Abstract
Pluripotent mouse embryonic stem cells maintain their identity throughout virtually infinite cell divisions. This phenomenon, referred to as self-renewal, depends on a network of sequence-specific transcription factors (TFs) and requires daughter cells to accurately reproduce the gene expression pattern of the mother. However, dramatic chromosomal changes take place in mitosis, generally leading to the eviction of TFs from chromatin. Here, we report that Esrrb, a major pluripotency TF, remains bound to key regulatory regions during mitosis. We show that mitotic Esrrb binding is highly dynamic, driven by specific recognition of its DNA-binding motif and is associated with early transcriptional activation of target genes after completion of mitosis. These results indicate that Esrrb may act as a mitotic bookmarking factor, opening another perspective to molecularly understand the role of sequence-specific TFs in the epigenetic control of self-renewal, pluripotency and genome reprogramming.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout







Similar content being viewed by others
References
Smith, A. G. et al. Inhibition of pluripotential embryonic stem cell differentiation by purified polypeptides. Nature 336, 688–690 (1988).
Kim, J., Chu, J., Shen, X., Wang, J. & Orkin, S. H. An extended transcriptional network for pluripotency of embryonic stem cells. Cell 132, 1049–1061 (2008).
Chen, X. et al. Integration of external signaling pathways with the core transcriptional network in embryonic stem cells. Cell 133, 1106–1117 (2008).
Chambers, I. et al. Nanog safeguards pluripotency and mediates germline development. Nature 450, 1230–1234 (2007).
Nichols, J. et al. Formation of pluripotent stem cells in the mammalian embryo depends on the POU transcription factor Oct4. Cell 95, 379–391 (1998).
Niwa, H., Miyazaki, J. & Smith, A. G. Quantitative expression of Oct-3/4 defines differentiation, dedifferentiation or self-renewal of ES cells. Nat. Genet. 24, 372–376 (2000).
Masui, S. et al. Pluripotency governed by Sox2 via regulation of Oct3/4 expression in mouse embryonic stem cells. Nat. Cell Biol. 9, 625–635 (2007).
Festuccia, N. et al. Esrrb is a direct Nanog target gene that can substitute for Nanog function in pluripotent cells. Cell Stem Cell 11, 477–490 (2012).
Martello, G. et al. Esrrb is a pivotal target of the gsk3/tcf3 axis regulating embryonic stem cell self-renewal. Cell Stem Cell 11, 491–504 (2012).
Kopp, J. L., Ormsbee, B. D., Desler, M. & Rizzino, A. Small increases in the level of Sox2 trigger the differentiation of mouse embryonic stem cells. Stem Cells 26, 903–911 (2008).
Chambers, I. et al. Functional expression cloning of Nanog, a pluripotency sustaining factor in embryonic stem cells. Cell 113, 643–655 (2003).
Stead, E. et al. Pluripotent cell division cycles are driven by ectopic Cdk2, cyclin A/E and E2F activities. Oncogene 21, 8320–8333 (2002).
White, J. et al. Developmental activation of the Rb-E2F pathway and establishment of cell cycle-regulated cyclin-dependent kinase activity during embryonic stem cell differentiation. Mol. Biol. Cell 16, 2018–2027 (2005).
Faast, R. et al. Cdk6-cyclin D3 activity in murine ES cells is resistant to inhibition by p16(INK4a). Oncogene 23, 491–502 (2004).
Dalton, S. Linking the cell cycle to cell fate decisions. Trends Cell Biol. 25, 592–600 (2015).
Martinez-Balbas, M. A., Dey, A., Rabindran, S. K., Ozato, K. & Wu, C. Displacement of sequence-specific transcription factors from mitotic chromatin. Cell 83, 29–38 (1995).
Tsumura, A. et al. Maintenance of self-renewal ability of mouse embryonic stem cells in the absence of DNA methyltransferases Dnmt1, Dnmt3a and Dnmt3b. Genes Cells 11, 805–814 (2006).
Leeb, M. et al. Polycomb complexes act redundantly to repress genomic repeats and genes. Genes Dev. 24, 265–276 (2010).
Lehnertz, B. et al. Suv39h-mediated histone H3 lysine 9 methylation directs DNA methylation to major satellite repeats at pericentric heterochromatin. Curr. Biol. 13, 1192–1200 (2003).
