Abstract
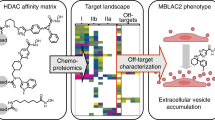
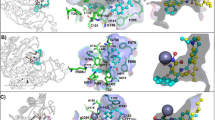
In contrast to studies on class I histone deacetylase (HDAC) inhibitors, the elucidation of the molecular mechanisms and therapeutic potential of class IIa HDACs (HDAC4, HDAC5, HDAC7 and HDAC9) is impaired by the lack of potent and selective chemical probes. Here we report the discovery of inhibitors that fill this void with an unprecedented metal-binding group, trifluoromethyloxadiazole (TFMO), which circumvents the selectivity and pharmacologic liabilities of hydroxamates. We confirm direct metal binding of the TFMO through crystallographic approaches and use chemoproteomics to demonstrate the superior selectivity of the TFMO series relative to a hydroxamate-substituted analog. We further apply these tool compounds to reveal gene regulation dependent on the catalytic active site of class IIa HDACs. The discovery of these inhibitors challenges the design process for targeting metalloenzymes through a chelating metal-binding group and suggests therapeutic potential for class IIa HDAC enzyme blockers distinct in mechanism and application compared to current HDAC inhibitors.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Bradner, J.E. et al. Chemical phylogenetics of histone deacetylases. Nat. Chem. Biol. 6, 238–243 (2010).
Fischle, W. et al. Human HDAC7 histone deacetylase activity is associated with HDAC3 in vivo. J. Biol. Chem. 276, 35826–35835 (2001).
Choudhary, C. et al. Lysine acetylation targets protein complexes and co-regulates major cellular functions. Science 325, 834–840 (2009).
Kim, S.C. et al. Substrate and functional diversity of lysine acetylation revealed by a proteomics survey. Mol. Cell 23, 607–618 (2006).
Yuan, Z., Peng, L., Radhakrishnan, R. & Seto, E. Histone deacetylase 9 (HDAC9) regulates the functions of the ATDC (TRIM29) protein. J. Biol. Chem. 285, 39329–39338 (2010).
Fischle, W. et al. Enzymatic activity associated with class II HDACs is dependent on a multiprotein complex containing HDAC3 and SMRT/N-CoR. Mol. Cell 9, 45–57 (2002).
McKinsey, T.A., Zhang, C.L. & Olson, E.N. Activation of the myocyte enhancer factor-2 transcription factor by calcium/calmodulin-dependent protein kinase-stimulated binding of 14–3-3 to histone deacetylase 5. Proc. Natl. Acad. Sci. USA 97, 14400–14405 (2000).
Zhang, C.L. et al. Class II histone deacetylases act as signal-responsive repressors of cardiac hypertrophy. Cell 110, 479–488 (2002).
Verdin, E., Dequiedt, F. & Kasler, H.G. Class II histone deacetylases: versatile regulators. Trends Genet. 19, 286–293 (2003).
Arrowsmith, C.H., Bountra, C., Fish, P.V., Lee, K. & Schapira, M. Epigenetic protein families: a new frontier for drug discovery. Nat. Rev. Drug Discov. 11, 384–400 (2012).
Brockschmidt, F.F. et al. Susceptibility variants on chromosome 7p21.1 suggest HDAC9 as a new candidate gene for male-pattern baldness. Br. J. Dermatol. 165, 1293–1302 (2011).
Mielcarek, M. et al. SAHA decreases HDAC 2 and 4 levels in vivo and improves molecular phenotypes in the R6/2 mouse model of Huntington's disease. PLoS One 6, e27746 (2011).
Quinti, L. et al. Evaluation of histone deacetylases as drug targets in Huntington's disease models. Study of HDACs in brain tissues from R6/2 and CAG140 knock-in HD mouse models and human patients and in a neuronal HD cell model. PLoS Curr. 2, pii:RRN1172 (2010).
Berdeaux, R. et al. SIK1 is a class II HDAC kinase that promotes survival of skeletal myocytes. Nat. Med. 13, 597–603 (2007).
Tao, R. et al. Deacetylase inhibition promotes the generation and function of regulatory T cells. Nat. Med. 13, 1299–1307 (2007).
International Stroke Genetics Consortium (ISGC). et al. Genome-wide association study identifies a variant in HDAC9 associated with large vessel ischemic stroke. Nat. Genet. 44 , 328–333 (2012).
DasGupta, S., Murumkar, P.R., Giridhar, R. & Yadav, M.R. Current perspective of TACE inhibitors: a review. Bioorg. Med. Chem. 17, 444–459 (2009).
Nuti, E., Tuccinardi, T. & Rossello, A. Matrix metalloproteinase inhibitors: new challenges in the era of post broad-spectrum inhibitors. Curr. Pharm. Des. 13, 2087–2100 (2007).
Lahm, A. et al. Unraveling the hidden catalytic activity of vertebrate class IIa histone deacetylases. Proc. Natl. Acad. Sci, USA 104, 17335–17340 (2007).
Baloglu, E., Ghosh, S., Lobera, M. & Schmidt,, D.R. Compounds and Methods. US Provisional Patent Application PCT/US2011/021089 (2011).
Ma, X., Ezzeldin, H.H. & Diasio, R.B. Histone deacetylase inhibitors: current status and overview of recent clinical trials. Drugs 69, 1911–1934 (2009).
