Abstract
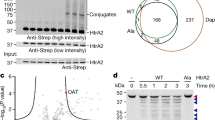
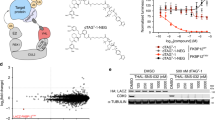
The ability to regulate any protein of interest in living systems with small molecules remains a challenge. We hypothesized that appending a hydrophobic moiety to the surface of a protein would mimic the partially denatured state of the protein, thus engaging the cellular quality control machinery to induce its proteasomal degradation. We designed and synthesized bifunctional small molecules to bind a bacterial dehalogenase (the HaloTag protein) and present a hydrophobic group on its surface. Hydrophobic tagging of the HaloTag protein with an adamantyl moiety induced the degradation of cytosolic, isoprenylated and transmembrane HaloTag fusion proteins in cell culture. We demonstrated the in vivo utility of hydrophobic tagging by degrading proteins expressed in zebrafish embryos and by inhibiting Hras1G12V-driven tumor progression in mice. Therefore, hydrophobic tagging of HaloTag fusion proteins affords small-molecule control over any protein of interest, making it an ideal system for validating potential drug targets in disease models.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout




Similar content being viewed by others
References
Overington, J.P., Al-Lazikani, B. & Hopkins, A.L. How many drug targets are there? Nat. Rev. Drug Discov. 5, 993–996 (2006).
Russ, A.P. & Lampel, S. The druggable genome: an update. Drug Discov. Today 10, 1607–1610 (2005).
Dixon, S.J. & Stockwell, B.R. Identifying druggable disease-modifying gene products. Curr. Opin. Chem. Biol. 13, 549–555 (2009).
Crews, C.M. Targeting the undruggable proteome: the small molecules of my dreams. Chem. Biol. 17, 551–555 (2010).
Luo, J. et al. A genome-wide RNAi screen identifies multiple synthetic lethal interactions with the Ras oncogene. Cell 137, 835–848 (2009).
Krishnan, M.N. et al. RNA interference screen for human genes associated with West Nile virus infection. Nature 455, 242–245 (2008).
Karlas, A. et al. Genome-wide RNAi screen identifies human host factors crucial for influenza virus replication. Nature 463, 818–822 (2010).
Whitehead, K.A., Langer, R. & Anderson, D.G. Knocking down barriers: advances in siRNA delivery. Nat. Rev. Drug Discov. 8, 129–138 (2009).
Schrader, E.K., Wilmington, S.R. & Matouschek, A. Making it easier to regulate protein stability. Chem. Biol. 17, 917–918 (2010).
Nishimura, K., Fukagawa, T., Takisawa, H., Kakimoto, T. & Kanemaki, M. An auxin-based degron system for the rapid depletion of proteins in nonplant cells. Nat. Methods 6, 917–922 (2009).
Schneekloth, J.S. Jr. et al. Chemical genetic control of protein levels: selective in vivo targeted degradation. J. Am. Chem. Soc. 126, 3748–3754 (2004).
Sakamoto, K.M. et al. Protacs: chimeric molecules that target proteins to the Skp1-Cullin-F box complex for ubiquitination and degradation. Proc. Natl. Acad. Sci. USA 98, 8554–8559 (2001).
Robinson, M.S., Sahlender, D.A. & Foster, S.D. Rapid inactivation of proteins by rapamycin-induced rerouting to mitochondria. Dev. Cell 18, 324–331 (2010).
Iwamoto, M., Bjorklund, T., Lundberg, C., Kirik, D. & Wandless, T.J. A general chemical method to regulate protein stability in the mammalian central nervous system. Chem. Biol. 17, 981–988 (2010).
Banaszynski, L.A., Chen, L.C., Maynard-Smith, L.A., Ooi, A.G. & Wandless, T.J. A rapid, reversible, and tunable method to regulate protein function in living cells using synthetic small molecules. Cell 126, 995–1004 (2006).
Clackson, T. et al. Redesigning an FKBP-ligand interface to generate chemical dimerizers with novel specificity. Proc. Natl. Acad. Sci. USA 95, 10437–10442 (1998).
Herm-Götz, A. et al. Rapid control of protein level in the apicomplexan Toxoplasma gondii. Nat. Methods 4, 1003–1005 (2007).
Banaszynski, L.A., Sellmyer, M.A., Contag, C.H., Wandless, T.J. & Thorne, S.H. Chemical control of protein stability and function in living mice. Nat. Med. 14, 1123–1127 (2008).
Dvorin, J.D. et al. A plant-like kinase in Plasmodium falciparum regulates parasite egress from erythrocytes. Science 328, 910–912 (2010).
Pruett-Miller, S.M., Reading, D.W., Porter, S.N. & Porteus, M.H. Attenuation of zinc finger nuclease toxicity by small-molecule regulation of protein levels. PLoS Genet. 5, e1000376 (2009).
Agashe, V.R., Shastry, M.C. & Udgaonkar, J.B. Initial hydrophobic collapse in the folding of barstar. Nature 377, 754–757 (1995).
Gething, M.J. Role and regulation of the ER chaperone BiP. Semin. Cell Dev. Biol. 10, 465–472 (1999).
Lins, L. & Brasseur, R. The hydrophobic effect in protein folding. FASEB J. 9, 535–540 (1995).
Blond-Elguindi, S. et al. Affinity panning of a library of peptides displayed on bacteriophages reveals the binding specificity of BiP. Cell 75, 717–728 (1993).
Kubota, H. Quality control against misfolded proteins in the cytosol: a network for cell survival. J. Biochem. 146, 609–616 (2009).