Tachibana, M. et al. G9a histone methyltransferase plays a dominant role in euchromatic histone H3 lysine 9 methylation and is essential for early embryogenesis. Genes Dev. 16, 1779–1791 (2002).
De Los Angeles, A. et al. Hallmarks of pluripotency. Nature 525, 469–478 (2015).
Michelotti, E. F., Sanford, S. & Levens, D. Marking of active genes on mitotic chromosomes. Nature 388, 895–899 (1997).
Xing, H., Vanderford, N. L. & Sarge, K. D. The TBP-PP2A mitotic complex bookmarks genes by preventing condensin action. Nat. Cell Biol. 10, 1318–1323 (2008).
Zaidi, S. K. et al. Mitotic partitioning and selective reorganization of tissue-specific transcription factors in progeny cells. Proc. Natl Acad. Sci. USA 100, 14852–14857 (2003).
Blobel, G. A. et al. A reconfigured pattern of MLL occupancy within mitotic chromatin promotes rapid transcriptional reactivation following mitotic exit. Mol. Cell 36, 970–983 (2009).
Young, D. W. et al. Mitotic occupancy and lineage-specific transcriptional control of rRNA genes by Runx2. Nature 445, 442–446 (2007).
Verdeguer, F. et al. A mitotic transcriptional switch in polycystic kidney disease. Nat. Med. 16, 106–110 (2009).
Caravaca, J. M. et al. Bookmarking by specific and nonspecific binding of FoxA1 pioneer factor to mitotic chromosomes. Genes Dev. 27, 251–260 (2013).
Kadauke, S. et al. Tissue-specific mitotic bookmarking by hematopoietic transcription factor GATA1. Cell 150, 725–737 (2012).
Guo, G. et al. Resolution of cell fate decisions revealed by single-cell gene expression analysis from zygote to blastocyst. Dev. Cell 18, 675–685 (2010).
Whyte, W. A. et al. Master transcription factors and mediator establish super-enhancers at key cell identity genes. Cell 153, 307–319 (2013).
Shen, Y. et al. A map of the cis-regulatory sequences in the mouse genome. Nature 488, 116–120 (2012).
Brookes, E. et al. Polycomb associates genome-wide with a specific RNA polymerase II variant, and regulates metabolic genes in ESCs. Cell Stem Cell 10, 157–170 (2012).
Bulut-Karslioglu, A. Suv39h-dependent H3K9me3 marks intact retrotransposons and silences LINE elements in mouse embryonic stem cells. Mol. Cell 55, 277–290 (2014).
Farla, P. et al. The androgen receptor ligand-binding domain stabilizes DNA binding in living cells. J. Struct. Biol. 147, 50–61 (2004).
Sauer, M. et al. C-terminal diversity within the p53 family accounts for differences in DNA binding and transcriptional activity. Nucleic Acids Res. 36, 1900–1912 (2008).
Mueller, F., Stasevich, T. J., Mazza, D. & McNally, J. G. Quantifying transcription factor kinetics: at work or at play? Crit. Rev. Biochem. Mol. Biol. 48, 492–514 (2013).
den Elzen, N. & Pines, J. Cyclin A is destroyed in prometaphase and can delay chromosome alignment and anaphase. J. Cell Biol. 153, 121–136 (2001).
Geley, S. et al. Anaphase-promoting complex/cyclosome-dependent proteolysis of human cyclin A starts at the beginning of mitosis and is not subject to the spindle assembly checkpoint. J. Cell Biol. 153, 137–148 (2001).
Marks, H. et al. The transcriptional and epigenomic foundations of ground state pluripotency. Cell 149, 590–604 (2012).
Dixon, J. R. et al. Topological domains in mammalian genomes identified by analysis of chromatin interactions. Nature 485, 376–380 (2012).
Feng, B. et al. Reprogramming of fibroblasts into induced pluripotent stem cells with orphan nuclear receptor Esrrb. Nat. Cell Biol. 11, 197–203 (2009).
Lerner, J. et al. Human mutations affect the epigenetic/bookmarking function of HNF1B. Nucleic Acids Res. http://dx.doi.org/10.1093/nar/gkw467 (2016).
Soufi, A., Donahue, G. & Zaret, K. S. Facilitators and impediments of the pluripotency reprogramming factors’ initial engagement with the genome. Cell 151, 994–1004 (2012).