Dormán, G. et al. Matrix metalloproteinase inhibitors: a critical appraisal of design principles and proposed therapeutic utility. Drugs 70, 949–964 (2010).
Schuetz, A. et al. Human HDAC7 harbors a class IIa histone deacetylase–specific zinc binding motif and cryptic deacetylase activity. J. Biol. Chem. 283, 11355–11363 (2008).
Bottomley, M.J. et al. Structural and functional analysis of the human HDAC4 catalytic domain reveals a regulatory structural zinc-binding domain. J. Biol. Chem. 283, 26694–26704 (2008).
Allen, F.H. The Cambridge Structural Database: a quarter of a million crystal structures and rising. Acta Crystallogr. B 58, 380–388 (2002).
Bantscheff, M. et al. Chemoproteomics profiling of HDAC inhibitors reveals selective targeting of HDAC complexes. Nat. Biotechnol. 29, 255–265 (2011).
Bertos, N.R., Wang, A.H. & Yang, X.J. Class II histone deacetylases: structure, function, and regulation. Biochem. Cell Biol. 79, 243–252 (2001).
Emanuele, S., Lauricella, M. & Tesoriere, G. Histone deacetylase inhibitors: apoptotic effects and clinical implications (Review). Int. J. Oncol. 33, 637–646 (2008).
Fleetwood, A.J., Lawrence, T., Hamilton, J.A. & Cook, A.D. Granulocyte-macrophage colony-stimulating factor (CSF) and macrophage CSF–dependent macrophage phenotypes display differences in cytokine profiles and transcription factor activities: implications for CSF blockade in inflammation. J. Immunol. 178, 5245–5252 (2007).
Lacey, D.C. et al. Defining GM-CSF– and macrophage-CSF–dependent macrophage responses by in vitro models. J. Immunol. 188, 5752–5765 (2012).
Parra, M. & Verdin, E. Regulatory signal transduction pathways for class IIa histone deacetylases. Curr. Opin. Pharmacol. 10, 454–460 (2010).
Spiegel, S., Milstien, S. & Grant, S. Endogenous modulators and pharmacological inhibitors of histone deacetylases in cancer therapy. Oncogene 31, 537–551 (2012).
Kewitz, S., Bernig, T. & Staege, M.S. Histone deacetylase inhibition restores cisplatin sensitivity of Hodgkin's lymphoma cells. Leuk. Res. 36, 773–778 (2012).
Acknowledgements
The authors would like to thank A.Y. Rudensky, D. Mathis, C. Benoist and V.K. Kuchroo for their comments during manuscript preparation. We also thank M.S. Sundrud and J.A. Hill for discussions and insight through the execution of this work. We also thank Y. Alekseyev and A. LeClerc of the Boston University Medical Campus Microarray Core facility.
Author information
Authors and Affiliations
Contributions
M.L. and M.A.N. performed experiments, directed the research, and wrote the manuscript. D.S. and S.G. contributed to experimental design. D.R.S., E.B., C. Wells and R.P.T. contributed to chemical synthesis. Q.G.W., M.T., P.K.M. and L.K. contributed to the biological experiments. K.P.M. and D.T.P. performed the crystallographic and computational efforts, and X.H., R.A.R., W.B. and M.S.H. contributed to the crystallographic efforts. G.A.H., M.M.-T., B.S., S.C. and Z.W. contributed to the high-throughput screening efforts. R.P.T. designed and B.J.T., J.X.R., C. Wagner, M.B.M., J.T.M. and J.D.W. contributed to the chemoproteomics efforts.
Corresponding author
Ethics declarations
Competing interests
All authors are present or former employees of Tempero Pharmaceuticals, Inc. or GlaxoSmithKline, as indicated in the affiliations.
Supplementary information
Supplementary Text and Figures
Supplementary Results (PDF 6031 kb)
Supplementary Data Set 1
Supplementary Dataset 1 (PDF 102 kb)
Supplementary Data Set 2
Supplementary Dataset 2 (PDF 1671 kb)
Supplementary Data Set 3
Supplementary Dataset 3 (PDF 81 kb)
Supplementary Data Set 4
Supplementary Dataset 4 (PDF 86 kb)
Supplementary Data Set 5
Supplementary Dataset 5 (PDF 259 kb)
Supplementary Note
Supplementary Note (PDF 283 kb)
Rights and permissions
About this article
Cite this article
Lobera, M., Madauss, K., Pohlhaus, D. et al. Selective class IIa histone deacetylase inhibition via a nonchelating zinc-binding group. Nat Chem Biol 9, 319–325 (2013). https://doi.org/10.1038/nchembio.1223
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1223
This article is cited by
-
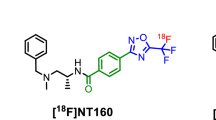
Design and radiosynthesis of class-IIa HDAC inhibitor with high molar activity via repositioning the 18F-radiolabel
Scientific Reports (2024)
-
Properties and biotechnological applications of microbial deacetylase
Applied Microbiology and Biotechnology (2023)
-
In the quest for histone deacetylase inhibitors: current trends in the application of multilayered computational methods
Amino Acids (2023)
-
Synergistic antitumor effect of histone deacetylase class IIa inhibitor with lenvatinib in hepatocellular carcinoma
Hepatology International (2023)
-
Recent advancement of HDAC inhibitors against breast cancer
Medical Oncology (2023)