Abstract
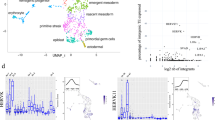
Detection of new genomic control elements is critical in understanding transcriptional regulatory networks in their entirety. We studied the genome-wide binding locations of three key regulatory proteins (POU5F1, also known as OCT4; NANOG; and CTCF) in human and mouse embryonic stem cells. In contrast to CTCF, we found that the binding profiles of OCT4 and NANOG are markedly different, with only ∼5% of the regions being homologously occupied. We show that transposable elements contributed up to 25% of the bound sites in humans and mice and have wired new genes into the core regulatory network of embryonic stem cells. These data indicate that species-specific transposable elements have substantially altered the transcriptional circuitry of pluripotent stem cells.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
Accession codes
Change history
13 June 2010
In the version of this article initially published online, the phrase “the genes encoding” was erroneously inserted into the second sentence of the abstract and the protein name of POU5F1 was incorrect. The error has been corrected for the print, PDF and HTML versions of this article.
References
Dermitzakis, E.T. & Clark, A.G. Evolution of transcription factor binding sites in Mammalian gene regulatory regions: conservation and turnover. Mol. Biol. Evol. 19, 1114–1121 (2002).
Borneman, A.R. et al. Divergence of transcription factor binding sites across related yeast species. Science 317, 815–819 (2007).
Moses, A.M. et al. Large-scale turnover of functional transcription factor binding sites in Drosophila. PLOS Comput. Biol. 2, e130 (2006).
Bourque, G. et al. Evolution of the mammalian transcription factor binding repertoire via transposable elements. Genome Res. 18, 1752–1762 (2008).
Birney, E. et al. Identification and analysis of functional elements in 1% of the human genome by the ENCODE pilot project. Nature 447, 799–816 (2007).
Odom, D.T. et al. Tissue-specific transcriptional regulation has diverged significantly between human and mouse. Nat. Genet. 39, 730–732 (2007).
Loh, Y.H. et al. The Oct4 and Nanog transcription network regulates pluripotency in mouse embryonic stem cells. Nat. Genet. 38, 431–440 (2006).
Boyer, L.A. et al. Core transcriptional regulatory circuitry in human embryonic stem cells. Cell 122, 947–956 (2005).
Bell, A.C., West, A.G. & Felsenfeld, G. The protein CTCF is required for the enhancer blocking activity of vertebrate insulators. Cell 98, 387–396 (1999).
Brons, I.G. et al. Derivation of pluripotent epiblast stem cells from mammalian embryos. Nature 448, 191–195 (2007).
Tesar, P.J. et al. New cell lines from mouse epiblast share defining features with human embryonic stem cells. Nature 448, 196–199 (2007).
Rossant, J. Stem cells and early lineage development. Cell 132, 527–531 (2008).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Wang, T. et al. Species-specific endogenous retroviruses shape the transcriptional network of the human tumor suppressor protein p53. Proc. Natl. Acad. Sci. USA 104, 18613–18618 (2007).
Cohen, C.J., Lock, W.M. & Mager, D.L. Endogenous retroviral LTRs as promoters for human genes: a critical assessment. Gene 448, 105–114 (2009).
Feschotte, C. Transposable elements and the evolution of regulatory networks. Nat. Rev. Genet. 9, 397–405 (2008).
Cao, R. & Zhang, Y. SUZ12 is required for both the histone methyltransferase activity and the silencing function of the EED-EZH2 complex. Mol. Cell 15, 57–67 (2004).
Sperger, J.M. et al. Gene expression patterns in human embryonic stem cells and human pluripotent germ cell tumors. Proc. Natl. Acad. Sci. USA 100, 13350–13355 (2003).
Kim, T.H. et al. Analysis of the vertebrate insulator protein CTCF-binding sites in the human genome. Cell 128, 1231–1245 (2007).
Davidson, E.H. & Britten, R.J. Regulation of gene expression: possible role of repetitive sequences. Science 204, 1052–1059 (1979).
McClintock, B. The significance of responses of the genome to challenge. Science 226, 792–801 (1984).
Brosius, J. Retroposons—seeds of evolution. Science 251, 753 (1991).
Thomson, J.A. et al. Embryonic stem cell lines derived from human blastocysts. Science 282, 1145–1147 (1998).
Xu, C. et al. Feeder-free growth of undifferentiated human embryonic stem cells. Nat. Biotechnol. 19, 971–974 (2001).
Zhang, Y. et al. Model-based analysis of ChIP-Seq (MACS). Genome Biol. 9, R137 (2008).
Conlon, E.M., Liu, X.S., Lieb, J.D. & Liu, J.S. Integrating regulatory motif discovery and genome-wide expression analysis. Proc. Natl. Acad. Sci. USA 100, 3339–3344 (2003).
Kent, W.J. et al. The human genome browser at UCSC. Genome Res. 12, 996–1006 (2002).
Siepel, A. et al. Evolutionarily conserved elements in vertebrate, insect, worm, and yeast genomes. Genome Res. 15, 1034–1050 (2005).
Tusher, V.G., Tibshirani, R. & Chu, G. Significance analysis of microarrays applied to the ionizing radiation response. Proc. Natl. Acad. Sci. USA 98, 5116–5121 (2001).
Acknowledgements
This work was supported by the Agency for Science, Technology and Research (A*STAR) of Singapore.
Author information
Authors and Affiliations
Contributions
H.-H.N. and G.B. designed the experiments. N.-Y.C., X.L. and Y.-S.C. performed the experiments. G.K. performed the data analysis with contributions from J.J. and C.H. G.B. wrote the manuscript with contributions from H.-H.N. and G.K.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Figures 1–7, Supplementary Tables 1–10 and Supplementary Note. (PDF 723 kb)
Supplementary Table 1
Human RABS for OCT4, NANOG and CTCF (XLS 59 kb)
Supplementary Table 2
Mouse RABS for Oct4, Nanog and Ctcf (XLS 23 kb)
Supplementary Table 3
Human and mouse POU5F1 and Pou5f1 RNAi results (XLS 492 kb)
Supplementary Table 5
OCT4 binding regions around the conserved and the human-specific OCT4 target genes (XLS 158 kb)
Supplementary Table 6
OCT4 binding regions (XLS 3530 kb)
Supplementary Table 7
NANOG binding regions (ZIP 1994 kb)
Supplementary Table 8
CTCF binding regions (ZIP 2003 kb)
Rights and permissions
About this article
Cite this article
Kunarso, G., Chia, NY., Jeyakani, J. et al. Transposable elements have rewired the core regulatory network of human embryonic stem cells. Nat Genet 42, 631–634 (2010). https://doi.org/10.1038/ng.600
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ng.600
This article is cited by
-
Activation of human endogenous retroviruses and its physiological consequences
Nature Reviews Molecular Cell Biology (2024)
-
A SINE-VNTR-Alu at the LRIG2 locus is associated with proximal and distal gene expression in CRISPR and population models
Scientific Reports (2024)
-
Selective binding of retrotransposons by ZFP352 facilitates the timely dissolution of totipotency network
Nature Communications (2023)
-
Widespread contribution of transposable elements to the rewiring of mammalian 3D genomes
Nature Communications (2023)
-
Comparative analysis of bats and rodents’ genomes suggests a relation between non-LTR retrotransposons, cancer incidence, and ageing
Scientific Reports (2023)