Soufi, A. et al. Pioneer transcription factors target partial DNA motifs on nucleosomes to initiate reprogramming. Cell 161, 555–568 (2015).
van den Berg, D. L. et al. An Oct4-centered protein interaction network in embryonic stem cells. Cell Stem Cell 6, 369–381 (2010).
Zaidi, S. K. et al. Mitotic bookmarking of genes: a novel dimension to epigenetic control. Nat. Rev. Genet. 11, 583–589 (2010).
Kadauke, S. & Blobel, G. A. Mitotic bookmarking by transcription factors. Epigenet. Chromatin 6, 6 (2013).
Zaret, K. S. Genome reactivation after the silence in mitosis: recapitulating mechanisms of development? Dev. Cell 29, 132–134 (2014).
Sela, Y., Molotski, N., Golan, S., Itskovitz-Eldor, J. & Soen, Y. Human embryonic stem cells exhibit increased propensity to differentiate during the G1 phase prior to phosphorylation of retinoblastoma protein. Stem Cells 30, 1097–1108 (2012).
Pauklin, S. & Vallier, L. The cell-cycle state of stem cells determines cell fate propensity. Cell 155, 135–147 (2013).
Boroviak, T., Loos, R., Bertone, P., Smith, A. & Nichols, J. The ability of inner-cell-mass cells to self-renew as embryonic stem cells is acquired following epiblast specification. Nat. Cell Biol. 16, 516–528 (2014).
Buganim, Y. et al. Single-cell expression analyses during cellular reprogramming reveal an early stochastic and a late hierarchic phase. Cell 150, 1209–1222 (2012).
Gearhart, M. D., Holmbeck, S. M., Evans, R. M., Dyson, H. J. & Wright, P. E. Monomeric complex of human orphan estrogen related receptor-2 with DNA: a pseudo-dimer interface mediates extended half-site recognition. J. Mol. Biol. 327, 819–832 (2003).
Mueller, F., Karpova, T. S., Mazza, D. & McNally, J. G. Monitoring dynamic binding of chromatin proteins in vivo by fluorescence recovery after photobleaching. Methods Mol. Biol. 833, 153–176 (2012).
van den Berg, D. L. et al. Estrogen-related receptor β interacts with Oct4 to positively regulate Nanog gene expression. Mol. Cell Biol. 28, 5986–5995 (2008).
Halbritter, F., Vaidya, H. J. & Tomlinson, S. R. GeneProf: analysis of high-throughput sequencing experiments. Nat. Methods 9, 7–8 (2012).
McLean, C. Y. et al. GREAT improves functional interpretation of cis-regulatory regions. Nat. Biotechnol. 28, 495–501 (2010).
Medina-Rivera, A. et al. RSAT 2015: regulatory sequence analysis tools. Nucleic Acids Res. 43, W50–W56 (2015).
Madsen, J. G. et al. iRNA-seq: computational method for genome-wide assessment of acute transcriptional regulation from total RNA-seq data. Nucleic Acids Res. 43, e40 (2015).
Acknowledgements
We are grateful to I. Chambers for allowing N.F. and P.N. to pursue observations made initially in his laboratory on the chromosomal retention of Esrrb during mitosis. We are grateful to A. Rada-Iglesias, G. Cecere, S. Tajbakhsh, M. Yaniv and members of our laboratory for critical reading of the manuscript. We acknowledge B. Jagla for running the iRNA-Seq pipeline and P. Clerc for assistance with fluorescence microscopy. P.N. acknowledges financial support to the Institut Pasteur, CNRS and the ANR Laboratoire d’Excellence Revive (Investissement d’Avenir; ANR-10-LABX-73). N.F was funded by an EMBO Long Term Fellowship (ALTF 876-2013) and a Marie-Curie Intra-European Fellowship (EFIMB – 626705).
Author information
Authors and Affiliations
Contributions
This study was conceived by N.F. and P.N. All experiments were carried out by N.F. with the exception of embryo work performed by S.V.-P., S.B. and M.C.-T. In addition, A.D., E.G.T. and A.M. provided technical help and F.M. contributed to the analysis of FRAP assays. C.P. performed next-generation sequencing and P.N. conducted the Bioinformatic analyses. All experiments were analysed and interpreted by N.F. and P.N. The manuscript was written by P.N. with input from N.F.
Corresponding author
Ethics declarations
Competing interests
The authors declare no competing financial interests.
Integrated supplementary information
Supplementary Figure 1 Cell lines used in this study.