Los, G.V. et al. HaloTag: a novel protein labeling technology for cell imaging and protein analysis. ACS Chem. Biol. 3, 373–382 (2008).
Mathias, L.J., Jensen, J.J., Reichert, V.T., Lewis, C.M. & Tullos, G.L. Adamantane-containing polymers. ACS Symp. Ser. 624, 197–207 (1996).
Tsuzuki, N. et al. Adamantane as a brain-directed drug carrier for poorly absorbed drug. 2. AZT derivatives conjugated with the 1-adamantane moiety. J. Pharm. Sci. 83, 481–484 (1994).
Elofsson, M., Splittgerber, U., Myung, J., Mohan, R. & Crews, C.M. Towards subunit-specific proteasome inhibitors: synthesis and evaluation of peptide alpha',beta'-epoxyketones. Chem. Biol. 6, 811–822 (1999).
Oishi, I. et al. The receptor tyrosine kinase Ror2 is involved in non-canonical Wnt5a/JNK signalling pathway. Genes Cells 8, 645–654 (2003).
DeJarnette, J.B. et al. Specific requirement for CD3ɛ in T cell development. Proc. Natl. Acad. Sci. USA 95, 14909–14914 (1998).
Masellis-Smith, A. & Shaw, A.R. CD9-regulated adhesion. Anti-CD9 monoclonal antibody induce pre-B cell adhesion to bone marrow fibroblasts through de novo recognition of fibronectin. J. Immunol. 152, 2768–2777 (1994).
Briscoe, C.P. et al. The orphan G protein–coupled receptor GPR40 is activated by medium and long chain fatty acids. J. Biol. Chem. 278, 11303–11311 (2003).
Kirikoshi, H. et al. Molecular cloning and characterization of human Frizzled-4 on chromosome 11q14-q21. Biochem. Biophys. Res. Commun. 264, 955–961 (1999).
Bos, J.L. ras oncogenes in human cancer: a review. Cancer Res. 49, 4682–4689 (1989).
Parada, L.F., Tabin, C.J., Shih, C. & Weinberg, R.A. Human EJ bladder carcinoma oncogene is homologue of Harvey sarcoma virus ras gene. Nature 297, 474–478 (1982).
Shih, C. & Weinberg, R.A. Isolation of a transforming sequence from a human bladder carcinoma cell line. Cell 29, 161–169 (1982).
Porteus, M. Design and testing of zinc finger nucleases for use in mammalian cells. Methods Mol. Biol. 435, 47–61 (2008).
Ostrand-Rosenberg, S. Animal models of tumor immunity, immunotherapy and cancer vaccines. Curr. Opin. Immunol. 16, 143–150 (2004).
Rago, C., Vogelstein, B. & Bunz, F. Genetic knockouts and knockins in human somatic cells. Nat. Protoc. 2, 2734–2746 (2007).
Koh, E.Y., Chen, T. & Daley, G.Q. Novel retroviral vectors to facilitate expression screens in mammalian cells. Nucleic Acids Res. 30, e142 (2002).
Gies, E. et al. Niclosamide prevents the formation of large ubiquitin-containing aggregates caused by proteasome inhibition. PLoS ONE 5, e14410 (2010).
Link, V., Shevchenko, A. & Heisenberg, C.P. Proteomics of early zebrafish embryos. BMC Dev. Biol. 6, 1 (2006).
Acknowledgements
We wish to acknowledge financial support from the US National Institutes of Health (R01AI084140) and to thank the members of the Crews lab for critical reading of the manuscript. S. Stricker at the Max Planck Institute for Molecular Genetics kindly provided Mouse Ror2, and R. Weinberg at the Massachusetts Institute of Technology (MIT) contributed Addgene plasmid 9051 from which Hras1G12V was obtained. G. Daley at MIT kindly provided the retroviral pEYK3.1 vector. T.W.C. was the Canadian Institutes of Health Research Jean-François St-Denis Fellow in Cancer Research and a Bisby Fellow. T.B.S. is a recipient of an American Cancer Society fellowship.
Author information
Authors and Affiliations
Contributions
T.K.N., H.S.T., A.R.S. and C.M.C. designed the research. T.K.N., H.S.T., A.R.S., M.J.S., T.W.C., T.B.S. and K.R. performed the experiments. T.K.N., H.S.T., S.A.H. and C.M.C. analyzed the data. T.K.N., H.S.T. and C.M.C. wrote and edited the manuscript.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Methods and Supplementary Results (PDF 4459 kb)
Rights and permissions
About this article
Cite this article
Neklesa, T., Tae, H., Schneekloth, A. et al. Small-molecule hydrophobic tagging–induced degradation of HaloTag fusion proteins. Nat Chem Biol 7, 538–543 (2011). https://doi.org/10.1038/nchembio.597
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.597
This article is cited by
-
Exploiting E3 ubiquitin ligases to reeducate the tumor microenvironment for cancer therapy
Experimental Hematology & Oncology (2023)
-
Targeting selenoprotein H in the nucleolus suppresses tumors and metastases by Isovalerylspiramycin I
Journal of Experimental & Clinical Cancer Research (2022)
-
A proteolysis-targeting chimera molecule selectively degrades ENL and inhibits malignant gene expression and tumor growth
Journal of Hematology & Oncology (2022)
-
Emerging strategies to target RAS signaling in human cancer therapy
Journal of Hematology & Oncology (2021)
-
Protein degradation technology: a strategic paradigm shift in drug discovery
Journal of Hematology & Oncology (2021)