Abstract
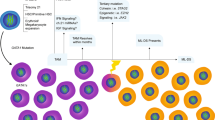
Transient abnormal myelopoiesis (TAM) is a myeloid proliferation resembling acute megakaryoblastic leukemia (AMKL), mostly affecting perinatal infants with Down syndrome. Although self-limiting in a majority of cases, TAM may evolve as non-self-limiting AMKL after spontaneous remission (DS-AMKL). Pathogenesis of these Down syndrome–related myeloid disorders is poorly understood, except for GATA1 mutations found in most cases. Here we report genomic profiling of 41 TAM, 49 DS-AMKL and 19 non-DS-AMKL samples, including whole-genome and/or whole-exome sequencing of 15 TAM and 14 DS-AMKL samples. TAM appears to be caused by a single GATA1 mutation and constitutive trisomy 21. Subsequent AMKL evolves from a pre-existing TAM clone through the acquisition of additional mutations, with major mutational targets including multiple cohesin components (53%), CTCF (20%), and EZH2, KANSL1 and other epigenetic regulators (45%), as well as common signaling pathways, such as the JAK family kinases, MPL, SH2B3 (LNK) and multiple RAS pathway genes (47%).
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout





Similar content being viewed by others
Accession codes
Accessions
NCBI Reference Sequence
Change history
30 October 2013
In the version of this article initially published, the discussion of cited reference 52 should also have noted that the work "reported accumulation of additional somatic mutations (including single cases of SMC3 and EZH2 mutation) during progression from TAM to DS-AMKL." The error has been corrected in the HTML and PDF versions of the article.
References
Khan, I., Malinge, S. & Crispino, J. Myeloid leukemia in Down syndrome. Crit. Rev. Oncog. 16, 25–36 (2011).
Massey, G.V. et al. A prospective study of the natural history of transient leukemia (TL) in neonates with Down syndrome (DS): Children's Oncology Group (COG) study POG-9481. Blood 107, 4606–4613 (2006).
Muramatsu, H. et al. Risk factors for early death in neonates with Down syndrome and transient leukaemia. Br. J. Haematol. 142, 610–615 (2008).
Klusmann, J.H. et al. Treatment and prognostic impact of transient leukemia in neonates with Down syndrome. Blood 111, 2991–2998 (2008).
Xu, G. et al. Frequent mutations in the GATA-1 gene in the transient myeloproliferative disorder of Down syndrome. Blood 102, 2960–2968 (2003).
Wechsler, J. et al. Acquired mutations in GATA1 in the megakaryoblastic leukemia of Down syndrome. Nat. Genet. 32, 148–152 (2002).
Walters, D.K. et al. Activating alleles of JAK3 in acute megakaryoblastic leukemia. Cancer Cell 10, 65–75 (2006).
Malinge, S. et al. Activating mutations in human acute megakaryoblastic leukemia. Blood 112, 4220–4226 (2008).
Blink, M. et al. Frequency and prognostic implications of JAK 1–3 aberrations in Down syndrome acute lymphoblastic and myeloid leukemia. Leukemia 25, 1365–1368 (2011).
Hama, A. et al. Molecular lesions in childhood and adult acute megakaryoblastic leukaemia. Br. J. Haematol. 156, 316–325 (2012).
Malkin, D., Brown, E.J. & Zipursky, A. The role of p53 in megakaryocyte differentiation and the megakaryocytic leukemias of Down syndrome. Cancer Genet. Cytogenet. 116, 1–5 (2000).
Hussein, K. et al. MPLW515L mutation in acute megakaryoblastic leukaemia. Leukemia 23, 852–855 (2009).
Greenman, C. et al. Patterns of somatic mutation in human cancer genomes. Nature 446, 153–158 (2007).
Welch, J.S. et al. The origin and evolution of mutations in acute myeloid leukemia. Cell 150, 264–278 (2012).
