Abstract
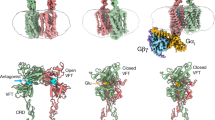
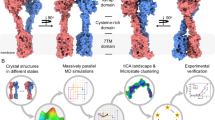
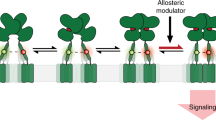
G protein–coupled receptors (GPCRs) are major players in cell communication. Although they form functional monomers, increasing evidence indicates that GPCR dimerization has a critical role in cooperative phenomena that are important for cell signal integration. However, the structural bases of these phenomena remain elusive. Here, using well-characterized receptor dimers, the metabotropic glutamate receptors (mGluRs), we show that structural changes at the dimer interface are linked to receptor activation. We demonstrate that the main dimer interface is formed by transmembrane α helix 4 (TM4) and TM5 in the inactive state and by TM6 in the active state. This major change in the dimer interface is required for receptor activity because locking the TM4-TM5 interface prevents activation by agonist, whereas locking the TM6 interface leads to a constitutively active receptor. These data provide important information on the activation mechanism of mGluRs and improve our understanding of the structural basis of the negative cooperativity observed in these GPCR dimers.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$259.00 per year
only $21.58 per issue
Buy this article
- Purchase on SpringerLink
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout






Similar content being viewed by others
References
Lagerström, M.C. & Schiöth, H.B. Structural diversity of G protein–coupled receptors and significance for drug discovery. Nat. Rev. Drug Discov. 7, 339–357 (2008).
Prézeau, L. et al. Functional crosstalk between GPCRs: with or without oligomerization. Curr. Opin. Pharmacol. 10, 6–13 (2010).
Ferré, S. et al. G protein–coupled receptor oligomerization revisited: functional and pharmacological perspectives. Pharmacol. Rev. 66, 413–434 (2014).
Smith, N.J. & Milligan, G. Allostery at G protein–coupled receptor homo- and heteromers: uncharted pharmacological landscapes. Pharmacol. Rev. 62, 701–725 (2010).
Albizu, L. et al. Time-resolved FRET between GPCR ligands reveals oligomers in native tissues. Nat. Chem. Biol. 6, 587–594 (2010).
Birdsall, N.J. Class A GPCR heterodimers: evidence from binding studies. Trends Pharmacol. Sci. 31, 499–508 (2010).
Springael, J.Y., Urizar, E., Costagliola, S., Vassart, G. & Parmentier, M. Allosteric properties of G protein–coupled receptor oligomers. Pharmacol. Ther. 115, 410–418 (2007).
Cherezov, V. et al. High-resolution crystal structure of an engineered human β2-adrenergic G protein–coupled receptor. Science 318, 1258–1265 (2007).
Huang, J., Chen, S., Zhang, J.J. & Huang, X.Y. Crystal structure of oligomeric β1-adrenergic G protein–coupled receptors in ligand-free basal state. Nat. Struct. Mol. Biol. 20, 419–425 (2013).
Liu, W. et al. Structural basis for allosteric regulation of GPCRs by sodium ions. Science 337, 232–236 (2012).
Manglik, A. et al. Crystal structure of the μ-opioid receptor bound to a morphinan antagonist. Nature 485, 321–326 (2012).
Wu, B. et al. Structures of the CXCR4 chemokine GPCR with small-molecule and cyclic peptide antagonists. Science 330, 1066–1071 (2010).
Ruprecht, J.J., Mielke, T., Vogel, R., Villa, C. & Schertler, G.F. Electron crystallography reveals the structure of metarhodopsin I. EMBO J. 23, 3609–3620 (2004).
Guo, W., Shi, L., Filizola, M., Weinstein, H. & Javitch, J.A. Crosstalk in G protein–coupled receptors: changes at the transmembrane homodimer interface determine activation. Proc. Natl. Acad. Sci. USA 102, 17495–17500 (2005).
Mancia, F., Assur, Z., Herman, A.G., Siegel, R. & Hendrickson, W.A. Ligand sensitivity in dimeric associations of the serotonin 5HT2c receptor. EMBO Rep. 9, 363–369 (2008).
Hebert, T.E. et al. A peptide derived from a β2-adrenergic receptor transmembrane domain inhibits both receptor dimerization and activation. J. Biol. Chem. 271, 16384–16392 (1996).
Guo, W. et al. Dopamine D2 receptors form higher order oligomers at physiological expression levels. EMBO J. 27, 2293–2304 (2008).
Calebiro, D. et al. Single-molecule analysis of fluorescently labeled G-protein–coupled receptors reveals complexes with distinct dynamics and organization. Proc. Natl. Acad. Sci. USA 110, 743–748 (2013).
Hern, J.A. et al. Formation and dissociation of M1 muscarinic receptor dimers seen by total internal reflection fluorescence imaging of single molecules. Proc. Natl. Acad. Sci. USA 107, 2693–2698 (2010).
Kasai, R.S. & Kusumi, A. Single-molecule imaging revealed dynamic GPCR dimerization. Curr. Opin. Cell Biol. 27, 78–86 (2014).