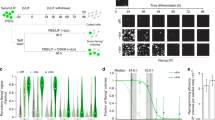
(a) Esrrb immuno-fluorescence on EKOiE cells cultured in the presence or absence of Dox (24 h withdrawal). Scale bars represent 30 μm. Images are representative of multiple field acquired in parallel. (b) The self-renewal efficiency of wild-type E14Tg2a, TNG, EKOiE and the respective derivative cell lines was assessed by Alkaline Phosphatase staining of cell colonies obtained 6 days after clonal plating in the indicated conditions. The graph shows the average and s.d. of two independent experiments, each performed in duplicate wells (n = 4). E14Tg2a cells self-renew exclusively in the presence of LIF. Upon introduction of a Nanog-GFP transgene, self-renewal efficiency increases in the presence of LIF and is rescued in its absence. As expected, TNG cells (Nanog heterozygous) exhibit reduced self-renewal as compared to E14Tg2a4. Upon knock-in of TdTomato at both Esrrb alleles, so that only Esrrb-TdT fusion protein is expressed in the cells, self-renewal efficiency is unaffected. EKOiE cells cultured without Dox show extremely severe defects in self-renewal efficiency as compared to Dox-treated cells, demonstrating the importance of Esrrb. In the presence of Dox, EKOiE cells also exhibit LIF-independent self-renewal. Upon the introduction of a constitutive Esrrb-GFP transgene, EKOiE cells are able to self-renew both in presence or absence of Dox; they also exhibit LIF-independent self-renewal. We conclude from this analysis that all the GFP and TdT fusion proteins (Nanog and Esrrb) are functional.
Supplementary Figure 2 Validation of Esrrb retention on mitotic chromosomes.
(a) Representative live-imaging of nocodazole-treated EKOiE cells carrying an Esrrb-GFP transgene and cultured in the absence of Dox. DNA was counterstained with Hoechst. Note that all mitotic cells display chromosomal retention of Esrrb. (b) Representative live-imaging of E14Tg2a cells expressing a Nanog-GFP transgene. DNA was counterstained with Hoechst. All mitotic cells display exclusion of Nanog from the chromosomes. (c) Representative live-imaging of nocodazole-treated Esrrb-TdTomato cells. DNA was counterstained with Hoechst. All mitotic cells show enrichment of Esrrb on the chromosomes. (d) Representative Esrrb immuno-staining on a full karyotype from EKOiE cells cultured in the presence (top) or absence (bottom) of Dox. DNA was counterstained with DAPI. (e) SNAP-Cell TMR-Star staining on a karyotype from EKOiE cells constitutively expressing Snap-tagged Esrrb and cultured without Dox. (f) Simultaneous immuno-staining for Nanog and Esrrb on a karyotype obtained from wild-type E14Tg2a cells. Scale bars represent 10 μm. Images are representative of multiple fields acquired in 1 (a), 3 (b), 2 (c), 6 (d), 2 (e) and 3 (f) independent repeat experiments.
Supplementary Figure 3 Additional information on Esrrb binding regions.
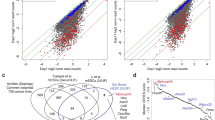
(a) The table highlights the criteria used to categorise peaks as bookmarked or lost. Classes_1,2,3 were considered as bookmarked (see Methods for further details). ∗Note that in class_1 all peaks (with one exception) were detected in at least one interphase dataset; they cannot be considered as mitotic-specific. (b) Esrrb average binding profile at the 5 classes of regions shown in a in both interphase (Int.) and mitosis (Mit.). All 5 classes show clear enrichment in interphase. Note that in mitosis Class_5 exhibits a complete depletion of Esrrb, whilst Class_4 shows low levels of Esrrb binding. The Y -axis represents alignment coverage expressed in bases per kb per million. (c) Aligned heat map of Esrrb density across the 5 classes of Esrrb binding regions. Note the focal accumulation of Esrrb at the centre of regions from all 5 classes in interphase (blue). In contrast, only Classes_1,2,3 show clear Esrrb binding in mitosis. (d) Heat-map of Esrrb enrichment levels (Log2_RPKM) at all binding regions in interphase (I) and mitosis (M) in all our datasets (E: Esrrb ChIP-Seq; C: Control obtained from Esrrb ChIP-Seq in EKOiE cultured without Dox; Mean represents the average of all shown replicates; Pooled represents enrichment levels after pooling together reads derived from all the replicates; Chen shows the enrichment levels in an independent ChIP-Seq3 for Esrrb in interphase (E: Esrrb ChIP-Seq; C: negative ChIP control against GFP). Note that across all regions the levels of Esrrb enrichment are significantly higher in interphase than in mitosis and all the controls. Also, note that in mitosis only Classes_1,2,3 show consistently high levels of enrichment. (e) Scatter plot representing enrichment levels in interphase (X-axis) versus mitosis (Y -axis) for the 5 classes of Esrrb binding regions. Note that Classes_1, 2 and 3 cluster together whilst Class_4 localises between Classes_1, 2 and 3 and Class_ 5. These analyses strongly support grouping together Classes_1,2,3 as bookmarked and Classes_4,5 as lost, even though a subset of Class_4 regions may display low levels of mitotic binding.