Ding, L. et al. Clonal evolution in relapsed acute myeloid leukaemia revealed by whole-genome sequencing. Nature 481, 506–510 (2012).
Creutzig, U. et al. Diagnosis and management of acute myeloid leukemia in children and adolescents: recommendations from an international expert panel. Blood 120, 3187–3205 (2012).
Swerdlow, S.H., Jaffe, E.S. & International Agency for Research on Cancer & World Health Organization WHO Classification of Tumours of Haematopoietic and Lymphoid Tissues (International Agency for Research on Cancer, Lyon, France, 2008).
Wu, C. et al. BioGPS: an extensible and customizable portal for querying and organizing gene annotation resources. Genome Biol. 10, R130 (2009).
Bourquin, J.P. et al. Identification of distinct molecular phenotypes in acute megakaryoblastic leukemia by gene expression profiling. Proc. Natl. Acad. Sci. USA 103, 3339–3344 (2006).
Mercher, T. et al. Involvement of a human gene related to the Drosophila spen gene in the recurrent t(1;22) translocation of acute megakaryocytic leukemia. Proc. Natl. Acad. Sci. USA 98, 5776–5779 (2001).
Ma, Z. et al. Fusion of two novel genes, RBM15 and MKL1, in the t(1;22)(p13;q13) of acute megakaryoblastic leukemia. Nat. Genet. 28, 220–221 (2001).
Gruber, T.A. et al. An inv(16)(p13.3q24.3)-encoded CBFA2T3-GLIS2 fusion protein defines an aggressive subtype of pediatric acute megakaryoblastic leukemia. Cancer Cell 22, 683–697 (2012).
Thiollier, C. et al. Characterization of novel genomic alterations and therapeutic approaches using acute megakaryoblastic leukemia xenograft models. J. Exp. Med. 209, 2017–2031 (2012).
Gruber, S., Haering, C.H. & Nasmyth, K. Chromosomal cohesin forms a ring. Cell 112, 765–777 (2003).
Nasmyth, K. & Haering, C.H. Cohesin: its roles and mechanisms. Annu. Rev. Genet. 43, 525–558 (2009).
Wendt, K.S. et al. Cohesin mediates transcriptional insulation by CCCTC-binding factor. Nature 451, 796–801 (2008).
Ström, L. et al. Postreplicative formation of cohesion is required for repair and induced by a single DNA break. Science 317, 242–245 (2007).
Watrin, E. & Peters, J.M. The cohesin complex is required for the DNA damage–induced G2/M checkpoint in mammalian cells. EMBO J. 28, 2625–2635 (2009).
Dorsett, D. et al. Effects of sister chromatid cohesion proteins on cut gene expression during wing development in Drosophila. Development 132, 4743–4753 (2005).
Parelho, V. et al. Cohesins functionally associate with CTCF on mammalian chromosome arms. Cell 132, 422–433 (2008).
Solomon, D.A. et al. Mutational inactivation of STAG2 causes aneuploidy in human cancer. Science 333, 1039–1043 (2011).
Forestier, E. et al. Cytogenetic features of acute lymphoblastic and myeloid leukemias in pediatric patients with Down syndrome: an iBFM-SG study. Blood 111, 1575–1583 (2008).
Rubio, E.D. et al. CTCF physically links cohesin to chromatin. Proc. Natl. Acad. Sci. USA 105, 8309–8314 (2008).
Stedman, W. et al. Cohesins localize with CTCF at the KSHV latency control region and at cellular c-myc and H19/Igf2 insulators. EMBO J. 27, 654–666 (2008).
Ohlsson, R., Bartkuhn, M. & Renkawitz, R. CTCF shapes chromatin by multiple mechanisms: the impact of 20 years of CTCF research on understanding the workings of chromatin. Chromosoma 119, 351–360 (2010).
Phillips, J.E. & Corces, V.G. CTCF: master weaver of the genome. Cell 137, 1194–1211 (2009).
