genetics: the Xiphophorus melanoma model. Trends Genet. 22, 654–661 (2006).
Braasch, I., Brunet, F., Volff, J.N. & Schartl, M. Pigmentation pathway evolution after whole-genome duplication in fish. Genome Biol. Evol. 1, 479–493 (2009).
Laisney, J.A., Mueller, T.D., Schartl, M. & Meierjohann, S. Hyperactivation of constitutively dimerized oncogenic EGF receptors by autocrine loops. Oncogene published online; doi:10.1038/onc.2012.267 (2 July 2012).
Regneri, J. & Schartl, M. Expression regulation triggers oncogenicity of xmrk alleles in the Xiphophorus melanoma system. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 155, 71–80 (2012).
Kazianis, S. et al. Localization of a CDKN2 gene in linkage group V of Xiphophorus fishes defines it as a candidate for the DIFF tumor suppressor. Genes Chromosom. Cancer 22, 210–220 (1998).
Chatzinasiou, F. et al. Comprehensive field synopsis and systematic meta-analyses of genetic association studies in cutaneous melanoma. J. Natl. Cancer Inst. 103, 1227–1235 (2011).
Butler, A.P. et al. Regulation of CDKN2A/B and Retinoblastoma genes in Xiphophorus melanoma. Comp. Biochem. Physiol. C Toxicol. Pharmacol. 145, 145–155 (2007).
Walter, R.B. & Kazianis, S. Xiphophorus interspecies hybrids as genetic models of induced neoplasia. ILAR J. 42, 299–321 (2001).
Nairn, R.S. et al. Genetic analysis of susceptibility to spontaneous and UV-induced carcinogenesis in Xiphophorus hybrid fish. Mar. Biotechnol. (NY) 3, S24–S36 (2001).
Kazianis, S. et al. Genetic analysis of neoplasia induced by N-nitroso-N-methylurea in Xiphophorus hybrid fish. Mar. Biotechnol. (NY) 3, S37–S43 (2001).
Hrbek, T., Seckinger, J. & Meyer, A. A phylogenetic and biogeographic perspective on the evolution of poeciliid fishes. Mol. Phylogenet. Evol. 43, 986–998 (2007).
Pollux, B.J.A., Pires, M.N., Banet, A.I. & Reznick, D.N. Evolution of placentas in the fish family Poeciliidae: an empirical study of macroevolution. Annu. Rev. Ecol. Evol. Syst. 40, 271–289 (2009).
Turner, C.L. Pseudoamnion, pseudochorion, and follicular pseudoplacenta in poeciliid fishes. J. Morphol. 67, 59–87 (1940).
Tavolga, W.N. & Rugh, R. Development of the platyfish, Platypoecilius maculatus. Zoologica 32, 1–15 (1947).
Scrimshaw, N.S. Embryonic development in poeciliid fishes. Biol. Bull. 88, 233–246 (1945).
Brawand, D., Wahli, W. & Kaessmann, H. Loss of egg yolk genes in mammals and the origin of lactation and placentation. PLoS Biol. 6, e63 (2008).
Sibley, C.P. et al. Placental-specific insulin-like growth factor 2 (Igf2) regulates the diffusional exchange characteristics of the mouse placenta. Proc. Natl. Acad. Sci. USA 101, 8204–8208 (2004).
Vo, L.H. & Hedrick, J.L. Independent and hetero-oligomeric–dependent sperm binding to egg envelope glycoprotein ZPC in Xenopus laevis. Biol. Reprod. 62, 766–774 (2000).
Bshary, R., Wickler, W. & Fricke, H. Fish cognition: a primate's eye view. Anim. Cogn. 5, 1–13 (2002).
Force, A. et al. Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151, 1531–1545 (1999).
Braasch, I. & Postlethwait, J.H. Fish polyploidy and the teleost genome duplication. in Polyploidy and Genome Evolution (eds. Soltis, P.S. & Soltis, D.E.) 341–383 (Springer, Berlin, Heidelberg, 2012).
Galtier, N., Gouy, M. & Gautier, C. SEAVIEW and PHYLO_WIN: two graphic tools for sequence alignment and molecular phylogeny. Comput. Appl. Biosci. 12, 543–548 (1996).
Setiamarga, D.H. et al. Divergence time of the two regional medaka populations in Japan as a new time scale for comparative genomics of vertebrates. Biol. Lett. 5, 812–816 (2009).
Froschauer, A. et al. Construction and initial analysis of bacterial artificial chromosome (BAC) contigs from the sex-determining region of the platyfish Xiphophorus maculatus. Gene 295, 247–254 (2002).
Huang, X., Wang, J., Aluru, S., Yang, S.P. & Hillier, L. PCAP: a whole-genome assembly program. Genome Res. 13, 2164–2170 (2003).
Yao, G. et al. Graph accordance of next-generation sequence assemblies. Bioinformatics 28, 13–16 (2012).
Langmead, B., Trapnell, C., Pop, M. & Salzberg, S.L. Ultrafast and memory-efficient alignment of short DNA sequences to the human genome. Genome Biol. 10, R25 (2009).
Zerbino, D.R. & Birney, E. Velvet: algorithms for de novo short read assembly using de Bruijn graphs. Genome Res. 18, 821–829 (2008).
Stanke, M. & Waack, S. Gene prediction with a hidden Markov model and a new intron submodel. Bioinformatics 19 (suppl. 2), ii215–ii225 (2003).
Kent, W.J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Trapnell, C., Pachter, L. & Salzberg, S.L. TopHat: discovering splice junctions with RNA-Seq. Bioinformatics 25, 1105–1111 (2009).