Supplementary Figure 4 ChIP-Seq profiles and validation by qPCR.
(a) Six bookmarked positions selected for spanning different levels of mitotic binding were selected along with five lost positions and analysed by ChIP-qPCR for Esrrb binding (n = 3; error bars represent s.d) in interphase (blue) and mitosis (red) in EKOiE cells cultured with Dox. Interphasic EKOiE cells cultured in the absence of Dox were used to control for the specificity of our assay (dark grey). Moreover, 5 and 10% of chromatin prepared from Dox-treated EKOiE cells was spiked into-Dox chromatin to test whether the contaminant interphasic cells (<5%) in our mitotic preparations (see Methods) are sufficient to produce a level of enrichment similar to that observed in mitosis. At bookmarked positions we observe mitotic enrichment levels (percentage IP, Y -axis) significantly higher than at lost positions. Moreover, even after introducing 10% of contaminant chromatin to EKOiE -Dox controls, the observed %IP remains much lower than in mitotic cells for all bookmarked positions tested. (b) Binding profile at three loci belonging to Class_1 (putative mitotic-specific peaks, see Supplementary Fig. 3) illustrating that even though the peak detected in interphase is not statistically significant, it is of similar magnitude to that observed in mitotic cells. (c) ChIP-qPCR validation of interphasic and mitotic binding at the 3 loci shown in (b), presented as in (a). (d) Bookmarked regions include ∼55% of ES-specific super-enhancers (for example, Tbx3 and Klf4 or Sox2 in Fig. 2a). Other noticeable regions bookmarked by Esrrb correspond to developmental genes (Otx2), chromatin remodelers (Ino80), Polycomb Group proteins (Jarid2), regulators of DNA methylation (Tet1), signalling mediators (Lif receptor) and metabolic genes (Eno1). Mitosis and interphase are shown at the same scale.
Supplementary Figure 5 B1 regions are more active than L1 regions.
(a) Enrichment levels of the indicated factors and chromatin modifications at the 4 clusters identified in Fig. 3a. The upper and lower extremities of each shaded area are the 25th and 75th percentile of each dataset across the 4 clusters. (b) Average profiles of the indicated factors and marks across promoters and enhancers belonging to B1 and L1 clusters as shown in Fig. 3.
Supplementary Figure 6 Additional information on the relationships between Esrrb bookmarking and the presence of Esrrb binding motifs.