Wendt, K.S. & Peters, J.M. How cohesin and CTCF cooperate in regulating gene expression. Chromosome Res. 17, 201–214 (2009).
Cancer Genome Atlas Network. Comprehensive molecular portraits of human breast tumours. Nature 490, 61–70 (2012).
Cao, R. et al. Role of histone H3 lysine 27 methylation in Polycomb-group silencing. Science 298, 1039–1043 (2002).
Ernst, T. et al. Inactivating mutations of the histone methyltransferase gene EZH2 in myeloid disorders. Nat. Genet. 42, 722–726 (2010).
Patel, J.P. et al. Prognostic relevance of integrated genetic profiling in acute myeloid leukemia. N. Engl. J. Med. 366, 1079–1089 (2012).
Koolen, D.A. et al. Mutations in the chromatin modifier gene KANSL1 cause the 17q21.31 microdeletion syndrome. Nat. Genet. 44, 639–641 (2012).
Zollino, M. et al. Mutations in KANSL1 cause the 17q21.31 microdeletion syndrome phenotype. Nat. Genet. 44, 636–638 (2012).
Yang, X.J. The diverse superfamily of lysine acetyltransferases and their roles in leukemia and other diseases. Nucleic Acids Res. 32, 959–976 (2004).
Li, X., Wu, L., Corsa, C.A., Kunkel, S. & Dou, Y. Two mammalian MOF complexes regulate transcription activation by distinct mechanisms. Mol. Cell 36, 290–301 (2009).
Bercovich, D. et al. Mutations of JAK2 in acute lymphoblastic leukaemias associated with Down's syndrome. Lancet 372, 1484–1492 (2008).
Mullighan, C.G. et al. JAK mutations in high-risk childhood acute lymphoblastic leukemia. Proc. Natl. Acad. Sci. USA 106, 9414–9418 (2009).
Kratz, C.P. et al. Mutational screen reveals a novel JAK2 mutation, L611S, in a child with acute lymphoblastic leukemia. Leukemia 20, 381–383 (2006).
Nussenzveig, R.H. et al. Detection of JAK2 mutations in paraffin marrow biopsies by high resolution melting analysis: identification of L611S alone and in cis with V617F in polycythemia vera. Leuk. Lymphoma 53, 2479–2486 (2012).
Miyata, Y. & Nishida, E. DYRK1A binds to an evolutionarily conserved WD40-repeat protein WDR68 and induces its nuclear translocation. Biochim. Biophys. Acta 1813, 1728–1739 (2011).
de Rooij, J.D. et al. NUP98/JARID1A is a novel recurrent abnormality in pediatric acute megakaryoblastic leukemia with a distinct HOX gene expression pattern. Leukemia doi:10.1038/leu.2013.87 (27 March 2013).10.1038/leu.2013.87
Nikolaev, S.I. et al. Exome sequencing identifies putative drivers of progression of transient myeloproliferative disorder to AMKL in infants with Down Syndrome. Blood 122, 554–561 (2013).
Krzywinski, M. et al. Circos: an information aesthetic for comparative genomics. Genome Res. 19, 1639–1645 (2009).
Sato, Y. et al. Integrated molecular analysis of clear-cell renal cell carcinoma. Nat. Genet. 45, 860–867 (2013).
Yoshida, K. et al. Frequent pathway mutations of splicing machinery in myelodysplasia. Nature 478, 64–69 (2011).
Kent, W.J. BLAT—the BLAST-like alignment tool. Genome Res. 12, 656–664 (2002).
Shiraishi, Y. et al. An empirical Bayesian framework for somatic mutation detection from cancer genome sequencing data. Nucleic Acids Res. 41, e89 (2013).
Sakaguchi, H. et al. Exome sequencing identifies secondary mutations of SETBP1 and JAK3 in juvenile myelomonocytic leukemia. Nat. Genet. 45, 937–941 (2013).
Li, H. et al. The Sequence Alignment/Map format and SAMtools. Bioinformatics 25, 2078–2079 (2009).