Rondard, P., Goudet, C., Kniazeff, J., Pin, J.-P. & Prézeau, L. The complexity of their activation mechanism opens new possibilities for the modulation of mGlu and GABAB class C G protein–coupled receptors. Neuropharmacology 60, 82–92 (2011).
Brock, C. et al. Activation of a dimeric metabotropic glutamate receptor by intersubunit rearrangement. J. Biol. Chem. 282, 33000–33008 (2007).
Doumazane, E. et al. A new approach to analyze cell surface protein complexes reveals specific heterodimeric metabotropic glutamate receptors. FASEB J. 25, 66–77 (2011).
Maurel, D. et al. Cell-surface protein-protein interaction analysis with time-resolved FRET and snap-tag technologies: application to GPCR oligomerization. Nat. Methods 5, 561–567 (2008).
Hlavackova, V. et al. Evidence for a single heptahelical domain being turned on upon activation of a dimeric GPCR. EMBO J. 24, 499–509 (2005).
Hlavackova, V. et al. Sequential inter- and intrasubunit rearrangements during activation of dimeric metabotropic glutamate receptor 1. Sci. Signal. 5, ra59 (2012).
Tateyama, M., Abe, H., Nakata, H., Saito, O. & Kubo, Y. Ligand-induced rearrangement of the dimeric metabotropic glutamate receptor 1α. Nat. Struct. Mol. Biol. 11, 637–642 (2004).
El Moustaine, D. et al. Distinct roles of metabotropic glutamate receptor dimerization in agonist activation and G-protein coupling. Proc. Natl. Acad. Sci. USA 109, 16342–16347 (2012).
Doré, A.S. et al. Structure of class C GPCR metabotropic glutamate receptor 5 transmembrane domain. Nature 511, 557–562 (2014).
Wu, H. et al. Structure of a class C GPCR metabotropic glutamate receptor 1 bound to an allosteric modulator. Science 344, 58–64 (2014).
Binet, V. et al. Common structural requirements for heptahelical domain function in class A and class C G protein–coupled receptors. J. Biol. Chem. 282, 12154–12163 (2007).
Huang, S. et al. Interdomain movements in metabotropic glutamate receptor activation. Proc. Natl. Acad. Sci. USA 108, 15480–15485 (2011).
Doumazane, E. et al. Illuminating the activation mechanisms and allosteric properties of metabotropic glutamate receptors. Proc. Natl. Acad. Sci. USA 110, E1416–E1425 (2013).
Manglik, A. & Kobilka, B. The role of protein dynamics in GPCR function: insights from the β2AR and rhodopsin. Curr. Opin. Cell Biol. 27, 136–143 (2014).
Rasmussen, S.G. et al. Crystal structure of the β2 adrenergic receptor-Gs protein complex. Nature 477, 549–555 (2011).
Pin, J.-P., Galvez, T. & Prézeau, L. Evolution, structure and activation mechanism of family 3/C G-protein coupled receptors. Pharmacol. Ther. 98, 325–354 (2003).
Nordström, K.J., Sällman Almén, M., Edstam, M.M., Fredriksson, R. & Schiöth, H.B. Independent HHsearch, Needleman-Wunsch-based, and motif analyses reveal the overall hierarchy for most of the G protein–coupled receptor families. Mol. Biol. Evol. 28, 2471–2480 (2011).
Knepp, A.M., Periole, X., Marrink, S.J., Sakmar, T.P. & Huber, T. Rhodopsin forms a dimer with cytoplasmic helix 8 contacts in native membranes. Biochemistry 51, 1819–1821 (2012).
Periole, X., Knepp, A.M., Sakmar, T.P., Marrink, S.J. & Huber, T. Structural determinants of the supramolecular organization of G protein–coupled receptors in bilayers. J. Am. Chem. Soc. 134, 10959–10965 (2012).
Kunishima, N. et al. Structural basis of glutamate recognition by a dimeric metabotropic glutamate receptor. Nature 407, 971–977 (2000).
Muto, T., Tsuchiya, D., Morikawa, K. & Jingami, H. Structures of the extracellular regions of the group II/III metabotropic glutamate receptors. Proc. Natl. Acad. Sci. USA 104, 3759–3764 (2007).
Parmentier, M.-L., Prézeau, L., Bockaert, J. & Pin, J.-P. A model for the functioning of family 3 GPCRs. Trends Pharmacol. Sci. 23, 268–274 (2002).
Geng, Y., Bush, M., Mosyak, L., Wang, F. & Fan, Q.R. Structural mechanism of ligand activation in human GABAB receptor. Nature 504, 254–259 (2013).
Monnier, C. et al. Trans-activation between 7TM domains: implication in heterodimeric GABAB receptor activation. EMBO J. 30, 32–42 (2011).
Damian, M., Martin, A., Mesnier, D., Pin, J.-P. & Banères, J.L. Asymmetric conformational changes in a GPCR dimer controlled by G-proteins. EMBO J. 25, 5693–5702 (2006).