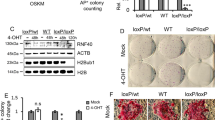
(a) ChIP-qPCR of wild-type (WT) and Mutant (Mut) Esrrb at several genomic positions (the graph shows the average of 2 independent experiments). The two variants are expressed in EKOiE or EKOiE Mut cells under the control of Dox (see methods; cells growing without Dox and not expressing Esrrb are used as a negative control – Ctl). (b) Representative live-imaging of mutant Esrrb-GFP carrying three mutations in residues known to establish specific contacts with the bases of the motif. DNA is counterstained with Hoechst. Note that no mitotic cell displays focal accumulation on the chromosomes. Scale bars represent 10 μm. Images are representative of multiple fields acquired in 4 independent repeat experiments. (c) The best motifs discovered are shown together with the percentage of regions containing at least one. Note that the regions containing a long motif may also contain a short motif. In contrast, the percentages given for the regions contain a short motif do not contain long motifs. (d) ChIP-Seq profile of Esrrb in interphase and mitosis across the Esrrb locus (coverage represents bases per kb per million). The arrow and dashed box indicate the position of the major enhancer of Esrrb shown at higher resolution on the right. The sequence of the long motif identified at this peak is shown; the 3 bases mutated to CCC are underlined. (e) Esrrb ChIP-qPCR at the Esrrb enhancer expressed as percentage of immunoprecipitation (for E14Tg2a n = 7 for interphase and n = 3 for mitosis; for EEt –edited to short n = 17 for interphase and n = 8 for mitosis; error bars represent s.d.) in E14Tg2a and in cells where the long motif was edited into a short motif (AGTTCAAGGTCA- > CCCTCAAGGTCA). No alteration of Esrrb binding in interphase or mitosis is observed (unpaired t-test). (f) Global quantification (RPKM) of Esrrb enrichment at bookmarked regions depending on whether they contain a long motif (red; n = 1246), only a short motif (blue; n = 537), or no Esrrb motif at all (grey; n = 197). Note that regions without short or long motifs display significantly higher enrichment in interphase and lower in mitosis. (g) Calculation of the percentage of mitotic retention at the three groups of Esrrb binding regions shown in (f). The presence of a long or a short binding site is associated with increased bookmarking efficiency. (h) Quantification of enrichment levels (RPKM) of several pluripotency TFs3 in the regions described in (f). All factors are statistically over-enriched at Esrrb binding regions lacking Esrrb motifs. The P-value of the difference between regions with long, only short or no motifs is indicated (Kruskal-Wallis ANOVA followed by Dunn’s multiple comparison test).
Supplementary Figure 7 Additional information of the Ccna-GFP reporter system and Esrrb-responsive genes.
(a) A transgene expressing GFP fused to the domain of Cyclin A2 (Ccna) responsible for its mitotic degradation41,42 was introduced in EKOiE cells that were stained with Hoechst to quantify DNA content. Ccna-GFP shows low fluorescence in M and early G1 and accumulates gradually before the cells enter into S phase. Asynchronous cells (before sorting) are presented along with sorted Early-G1, Late-G1 and G2 cells replated and cultured for additional 8 h. Approximate sorting gates are shown along with the percentages of cells falling in each gate to illustrate how EKOiE Ccna-GFP cells are able to resume cell cycle progression after sorting: the majority of cells sorted in early-G1 progressed to the beginning of the S phase; cells sorted in late-G1 largely entered S and many reached G2; cells in G2 were delayed but could enter mitosis. (b) Relative expression (Log2) of early-G1 responsive genes (Early-G1 cultured without Dox were set to 0) along the cell cycle. (c) Boxplot showing the log2 fold change of Esrrb-responsive early-G1 genes (n = 168 for upregulated genes; n = 206 for downregulated genes) in cells cultured in ground state conditions (2i) versus serum-containing medium43. The distribution of the fold changes followed a normal distribution (D’Agostino & Pearson normality test); therefore the P-values were calculated with a one-way Student test against a theoretical fold change of 0.
Supplementary information
Supplementary Information
Supplementary Information (PDF 2711 kb)
Supplementary Table 1
Supplementary Information (XLSX 2770 kb)
Supplementary Table 2
Supplementary Information (XLSX 9626 kb)
Supplementary Table 3
Supplementary Information (XLSX 276 kb)
Supplementary Table 4
Supplementary Information (XLSX 12 kb)
Supplementary Table 5
Supplementary Information (XLSX 20 kb)
Time-lapse of Esrrb-GFP cells.
The movie shows Esrrb-GFP cells and illustrates the continuous association of Esrrb with the mitotic chromosomes as cells enter, progress and exit mitosis. (WMV 1394 kb)
Rights and permissions
About this article
Cite this article
Festuccia, N., Dubois, A., Vandormael-Pournin, S. et al. Mitotic binding of Esrrb marks key regulatory regions of the pluripotency network. Nat Cell Biol 18, 1139–1148 (2016). https://doi.org/10.1038/ncb3418
Received:
Accepted:
Published:
Issue Date:
DOI: https://doi.org/10.1038/ncb3418
This article is cited by
-
Mitotic bookmarking by SWI/SNF subunits
Nature (2023)
-
Loss of H3K9 trimethylation alters chromosome compaction and transcription factor retention during mitosis
Nature Structural & Molecular Biology (2023)
-
The control of transcriptional memory by stable mitotic bookmarking
Nature Communications (2022)
-
Pioneer factors as master regulators of the epigenome and cell fate
Nature Reviews Molecular Cell Biology (2022)
-
Transcription factor Sp1 regulates mitotic chromosome assembly and segregation
Chromosoma (2022)