Wang, K., Li, M. & Hakonarson, H. ANNOVAR: functional annotation of genetic variants from high-throughput sequencing data. Nucleic Acids Res. 38, e164 (2010).
Forbes, S.A. et al. COSMIC: mining complete cancer genomes in the Catalogue of Somatic Mutations in Cancer. Nucleic Acids Res. 39, D945–D950 (2011).
Robinson, J.T. et al. Integrative genomics viewer. Nat. Biotechnol. 29, 24–26 (2011).
Torres, L. et al. Acute megakaryoblastic leukemia with a four-way variant translocation originating the RBM15-MKL1 fusion gene. Pediatr. Blood Cancer 56, 846–849 (2011).
Nannya, Y. et al. A robust algorithm for copy number detection using high-density oligonucleotide single nucleotide polymorphism genotyping arrays. Cancer Res. 65, 6071–6079 (2005).
Yamamoto, G. et al. Highly sensitive method for genomewide detection of allelic composition in nonpaired, primary tumor specimens by use of Affymetrix single-nucleotide-polymorphism genotyping microarrays. Am. J. Hum. Genet. 81, 114–126 (2007).
Acknowledgements
We thank Y. Mori, M. Nakamura, O. Hagiwara and N. Mizota for their technical assistance. This work was supported by the Research on Measures for Intractable Diseases Project and Health and Labor Sciences Research grants (Research on Intractable Diseases) from the Ministry of Health, Labour and Welfare, by Grants-in-Aid from the Ministry of Health, Labor and Welfare of Japan and KAKENHI (22134006, 23249052, 23118501, 23390266 and 25461579) and by the Japan Society for the Promotion of Science (JSPS) through the Funding Program for World-Leading Innovative Research and Development on Science and Technology (FIRST Program), initiated by the Council for Science and Technology Policy (CSTP) and research grants from the Japan Science and Technology Agency CREST.
Author information
Authors and Affiliations
Contributions
Y.O., Y. Shiraishi, A.S.-O., K.C., H.T. and S.M. performed bioinformatics analyses of the resequencing data. M.S., A.S.-O., Y. Sato, A.H. and H.M. performed microarray experiments and analyses. R.K. and A.H. performed RT-PCR analyses. M.P., K. Terui, R.W., D.H., K.N., H.K., K. Tsukamoto, S.A., K. Kawakami, K. Kato, R.N., S.I., Y.H., S.K. and E.I. collected specimens and were involved in planning the project. K.Y., T.T., H.S., Y.N. and N.S. processed and analyzed genetic materials, prepared the library and performed sequencing. K.Y., T.T., Y.O., A.K. and S.O. generated figures and tables. E.I. and S.O. led the entire project. K.Y. and S.O. wrote the manuscript. All authors participated in discussions and interpretation of the data and results.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–14, Supplementary Tables 1–14 and Supplementary Note (PDF 2603 kb)
Rights and permissions
About this article
Cite this article
Yoshida, K., Toki, T., Okuno, Y. et al. The landscape of somatic mutations in Down syndrome–related myeloid disorders. Nat Genet 45, 1293–1299 (2013). https://doi.org/10.1038/ng.2759
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.2759
This article is cited by
-
The abundance of the short GATA1 isoform affects megakaryocyte differentiation and leukemic predisposition in mice
Experimental Hematology & Oncology (2024)
-
Inherent genome instability underlies trisomy 21-associated myeloid malignancies
Leukemia (2024)
-
Prenatal diagnosis of Down syndrome combined with transient abnormal myelopoiesis in foetuses with a GATA1 gene variant: two case reports
Molecular Cytogenetics (2023)
-
Patients with Down syndrome can be included in early phase clinical trials- a report from the T2016-003 Therapeutic Advances in Childhood Leukemia and Lymphoma (TACL) study
Leukemia (2023)
-
Long-range gene regulation in hormone-dependent cancer
Nature Reviews Cancer (2023)