Trapnell, C. et al. Transcript assembly and quantification by RNA-Seq reveals unannotated transcripts and isoform switching during cell differentiation. Nat. Biotechnol. 28, 511–515 (2010).
Camacho, C. et al. BLAST+: architecture and applications. BMC Bioinformatics 10, 421 (2009).
Schulz, M.H., Zerbino, D.R., Vingron, M. & Birney, E. Oases: robust de novo RNA-seq assembly across the dynamic range of expression levels. Bioinformatics 28, 1086–1092 (2012).
Wu, T.D. & Watanabe, C.K. GMAP: a genomic mapping and alignment program for mRNA and EST sequences. Bioinformatics 21, 1859–1875 (2005).
Lowe, T.M. & Eddy, S.R. tRNAscan-SE: a program for improved detection of transfer RNA genes in genomic sequence. Nucleic Acids Res. 25, 955–964 (1997).
Wicker, T. et al. A unified classification system for eukaryotic transposable elements. Nat. Rev. Genet. 8, 973–982 (2007).
Kohany, O., Gentles, A.J., Hankus, L. & Jurka, J. Annotation, submission and screening of repetitive elements in Repbase: RepbaseSubmitter and Censor. BMC Bioinformatics 7, 474 (2006).
Xu, Z. & Wang, H. LTR_FINDER: an efficient tool for the prediction of full-length LTR retrotransposons. Nucleic Acids Res. 35, W265–W268 (2007).
Catchen, J.M., Amores, A., Hohenlohe, P., Cresko, W. & Postlethwait, J.H. Stacks: building and genotyping loci de novo from short-read sequences. G3 (Bethesda) 1, 171–182 (2011).
Hohenlohe, P.A. et al. Population genomics of parallel adaptation in threespine stickleback using sequenced RAD tags. PLoS Genet. 6, e1000862 (2010).
Yang, Z., Wong, W.S. & Nielsen, R. Bayes empirical bayes inference of amino acid sites under positive selection. Mol. Biol. Evol. 22, 1107–1118 (2005).
Acknowledgements
The authors would like to thank the staff of the Xiphophorus Genetic Stock Center (XGSC) and the Biocenter Fish Facility for maintaining the pedigreed fish lines used in this study. We gratefully acknowledge the sequencing efforts of C. Fronick, K. Delehaunty and the production sequencing group at the Genome Institute. This work was supported by US National Institutes of Health, National Center for Research Resources (NCRR) and Office of Research Infrastructure Programs (ORIP), Division of Comparative Medicine grants R24 RR024790 and R24 OD011120 (R.B.W.), including an American Recovery and Reinvestment Act supplement to this award, and R24OD011199 (R.B.W.), R24 RR032658 and R24 OD011198 (W.C.W.), R01 RR020833 and R01 OD011116 (J.H.P.), by the Deutsche Forschungsgemeinschaft, TRR 58/A5 (K.P.L.) and TRR 17 (M.S.) VolkswagenStiftung, grant I/84 815 (I.B.) and by the Agence Nationale de Recherche (J.-N.V.).
Author information
Authors and Affiliations
Contributions
M.S., R.B.W., J.H.P. and W.C.W. are the principal investigators who conceived the project, analyzed data and wrote the manuscript. R.B.W. and M.S. provided the inbred Jp163A fish and RNA samples. W.C.W. carried out BAC and whole-genome sequencing, produced assembly, testing and submission. P.M. and L.H. built the assembly. R.K.W. coordinated genome sequencing and assembly. T.G., R.B.W. and Y.S. provided RNA sequencing and assembled a JP163A reference transcriptome. J.B. and S.F. improved the genome assembly consensus by incorporating Illumina sequencing reads, created gene models and performed whole-genome evolutionary analysis using PHRINGE. S.S. led the Ensembl gene annotation. J.C. wrote software for RAD-tag mapping and aligned the transcriptome and genomic contigs to the genetic map. J.H.P. led the construction of the genetic map and performed the conserved synteny analyses. M.S. performed the mapping cross. A.A. constructed the genetic map. D.C., T.G. and J.-N.V. performed repeat analysis and noncoding RNA and TE annotation. Y.S. analyzed the viviparity genes. A.B. developed the TGD-cognition hypothesis. I.B. designed Figure 1b, analyzed TGD paralog retention and pigmentation gene locations and contributed to manuscript writing. I.B., K.-P.L. and M.S. analyzed cognition-related genes.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Note, Supplementary Tables 1–15 and Supplementary Figures 1–9 (PDF 4856 kb)
Rights and permissions
This work is licensed under a Creative Commons Attribution-NonCommercial-Share Alike 3.0 Unported License. To view a copy of this license, visit http://creativecommons.org/licenses/by-nc-sa/3.0/.
About this article
Cite this article
Schartl, M., Walter, R., Shen, Y. et al. The genome of the platyfish, Xiphophorus maculatus, provides insights into evolutionary adaptation and several complex traits. Nat Genet 45, 567–572 (2013). https://doi.org/10.1038/ng.2604
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2604
This article is cited by
-
A lethal mitonuclear incompatibility in complex I of natural hybrids
Nature (2024)
-
The medaka mutant deficient in eyes shut homolog exhibits opsin transport defects and enhanced autophagy in retinal photoreceptors
Cell and Tissue Research (2023)
-
Fish genomics and its impact on fundamental and applied research of vertebrate biology
Reviews in Fish Biology and Fisheries (2022)
-
Zebrafish excel in number discrimination under an operant conditioning paradigm
Animal Cognition (2022)
-
Whole-genome sequencing reveals sex determination and liver high-fat storage mechanisms of yellowstripe goby (Mugilogobius chulae)
Communications Biology (2021)