Han, Y., Moreira, I.S., Urizar, E., Weinstein, H. & Javitch, J.A. Allosteric communication between protomers of dopamine class A GPCR dimers modulates activation. Nat. Chem. Biol. 5, 688–695 (2009).
Rovira, X., Pin, J.-P. & Giraldo, J. The asymmetric/symmetric activation of GPCR dimers as a possible mechanistic rationale for multiple signalling pathways. Trends Pharmacol. Sci. 31, 15–21 (2010).
Comps-Agrar, L. et al. The oligomeric state sets GABAB receptor signalling efficacy. EMBO J. 30, 2336–2349 (2011).
Fribourg, M. et al. Decoding the signaling of a GPCR heteromeric complex reveals a unifying mechanism of action of antipsychotic drugs. Cell 147, 1011–1023 (2011).
Moreno, J.L. et al. Identification of three residues essential for 5-hydroxytryptamine 2A-metabotropic glutamate 2 (5–HT2A.mGlu2) receptor heteromerization and its psychoactive behavioral function. J. Biol. Chem. 287, 44301–44319 (2012).
Šali, A. & Blundell, T.L. Comparative protein modelling by satisfaction of spatial restraints. J. Mol. Biol. 234, 779–815 (1993).
Larkin, M.A. et al. Clustal W and Clustal X version 2.0. Bioinformatics 23, 2947–2948 (2007).
Shen, M.Y. & Sali, A. Statistical potential for assessment and prediction of protein structures. Protein Sci. 15, 2507–2524 (2006).
Pettersen, E.F. et al. UCSF Chimera—a visualization system for exploratory research and analysis. J. Comput. Chem. 25, 1605–1612 (2004).
Tsuchiya, D., Kunishima, N., Kamiya, N., Jingami, H. & Morikawa, K. Structural views of the ligand-binding cores of a metabotropic glutamate receptor complexed with an antagonist and both glutamate and Gd3+. Proc. Natl. Acad. Sci. USA 99, 2660–2665 (2002).
Acknowledgements
We thank the Cisbio Company for their support in providing reagents. The intracellular calcium release and FRET experiments were performed at the ARPEGE (Pharmacology Screening-Interactome) facility, Institut de Génomique Fonctionnelle (Montpellier, France). J.L. was supported by the National Natural Science Foundation of China (grant numbers 31130028 and 31225011), the Ministry of Science and Technology (grant number 2012CB518000), the Program for Introducing Talents of Discipline to the Universities of the Ministry of Education (grant number B08029) and the Mérieux Research Grants Program of the Institut Mérieux. P.R. and J.-P.P. are supported by the Centre National de la Recherche Scientifique, the Institut National de la Santé et de la Recherche Médicale and by grants from the Agence Nationale de la Recherche (ANR-09-PIRI-0011), the Fondation pour la Recherche Médicale (Equipe FRM DEQ20130326522) and the Fondation Bettencourt Schueller. L.X. was supported by a doctoral fellowship from the French Embassy in China and X.R. by a FEBS long-term fellowship and by a Agència de Gestió d'Ajuts Universitaris i de Recerca (AGAUR) BP post-doctoral fellowship.
Author information
Authors and Affiliations
Contributions
L.X., X.R., J.L., J.-P.P. and P.R. designed experiments; L.X. performed molecular biology, cross-linking, functional assays and FRET experiments; X.R. performed molecular modeling; P.S. performed FRET experiments; H.Z. performed molecular biology; J.L., J.-P.P. and P.R. wrote the manuscript.
Corresponding authors
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Supplementary information
Supplementary Text and Figures
Supplementary Results and Supplementary Figures 1–13. (PDF 12218 kb)
Supplementary Video 1
Movie to illustrate the changes of the mGluR2 ECD dimer between the resting and active states (side view). (MP4 510 kb)
Supplementary Video 2
Movie to illustrate the changes of the mGluR2 ECD dimer between the resting and active states (bottom view). (MP4 954 kb)
Supplementary Video 3
Movie to illustrate the major rearrangement of the mGluR2 7TM dimer between the resting and active states (top view). (MP4 1280 kb)
Rights and permissions
About this article
Cite this article
Xue, L., Rovira, X., Scholler, P. et al. Major ligand-induced rearrangement of the heptahelical domain interface in a GPCR dimer. Nat Chem Biol 11, 134–140 (2015). https://doi.org/10.1038/nchembio.1711
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/nchembio.1711
This article is cited by
-
Structural basis of dimerization of chemokine receptors CCR5 and CXCR4
Nature Communications (2023)
-
Kinetic fingerprinting of metabotropic glutamate receptors
Communications Biology (2023)
-
Mechanism of sensitivity modulation in the calcium-sensing receptor via electrostatic tuning
Nature Communications (2022)
-
Biased signaling due to oligomerization of the G protein-coupled platelet-activating factor receptor
Nature Communications (2022)
-
Disruption of 5-hydroxytryptamine 1A receptor and orexin receptor 1 heterodimer formation affects novel G protein-dependent signaling pathways and has antidepressant effects in vivo
Translational Psychiatry (2022)